Introduction to NCBI database and phylogeny in MEGA

Introduction to NCBI database and phylogeny in MEGA using SARS-Co. V-2 and other pathogens Nicholas Lorusso, Maria Shumskaya, Kean University, 1000 Morris Ave, Union, NJ 07083

• An evolutionary hypothesis of relationships in graphical form • Shows the origin and relationship of genetic groups at the level of population, species, or higher taxon. • The tips of the branches are called: TERMINALS • The intersections are called: NODES • Synonyms: phylogentic tree, phylogenetic hypothesis, cladogram, evolutionary tree. Hypothesis 1 Hypothesis 2

Parts of a phylogeny are used to show possible relationships Is C more closely related to D or E? Clade= Monophyletic Group= Natural Group =a group of species that includes an ancestral species and ALL of its descendants
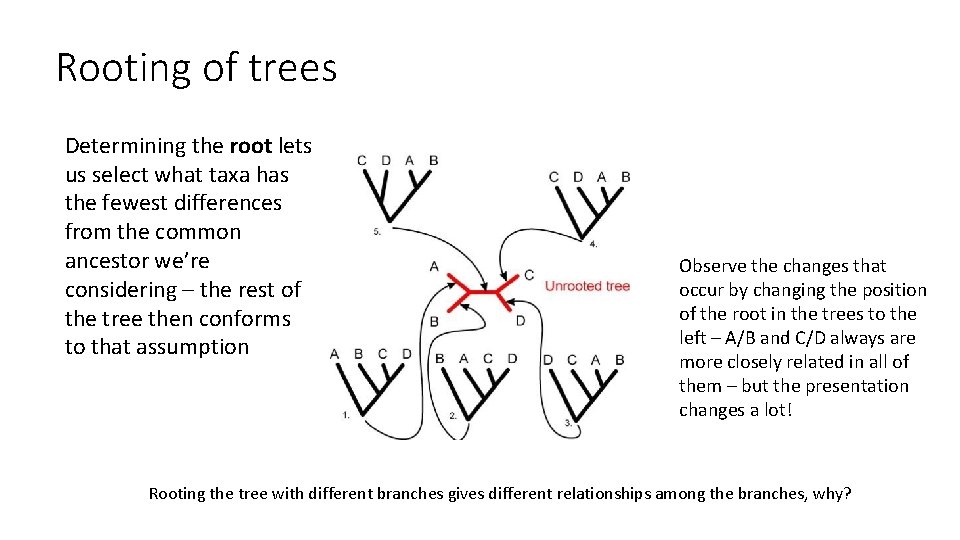
Rooting of trees Determining the root lets us select what taxa has the fewest differences from the common ancestor we’re considering – the rest of the tree then conforms to that assumption Observe the changes that occur by changing the position of the root in the trees to the left – A/B and C/D always are more closely related in all of them – but the presentation changes a lot! Rooting the tree with different branches gives different relationships among the branches, why?

Tree variations – Some trees look different but say the same thing Campbell Biology, 11 th
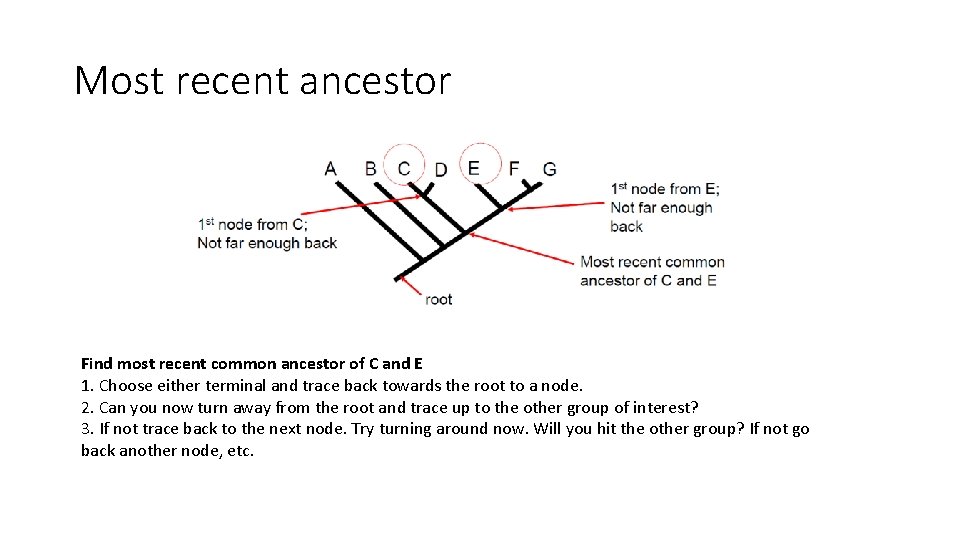
Most recent ancestor Find most recent common ancestor of C and E 1. Choose either terminal and trace back towards the root to a node. 2. Can you now turn away from the root and trace up to the other group of interest? 3. If not trace back to the next node. Try turning around now. Will you hit the other group? If not go back another node, etc.
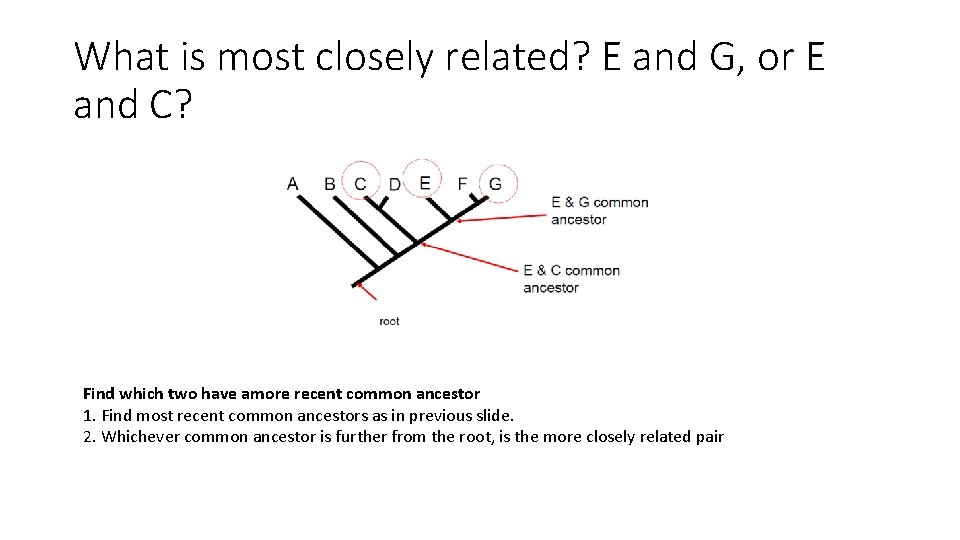
What is most closely related? E and G, or E and C? Find which two have amore recent common ancestor 1. Find most recent common ancestors as in previous slide. 2. Whichever common ancestor is further from the root, is the more closely related pair

Making phylogenies • Parsimony • the simplest explanation is the solution that requires the fewest assumptions, and is therefore the best supported explanation. • Fewest assumptions = Fewest character (ex. C, T, A, G) changes • Likelihood • statistical method; most probable tree is chosen, based on the data, and on the chosen evolutionary model. • Needs models for evolution of nucleotides such as knowing how likely a Cytosine changing to a Guanine is

Application: What can we learn about SARSCo. V-2 which causes COVID-19 • Viruses can be difficult to study due to their small size and rapid generation times • We need to given how viruses can effect health, agriculture, biodiversity… • Luckily their genomes are small! • We can compare their sequences to find out if a current virus has something in common with other viruses we know already • Using phylogenetic methods allows us to evaluate these relationships • This can speed things like vaccine development!

SARS-Co. V-2 is a novel coronavirus • Similar known coronaviruses: • SARS-Co. V, caused SARS epidemic in China in 2002 • MERS, caused MERS epidemic in Middle East in 2012 • Coronaviruses from other animals, such as bats, pangolin etc. • Let’s build a phylogeny to investigate the ancestry of SARS-Co. V-2 and establish its relationships to other coronaviruses
MEGA and publically available data • We’ll be using a program called MEGA (Molecular Evolutionary Genetics Analysis) to build our predictive phylogenies • We’ll take sequences, align them to each other (so we can compare them), then build two different types of predictive phylogeny • The data we’re using is thanks to the collaborative effort of scientists around the world • Sources like the National Center for Biotechnology Information (NCBI) allowed us to prepare a dataset of RNA sequences for you to analyze • Most of the sequences available for coronaviruses are full-length genomes, which we trimmed for teaching purposes
- Slides: 11