Introduction to Singlecrystal Diffraction Biology 555 Andrew J

Introduction to Single-crystal Diffraction Biology 555 Andrew J. Howard 11 September 2018

What we’ll discuss l Diffraction: why l Resolution l Crystallography: how l Experiment l Formalism – Lattice Symmetries – Cell matrices – Intensity relationships l Images and measurements l Rotation vs. Laue 09/11/2018 Single-crystal diffraction I p. 2 of 43

Determining 3 -D structures l We want to know the structures of macromolecules because we can understand function on the basis of structure – If we already know what the function is, we use the structure to figure out how it works – If we don’t know the function, we sometimes can use the structure to figure out what the function is. 09/11/2018 Single-crystal diffraction I p. 3 of 43

How do we do this? l If we already know something about structure, we can use a variety of techniques (e. g. light scattering, X-ray absorption, EPR , fiber diffraction) to answer specific structural questions l If we don’t know anything: – Low resolution (details > 3Å): cryo. EM, CD, SAXS, SANS – High resolution (details < 3Å) • single-crystal diffraction • multidimensional NMR 09/11/2018 Single-crystal diffraction I p. 4 of 43

What does resolution mean? l In general it refers to the distance associated with the smallest detail discernible from a particular experiment or approach l Specific significance depends on the technique involved l In crystallography it’s related to the widest angle of diffraction measured, via Bragg’s law 09/11/2018 Single-crystal diffraction I p. 5 of 43

Crystallography Determine 3 -D structure by: l Making a well-ordered 3 -D crystal l Identifying Bragg diffraction spots l Measuring their intensities l Correcting for systematic errors l Getting phase angles experimentally or by indirect reasoning l Calculating electron density by Fourier techniques l Relating electron density to atomic positions 09/11/2018 Single-crystal diffraction I p. 6 of 43

Crystal growth l We’ll discuss this in detail Tuesday l Recognize at the outset that protein crystals are artificial constructs: very few proteins naturally form 3 -D ordered arrays l But there are systematic ways to induce crystallization 09/11/2018 Single-crystal diffraction I p. 7 of 43

What does 3 -D order do? l Briefly, we say that the 3 -D order collapses what would otherwise be a fuzzy scattering pattern into a series of discrete spots l Each spot has 3 integer indices that characterize it (Why? 3 dimensions) l If we can identify each spot (determine the 3 integers) and measure the intensity of it, we have the Fourier amplitudes needed in the transform. 09/11/2018 Single-crystal diffraction I p. 8 of 43

How does the experiment work? l Last time we showed a simple-minded derivation of Bragg’s law based on planes of diffracting objects l This is an oversimplification but it yields nl = 2 D sinq l n = integer order of plane l l = incoming X-ray wavelength l D = spacing between planes l q = angle between diffracting plane and incoming or outgoing beam direction 09/11/2018 Single-crystal diffraction I p. 9 of 43
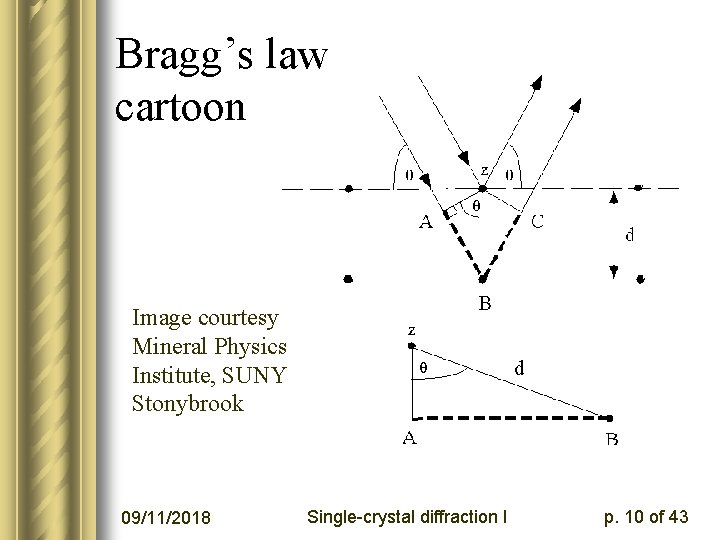
Bragg’s law cartoon Image courtesy Mineral Physics Institute, SUNY Stonybrook 09/11/2018 Single-crystal diffraction I p. 10 of 43
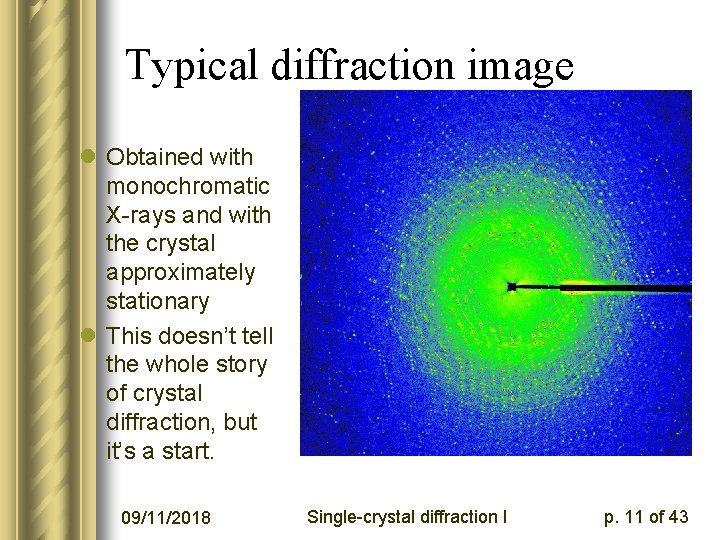
Typical diffraction image l Obtained with monochromatic X-rays and with the crystal approximately stationary l This doesn’t tell the whole story of crystal diffraction, but it’s a start. 09/11/2018 Single-crystal diffraction I p. 11 of 43

How would a small-molecule crystallographer react to that? l Too many spots! l In a typical small-molecule experiment with the crystal stationary, the Laue conditions are only satisfied for a very few Bragg spots—maybe 3. l So the small-molecule person would be impressed that hundreds of spots are visible in a still image 09/11/2018 Single-crystal diffraction I p. 12 of 43

Laue conditions for Bragg diffraction spots Laue condition for 3 -D crystals tells us: l Bragg diffraction occurs in discrete angular directions (rays spots) l Each spot can be identified with three integer indices (h, k, l), e. g. (-6, 11, -4) l For a given sample orientation, Laue condition is only satisfied for a few index values 09/11/2018 Single-crystal diffraction I p. 13 of 43

3 -space vectors l Get used to thinking about ordered triples of numbers as vectors l That’s natural with things like diffraction vectors, since they exist in a three-space l But we can even treat a triple of reflection indices (h, k, l) as a vector h. l Convention: boldface lowercase letters (e. g. s, k, h) denote vectors; boldface capital letters (e. g. A, R) denote square matrices 09/11/2018 Single-crystal diffraction I p. 14 of 43

What will we use this for? l We’re shooting electromagnetic waves exp(2 pik 0 • x) at a sample l Scatter with amplitude exp(2 pik • x) l Difference=exp(2 pis • x) for s = k - k 0 l I ~ |F|2 = [Sx f(x) exp(i(k • x-k 0 • x))]2 k k 0 09/11/2018 -k 0 s = k-k 0 Single-crystal diffraction I k p. 15 of 43

Using these structure factors to determine a structure l We need all of these structure factors to determine electron densities: l r(r) = Shkl Fhkl exp(-2 pi(s • r)) l Thus the electron density is the inverse Fourier transform of the structure factors l This is a triple sum over the three integer indices h = (h, k, l) l In principle it extends from -∞ to ∞ in all three indices; in practice it covers a narrower range 09/11/2018 Single-crystal diffraction I p. 16 of 43

Relating s to h l The diffraction vector s is related to the index vector h in a simple way: l s = l A • h where A is a 3 x 3 matrix describing the reciprocal-space unit cell associated with the real sample l This is a concept we’ll revisit later 09/11/2018 Single-crystal diffraction I p. 17 of 43

Relating h to s l A more intuitive description: l h = R • s / l where the index vector h (e. g. (6, -11, 4)) is related to the diffraction vector s via a 3 x 3 matrix R describing the real-space unit cell l Columns of R = unit cell lengths a, b, c l Dot products between columns yield the unit cell angles , , l Note that R = A-1 09/11/2018 Single-crystal diffraction I p. 18 of 43

Sample symmetry and R l The rotational symmetry of the unit cell gives rise to specific properties of the unit cell matrix R – If = =90º, ≠ 90º, it’s monoclinic – = = =90º, lengths unrestricted: orthorhombic – = = =90º, a=b: tetragonal – = =90º, 120º, a=b: hexagonal or trigonal – a=b=c, = = : rhombohedral – a=b=c, = = º: cubic l Those are the only possibilities, apart from redefinitions of what we mean by a, b, c. 09/11/2018 Single-crystal diffraction I p. 19 of 43
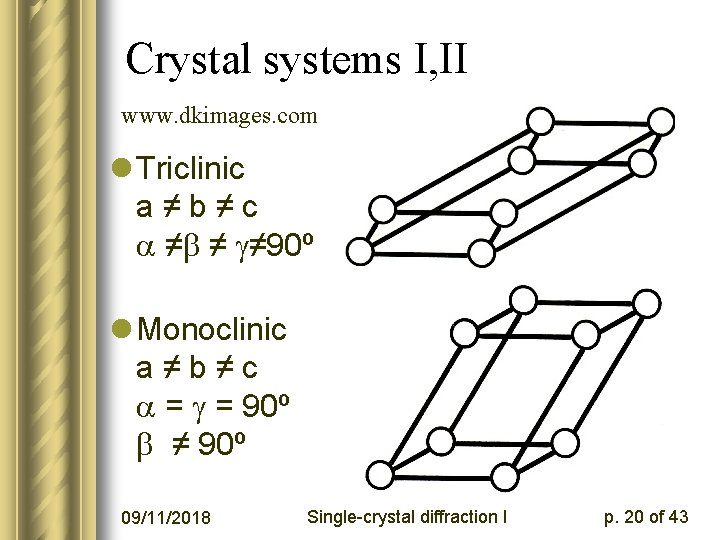
Crystal systems I, II www. dkimages. com l Triclinic a≠b≠c ≠ ≠ ≠ 90º l Monoclinic a≠b≠c = = 90º ≠ 90º 09/11/2018 Single-crystal diffraction I p. 20 of 43
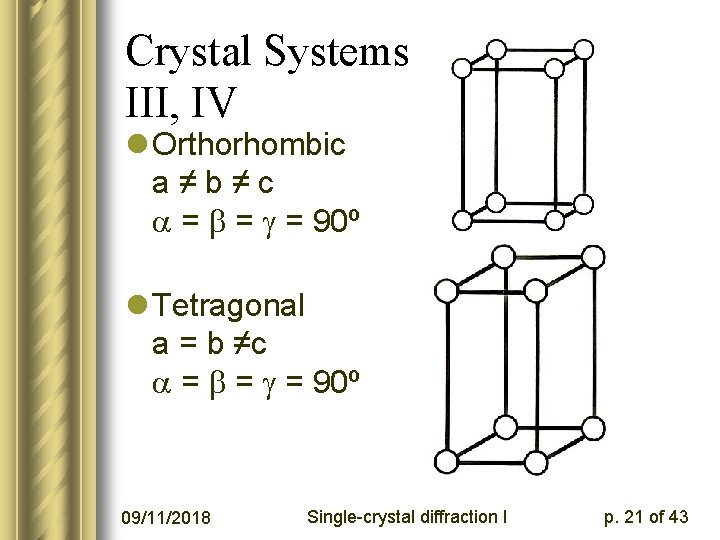
Crystal Systems III, IV l Orthorhombic a≠b≠c = = = 90º l Tetragonal a = b ≠c = = = 90º 09/11/2018 Single-crystal diffraction I p. 21 of 43
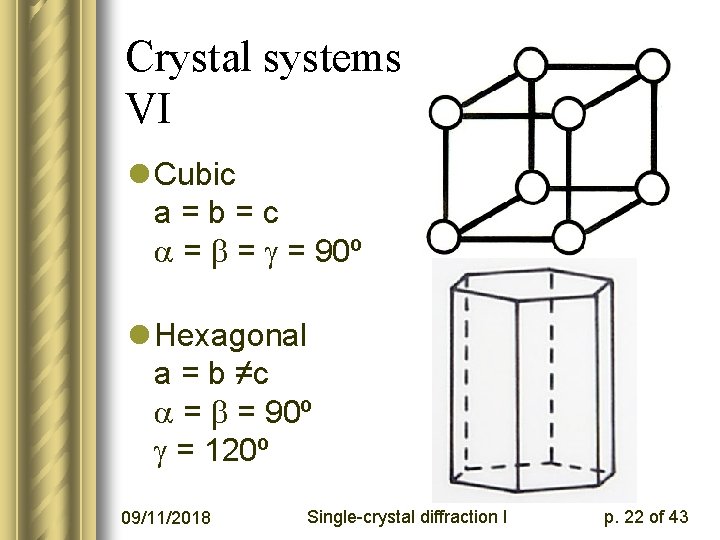
Crystal systems V, VI l Cubic a=b=c = = = 90º l Hexagonal a = b ≠c = = 90º = 120º 09/11/2018 Single-crystal diffraction I p. 22 of 43
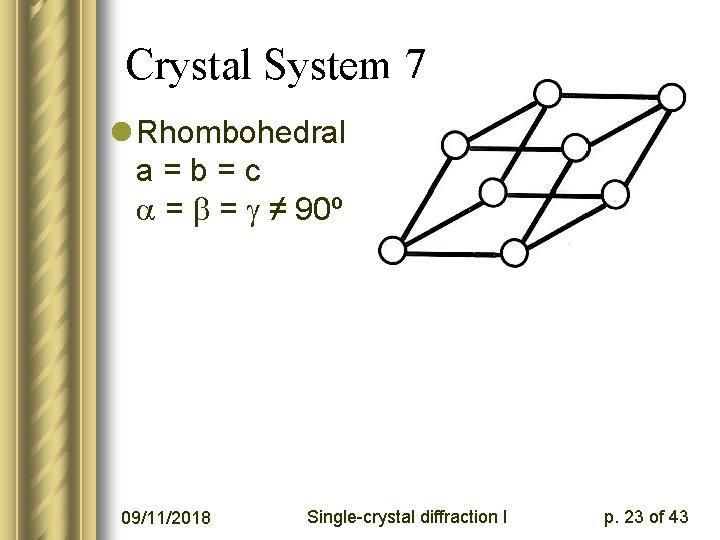
Crystal System 7 l Rhombohedral a=b=c = = ≠ 90º 09/11/2018 Single-crystal diffraction I p. 23 of 43

Crystal systems and symmetries l Crystals must have 3 -D translational order or they aren’t crystals l They may have rotational symmetries within the unit cell l The crystal systems are specifically associated with those symmetries 09/11/2018 Single-crystal diffraction I p. 24 of 43
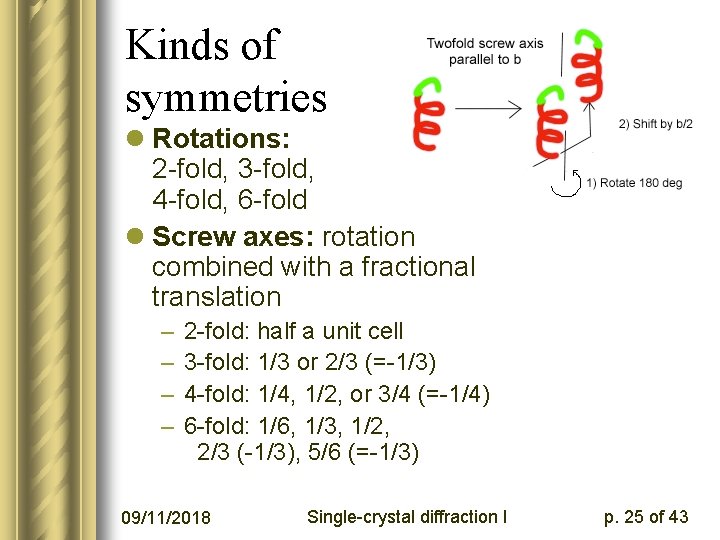
Kinds of symmetries l Rotations: 2 -fold, 3 -fold, 4 -fold, 6 -fold l Screw axes: rotation combined with a fractional translation – – 2 -fold: half a unit cell 3 -fold: 1/3 or 2/3 (=-1/3) 4 -fold: 1/4, 1/2, or 3/4 (=-1/4) 6 -fold: 1/6, 1/3, 1/2, 2/3 (-1/3), 5/6 (=-1/3) 09/11/2018 Single-crystal diffraction I p. 25 of 43

Other symmetries l Centers of symmetry: r -r l Mirrors: you can figure that out; A mirror through Y maps (x, y, z) into (x, -y, z) l Glide planes: center of symmetry combined with translation 09/11/2018 Single-crystal diffraction I p. 26 of 43

Specific symmetries l Triclinic: none l Monoclinic: twofold axis* about b l Orthorhombic: twofold axis* about a, b, c l Tetragonal: fourfold axis* about c l Cubic: threefold axis down body diagonal; twofold axis* perpendicular l Hexagonal: threefold* or sixfold axis* about c * can be screw or glide plane as well 09/11/2018 Single-crystal diffraction I p. 27 of 43

Additional symmetries l Tetragonal: Sometimes twofolds perpendicular to the fourfold axis of symmetry l Trigonal: Twofold perpendicular to threefold either of two positions l Hexagonal: twofold perpendicular to sixfold l Cubic: fourfold perpendicular to 3 and 2 09/11/2018 Single-crystal diffraction I p. 28 of 43

A consequence of chirality l Recognize that almost all biopolymers are chiral l That means that they cannot be identical to their mirror images l Therefore they can’t crystallize in centrosymmetric spacegroups, i. e. with symmetries that involve – Centers of symmetry (x, y, z) (-x, -y, -z) – Mirror planes (x, y, z) (x, -y, z) – Glide planes (trust me on this one) 09/11/2018 Single-crystal diffraction I p. 29 of 43

Is there a way around that? l Yes, if you’re dedicated enough. l Suppose you have a 105 -aa protein for which the phase problem (see later…) can’t be solved. l Synthesize your L-amino-acid protein using ribosomal synthesis and also synthesize a Damino acid version with nonbiological chemistry l Combine the L and D versions (1: 1 ratio) l Crystallize the resulting racemic mixture l If you’re lucky, it’ll crystallize in spacegroup P-1, so that the D protein and the L protein are at (x, y, z) and (-x, -y, -z): that then simplifies solving the phase problem! 09/11/2018 Single-crystal diffraction I p. 30 of 43

A simple R matrix l ( asin 0 a cos ) R= ( 0 b 0 ) ( 0 0 c ) l Monoclinic, b along Y, c along Z l Unit cell volume = det R = abcsin l As we rotate the crystal about X this matrix will be left-multiplied by a rotation matrix 09/11/2018 Single-crystal diffraction I p. 31 of 43

Rotated R matrix l The rotation matrix associated with a rotation W about X is (1 0 0 ) Q = ( 0 cos. W sin. W ) ( 0 -sin. W cos. W) l Thus R’ = QR = ( asin 0 a cos ) ( 0 bcos. W c sin. W ) ( 0 -bsin W c cos. W ) 09/11/2018 Single-crystal diffraction I p. 32 of 43

Realities of crystal symmetry l It occasionally happens that relationships in R that look like higher symmetries are not reflected in the underlying rotational symmetry l You can rely on relationships among intensities to sort that out. l These relationships among intensities are critical for merging data even without accidental correspondences. 09/11/2018 Single-crystal diffraction I p. 33 of 43

Intensity relationships l All unit cells: I(h, k, l) = I(-h, -k, -l) unless anomalous diffraction present (Friedel’s law) l Monoclinic: I(h, k, l) = I(-h, k, -l) l Orthorhombic: I(h, k, l) = I(h, -k, -l) = I(-h, k, -l)= I(-h, -k, l) l Tetragonal: I(h, k, l) = I(-k, h, l) = I(-h, -k, l)= I(k, -h, l) 09/11/2018 Single-crystal diffraction I p. 34 of 43

Intensity relationships, cont’d l Tetragonal with twofolds: All symmetry elements from orthorhombic and tetragonals l Cubic: I(h, k, l) = I(k, l, h) = I(l, h, k) plus orthorhombic relations l Cubic with fourfolds: As above, plus tetragonals l Hexagonal/ Trigonal: I(h, k, l) = I(k, i, l) = I(i, h, l) for i = -h-k 09/11/2018 Single-crystal diffraction I p. 35 of 43

Intensity relations, concluded l Trigonal with twofold (“ 312”): I(h, k, l) = I(k, i, l) = I(i, h, l) = I(-k, -h, -l) = I(-i, -k, -l) = I(-h, -i, -l) l Trigonal with twofold (“ 321”): I(h, k, l) = I(k, i, l) = I(i, h, l) = I(k, h, -l) = I(i, k, -l) = I(h, i, -l) l Hexagonal: I(h, k, l) = I(k, i, l) = I(i, h, l) = I(-h, -k, l) = I(-k, -i, l) = I(-i, -h, l) l Hexagonal with twofolds: as above plus those also equal I(-k, -h, -l) = I(-i, -k, -l) = I(-h, -i, -l) = I(k, h, -l) = I(i, k, -l) = I(h, i, -l) 09/11/2018 Single-crystal diffraction I p. 36 of 43

A single monochromatic shot won’t tell us about all the spots l Since the Laue condition is only satisfied for a tiny fraction of the total list of spots at any given sample position, to measure all the spots we need many different sample positions or many different wavelengths – Many positions: rotate the sample – Many wavelengths: polychromatic diffraction 09/11/2018 Single-crystal diffraction I p. 37 of 43

Which is better? They both have their uses l Polychromatic (“Laue”) takes advantage of inherently polychromatic incoming radiation l Rotation is easier to understand what the individual spots mean l Quality of data is often higher with rotation data: see next slide 09/11/2018 Single-crystal diffraction I p. 38 of 43
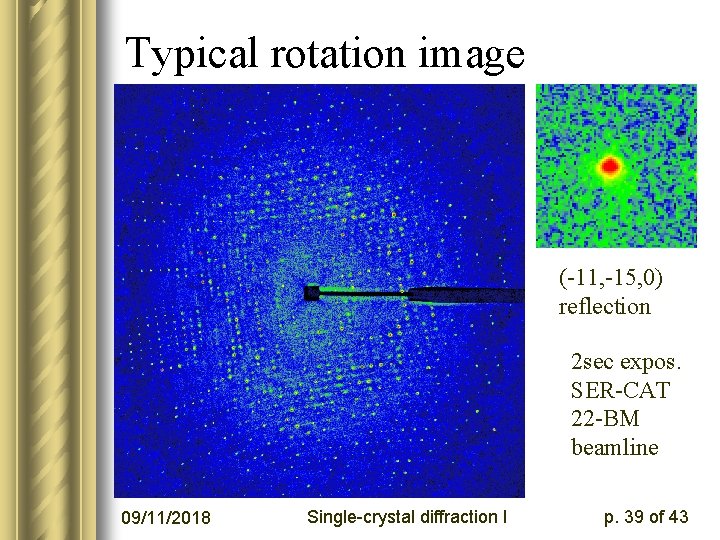
Typical rotation image (-11, -15, 0) reflection 2 sec expos. SER-CAT 22 -BM beamline 09/11/2018 Single-crystal diffraction I p. 39 of 43

Simulated Polychromatic Image Courtesy Renz Research, Inc. 09/11/2018 Single-crystal diffraction I p. 40 of 43

Why is rotation data often of higher quality? l Individual atomic scattering amplitudes f are themselves wavelength-dependent; that dependence scales away if l constant or close to constant l Detector response is wavelength dependent l Background scatter is often higher with Laue 09/11/2018 Single-crystal diffraction I p. 41 of 43

Rotation vs. Laue, concluded l Sometimes spots overlap harmonically in Laue; that problem doesn’t occur in monochromatic experiments l Spatial overlaps (even without harmonic overlaps) are worse in Laue l With conventional sources, we can’t really get much intensity from Bremsstrahlung, so we’re stuck 09/11/2018 Single-crystal diffraction I p. 42 of 43

Special use of Laue l The phenomenally high fluence achievable with Laue means that one can obtain a complete diffraction pattern in microseconds or even nanoseconds of exposure l If you only need one pattern to determine the structure (which is only occasionally true!) this enables you to determine a structure in a single shot l This allows for true time-resolved crystallography 09/11/2018 Single-crystal diffraction I p. 43 of 43
- Slides: 43