Introduction to Haplotype Estimation StatBiostat 550 The Haplotype

Introduction to Haplotype Estimation Stat/Biostat 550

The Haplotype Problem • Suppose we genotype individuals at a number of tightly linked SNPs. A C G C C T T T G C G A A C C C A G G C

The Haplotype Problem • Suppose we genotype individuals at a number of tightly linked SNPs. A C G C C T T T G C G A A C C C A G G C

The Haplotype Problem • Suppose we genotype individuals at a number of tightly linked SNPs.

The Haplotype Problem • What do the types on the two chromosomes look like?

The Haplotype Problem • What do the types on the two chromosomes look like?

The Haplotype Problem • What do the types on the two chromosomes look like?

The Haplotype Problem • What do the types on the two chromosomes look like?

The Haplotype Problem • What do the types on the two chromosomes look like?

Haplotypes: who cares? Many people, for many different reasons… • LD mapping: increase power? • LD mapping: decrease genotyping? • Evolutionary studies: selection, recombination, gene conversion, population structure, …

The Haplotype Problem – potential solutions • Molecular methods • Collect family data • Statistical methods for population data

The Simplest Case • What do the types on the two chromosomes look like?

The Next Simplest Case • What do the types on the two chromosomes look like?

The Next Simplest Case • What do the types on the two chromosomes look like?

The first difficult case… • What do the types on the two chromosomes look like?

The first difficult case… • What do the types on the two chromosomes look like?

Clark’s Method (1990) • Idea: use information obtained from other individuals in the population to determine the most probable haplotype pair.

1 2 3 Is it this configuration?

1 2 3 …or this one?

1 2 3 This one is more probable.

Clark’s Method (Clark, 1990) • Identify the unambiguous individuals. • Make a list of “known” haplotypes. • Go through list, and see whether ambiguous individuals can be made up from a “known” haplotype plus another “complementary” haplotype. If so, add the complementary haplotype to the list of “known” haplotypes.

Clark’s Method 1 2 3 List of known haps.

Clark’s Method 1 2 3 List of known haps.

Clark’s Method: Problem 1 1 2 3

Clark’s Method: Problem 1 1 2 3 List of known haps.

Clark’s Method: Problem 1 1 2 3 List of known haps.

Clark’s Method: Problem 1 1 2 3 List of known haps.

Clark’s Method: Problem 1 1 2 3 List of known haps.

Clark’s Method: Problem 1 1 List of known haps. 2 3 Answer depends on order list is considered…. … and frequency information is ignored

Clark’s Method: Problem 2 1 2 3

Clark’s Method: Problem 2 1 List of known haps. 2 3 Algorithm can fail to resolve all haplotypes… … because looks only for exact matches

Clark’s Algorithm: Summary • Results may depend on order individuals are considered. • Frequency information is ignored. • May fail to resolve all haplotypes. • Fails to assess uncertainty. • Looks only for exact matches. • Fast and intuitive(? ).

Maximum Likelihood (EM Algorithm) • Idea: find haplotype frequencies (f 1, …f. N) to maximise probability of observed genotype data (g 1, …, gn).

Bayesian version Modify Clark’s algorithm: • Replace single pass through data, with iterative scheme. • Allow for uncertainty in resolution. • Use frequency information. Resulting “naïve Gibbs sampler” produces results similar to EM (Stephens, Smith and Donnelly 2001).

Example 1 List of known haps. 3 1 2 3 Matches 1 known Does not match any Assigned moderate probability

Example 1 List of known haps. 3 1 2 3 Matches 3 known Does not match any Assigned higher probability

Example 1 List of known haps. 3 1 2 3 Does not match any Assigned low probability

Problems with EM/naïve Gibbs • Potentially (very) large number of parameters to estimate, leading to inaccurate estimates. • Can be time-consuming for large problems. • Can “converge” to poor local optima (alleviated by multiple runs).

Further modification • Take into account “near misses”, as well as exact matches. (PHASE v 1. 0: Stephens, Smith and Donnelly 2001)

Example 1 2 3 List of known haps. 3 1 Matches 1 known Differs by 2 from 3 known

Example 1 2 3 List of known haps. 3 1 Matches 3 known Differs by 2 from 1 known
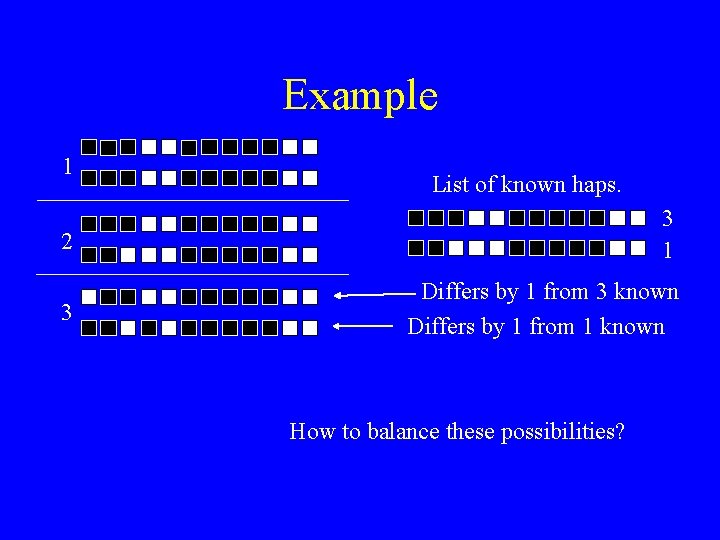
Example 1 List of known haps. 2 3 1 3 Differs by 1 from 3 known Differs by 1 from 1 known How to balance these possibilities?

The key question • What is the conditional distribution of the next haplotype, given a set of known haplotypes?

Example 1 2 Given the above haplotypes, what would you expect the next haplotype to look like?

Qualitative answer • The next haplotype will likely differ by a small number of mutations (possibly 0 mutations) from a (randomly-chosen) existing haplotype. • Use theory (Ewens sampling formula; coalescent theory) to roughly quantify the distribution of the “small number”.

Comparisons on simulated data


Problems • Time-consuming for large problems. • Can “converge” to poor local optima. • Ignores recombination (decay of LD with distance). • How should uncertainty in haplotype estimates be treated?

… to be continued.
- Slides: 49