Introduction to Free Surfer Dustin Clark clarkdcbu edu

Introduction to Free. Surfer Dustin Clark clarkdc@bu. edu Research Computing Services Information Services & Technology Boston University
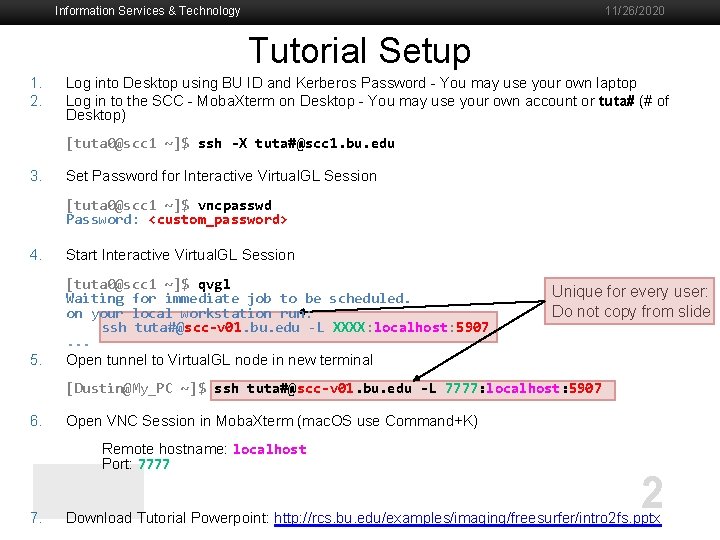
Information Services & Technology 11/26/2020 Tutorial Setup 1. 2. Log into Desktop using BU ID and Kerberos Password - You may use your own laptop Log in to the SCC - Moba. Xterm on Desktop - You may use your own account or tuta# (# of Desktop) [tuta 0@scc 1 ~]$ ssh -X tuta#@scc 1. bu. edu 3. Set Password for Interactive Virtual. GL Session [tuta 0@scc 1 ~]$ vncpasswd Password: <custom_password> 4. Start Interactive Virtual. GL Session 5. [tuta 0@scc 1 ~]$ qvgl Waiting for immediate job to be scheduled. on your local workstation run: ssh tuta#@scc-v 01. bu. edu -L XXXX: localhost: 5907. . . Open tunnel to Virtual. GL node in new terminal Unique for every user: Do not copy from slide [Dustin@My_PC ~]$ ssh tuta#@scc-v 01. bu. edu -L 7777: localhost: 5907 6. Open VNC Session in Moba. Xterm (mac. OS use Command+K) Remote hostname: localhost Port: 7777 7. 2 Download Tutorial Powerpoint: http: //rcs. bu. edu/examples/imaging/freesurfer/intro 2 fs. pptx
Information Services & Technology 11/26/2020 RCS Team and Expertise § Our Team § Scientific Programmers § Systems Administrators § Graphics/Visualization Specialists § Account/Project Managers § Special Initiatives (Grants) § Maintains and administers the Share Computing Cluster § Located in Holyoke, MA § ~14, 000 CPUs running Linux § Consulting Focus: § § Bioinformatics Data Analysis / Statistics Molecular Modeling Geographic Information Systems § Scientific / Engineering Simulation § Visualization § Contact Us: help@scv. bu. edu 3
Information Services & Technology 11/26/2020 What is Free. Surfer? … … 4
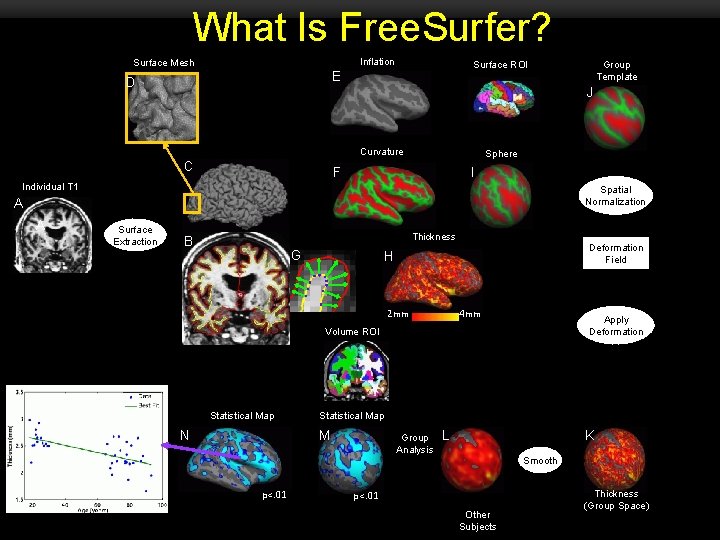
What Is Free. Surfer? Inflation Surface Mesh E D J Curvature C Sphere F I Individual T 1 Spatial Normalization A A Group Template Surface ROI Surface Extraction Thickness B G Deformation Field H 2 mm 4 mm Apply Deformation Volume ROI O Statistical Map N Statistical Map M p<. 01 Group Analysis L K Smooth p<. 01 Other Subjects Thickness (Group Space)
Information Services & Technology 11/26/2020 Goals Learn how to use the Free. Surfer for neuroimaging analysis § § § § § Introduction/Motivation Navigate FS Documentation Learn FS Jargon Approach Neuroimaging with both Volume and Surface models Optimize recon-all runtime Brain Editing ROI Analysis Anatomical Analysis Functional Analysis
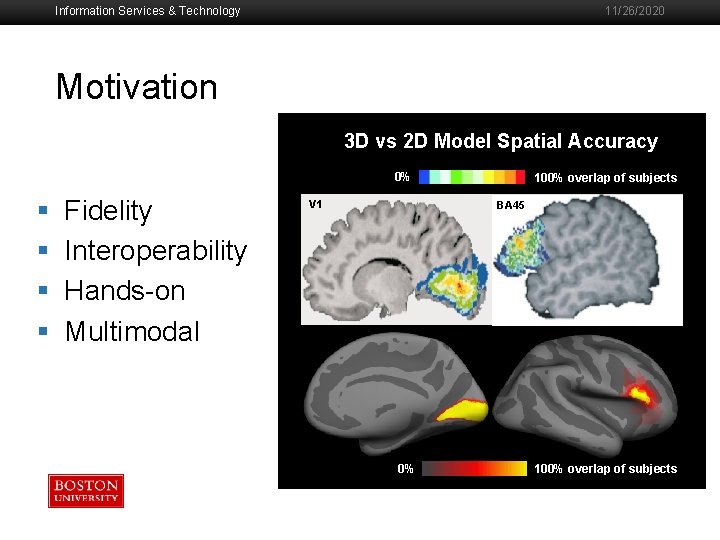
Information Services & Technology 11/26/2020 Motivation 3 D vs 2 D Model Spatial Accuracy 0% § § Fidelity Interoperability Hands-on Multimodal V 1 100% overlap of subjects BA 45 0% 100% overlap of subjects
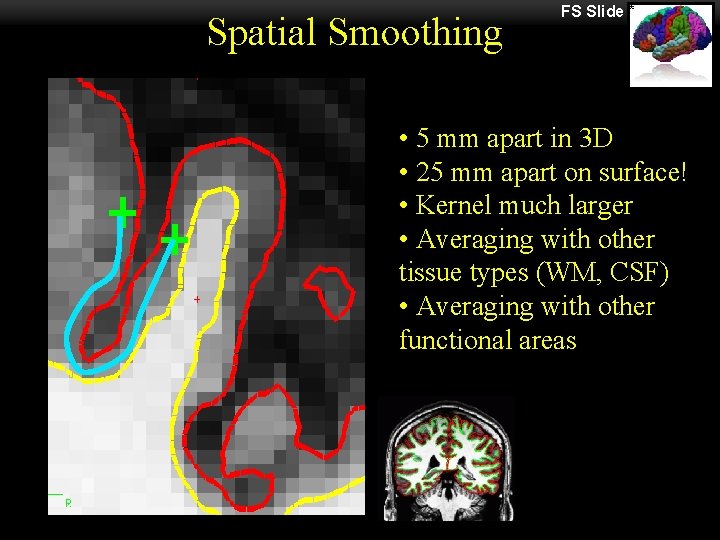
Spatial Smoothing FS Slide * • 5 mm apart in 3 D • 25 mm apart on surface! • Kernel much larger • Averaging with other tissue types (WM, CSF) • Averaging with other functional areas
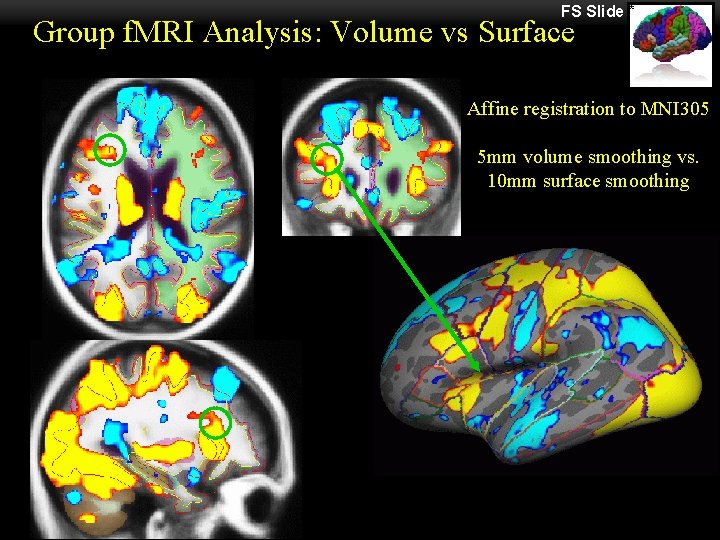
FS Slide * Group f. MRI Analysis: Volume vs Surface Affine registration to MNI 305 5 mm volume smoothing vs. 10 mm surface smoothing
Information Services & Technology Free. Surfer Resources 1. 2. 3. 4. 5. 6. 7. Free. Surfer Tutorial and Workshop Free. Surfer Tutorials Free. Surfer Videos Andrew Jahn Google Free. Surfer Archives Free. Surfer Wiki 11/26/2020
Information Services & Technology Free. Surfer Navigation Map 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. Introduction to Free. Surfer (video) Introduction to Free. Surfer Jargon (video) Analyzing the Individual Subject Part 1 (video) Analyzing the Individual Subject Part 2 (video) Inspection of Free. Surfer Output (walkthrough) Free. Surfer Troubleshooting (video) Troubleshooting your output (walkthrough) Surface-based analysis (video) Group Analysis 1 (video) Group Analysis 2 (video) Group Analysis (walkthrough) Multiple Comparisons Correction (video) Multiple Comparisons Correction (walkthrough) ROI Analysis (video) ROI Analysis (walkthrough) Multi-Modal Integration Part 1 (video) Multi-Modal Integration Part 2 (video) Multi-Modal Integration (walkthrough) 11/26/2020

Information Services & Technology 11/26/2020 Free. Surfer Jargon § § § Voxel - pixel with volume (location and § value) Surface - boundary (wm/gm, § gm(pial)/csf) § § Made up of many tiny points - vertices § Vertex - makes up surface, stores information Recon - output of data/make surfaces § Volumes - files (mgz, nii. gz) Cortical gm (ribbon) - grey matter Subcortical gm hippocampus/amygdala, etc. Segmentation - subcortical Parcellation - cortical ribbon Spatial alignment Tool/algorithm Find a common coordinate system for images being analyzed (. lta files) Intersubject/intrasubject FLIRT DOF § § CVS reg (combined volumetric and surface based) nonlinear § § § Rotation, displacement, scaling, shearing Same subject, same day - not expecting any scaling or shearing Higher local correspondence, better accuracy Input dataset - fixed/template/target, moving/subject Using registration matrix on other images - deform, morph, resample, transform

Information Services & Technology 11/26/2020 Analyzing the Individual Subject § All pipelines assume you have taken your T 1 image and reconstructed surfaces with it § Processing stream contains volume/surface § T 1 - white matter brighter than gray matter § MPRAGE on Siemens, SPGR on GE § High resolution could be worse (loss of snr in individual voxels, anatomical variation not needed like differences in white matter) § Reconstruction = ‘anatomical analysis of brain’ § Upon completion, creates directory structure
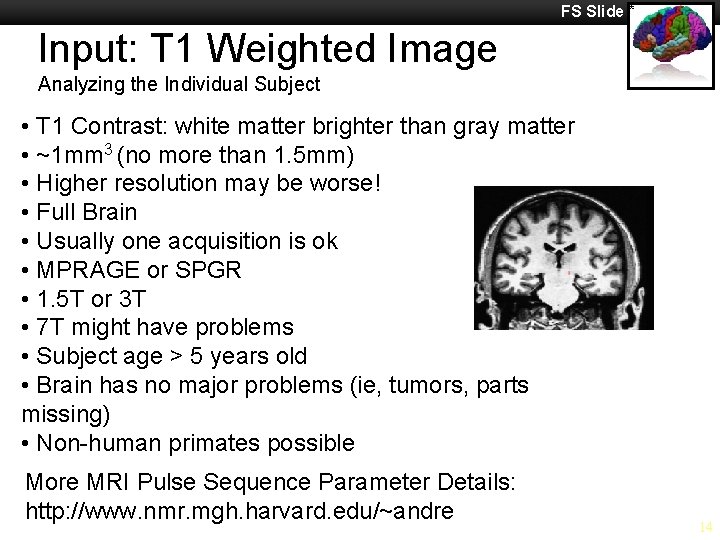
FS Slide * Input: T 1 Weighted Image Analyzing the Individual Subject • T 1 Contrast: white matter brighter than gray matter • ~1 mm 3 (no more than 1. 5 mm) • Higher resolution may be worse! • Full Brain • Usually one acquisition is ok • MPRAGE or SPGR • 1. 5 T or 3 T • 7 T might have problems • Subject age > 5 years old • Brain has no major problems (ie, tumors, parts missing) • Non-human primates possible More MRI Pulse Sequence Parameter Details: http: //www. nmr. mgh. harvard. edu/~andre 14

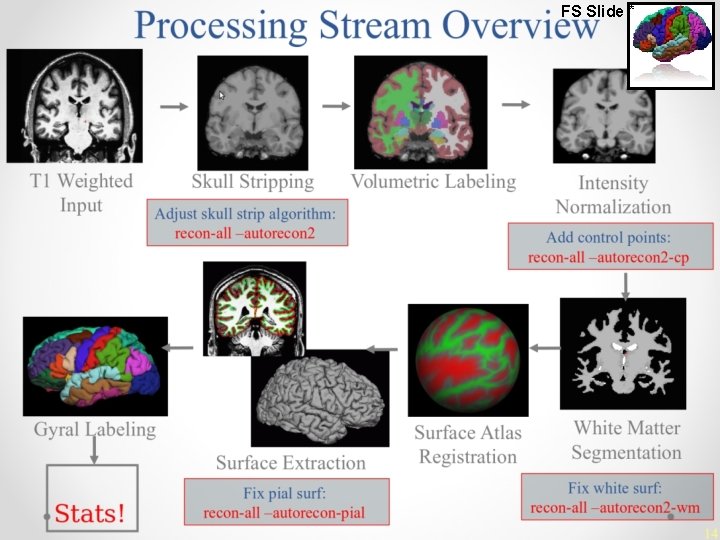
Information Services & Technology 11/26/2020 recon-all 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. Motion Correction and Conform 19. NU (Non-Uniform intensity normalization) 20. Talairach transform computation 21. Intensity Normalization 1 22. Skull Strip 23. EM Register (linear volumetric registration) 24. CA Intensity Normalization 25. CA Non-linear Volumetric Registration 26. Remove Neck LTA with Skull CA Label (Volumetric Labeling, ie Aseg) and 27. 28. Statistics Intensity Normalization 2 (start here for control 29. points) 30. White matter segmentation 31. Edit WM With ASeg Fill (start here for wm edits) Tessellation (begins per-hemisphere operations) Smooth 1 Inflate 1 QSphere Automatic Topology Fixer Final Surfs (start here for brain edits for pial surf) Smooth 2 Inflate 2 Spherical Mapping Spherical Registration, Contralateral hemisphere Map average curvature to subject Cortical Parcellation - Desikan_Killiany and Christophe (Labeling) Cortical Parcellation Statistics Cortical Ribbon Mask Cortical Parcellation mapping to Aseg 15
Information Services & Technology 11/26/2020 FS Slide * 16

Information Services & Technology 11/26/2020 Inspection of Free. Surfer Output § Introduces Free. Surfer Visualization tool: § Freeview (replaces tksurfer) § Introduces FS Volume and Surface nomenclature: § Volumes: T 1. mgz, wm. mgz, brainmask. mgz, aseg. mgz § Surfaces: Lh. white, rh. white, lh. pial, rh. pial § Navigating Freeview § Changing views, scroll through slices, manipulating volumes § Checking the Surfaces, Segmentation, Skull Strip, Intensity Normalization, WM Volume 17

FS Slide * Volume vs Surface Model Inspection of Free. Surfer Output Volume • uniform grid • voxel is an intersection of grid lines • columns, rows, slices • voxel size/distance • voxel assigned a value • XYZ Surface • NON-uniform grid • vertex is an intersection of triangles • each vertex has an index • distance between vertices ~1 mm • vertex assigned a value • XYZ • Vector of vertex values (~140, 000) 18

Volume vs Surface Model Inspection of Free. Surfer Output Volume FYI • Rawavg. mgz is in native anatomical space • Conformed space: 1 x 1 x 1 mm 3, 256 x 256 • Intensity bias from # coils corrected in T 1. mgz • Wm. mgz is everything but cortex, used for surface model Surface Steps • White surface • Orig surface • Pial surface • Inflation • Thickness • Curvature • Used for registration 19

Information Services & Technology 11/26/2020 Inspection of Free. Surfer Output § Viewing Volumes [tuta 0@scc-v 01 ~]$ module load freesurfer [tuta 0@scc-v 01 ~]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs [tuta 0@scc-v 01 ~]$ cd $SUBJECTS_DIR [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -v good_output/mri/T 1. mgz good_output/mri/wm. mgz good_output/mri/brainmask. mgz good_output/mri/aseg. mgz: colormap=lut: opacity=0. 2 -f good_output/surf/lh. white: edgecolor=blue good_output/surf/lh. pial: edgecolor=red good_output/surf/rh. white: edgecolor=blue good_output/surf/rh. pial: edgecolor=red & 20
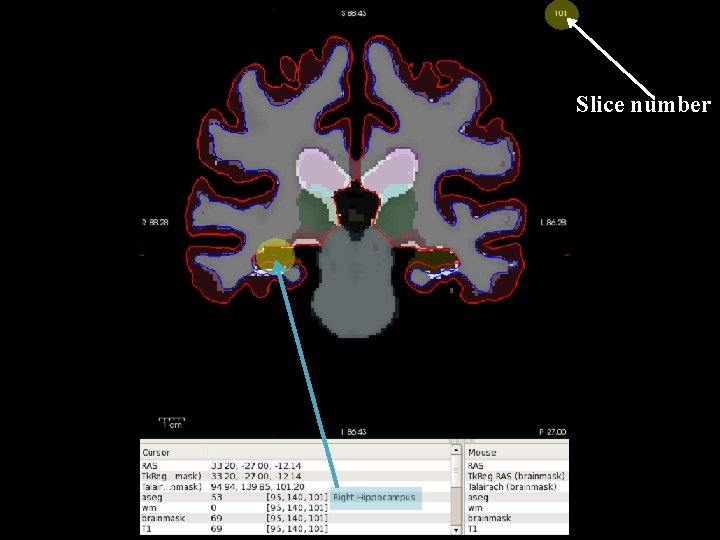
Slice number

Information Services & Technology 11/26/2020 Inspection of Free. Surfer Output § Viewing the Surface [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -f good_output/surf/lh. pial: annot=aparc. annot: name=pial_aparc: visible=0 good_output/surf/lh. pial: annot=aparc. a 2009 s. annot: name=pial_aparc_des: visible=0 good_output/surf/lh. inflated: overlay=lh. thickness: overlay_threshold=0. 1, 3: : name=inflated_ thickness: visible=0 good_output/surf/lh. inflated: visible=0 good_output/surf/lh. white: visible=0 good_output/surf/lh. pial --viewport 3 d & 22

Information Services & Technology 11/26/2020 Brain Editing Troubleshooting Your Output § “Brain Editing” § Fixing mistakes in white and pial surfaces are inaccurate § Edits cannot be directly on the surface, they are on volumes which are used to redraw the surfaces with another run of recon-all § “Do the surfaces follow gray matter/white matter borders? ” § “Does the subcortical segmentation follow intensity boundaries? 23
Information Services & Technology 11/26/2020 Brain Editing Troubleshooting Your Output § 5 main types of errors 1. 2. 3. 4. 5. § Pial Surface misplacement Segmentation Errors Skull Strip Errors Intensity Normalization Error Topological Defect 4 Fixes: 1. 2. 3. 4. Erase Voxels Fill Voxels Clone Voxels (copy from another volume) Add “Control Points” 24

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § Pial Surface Error: Erase Voxels § Pial Surface deviates from gray matter/CSF boundary § Editing: brainmask. mgz [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -v pial_edits_before/mri/T 1. mgz pial_edits_before/mri/brainmask. mgz -f pial_edits_before/surf/lh. white: edgecolor=yellow pial_edits_before/surf/lh. pial: edgecolor=red pial_edits_before/surf/rh. white: edgecolor=yellow pial_edits_before/surf/rh. pial: edgecolor=red & $ recon-all -autorecon-pial -subjid pial_edits_before $ cp brain. finalsurfs. mgz brain. finalsurfs. manedit. mgz 25
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § Segmentation Error: Fill Voxels § White matter surface cuts into wm. mgz § Editing: wm. mgz [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -v wm 1_edits_before/mri/brainmask. mgz wm 1_edits_before/mri/wm. mgz: colormap=heat: opacity=0. 4 -f wm 1_edits_before/surf/lh. white: edgecolor=yellow wm 1_edits_before/surf/lh. pial: edgecolor=red wm 1_edits_before/surf/rh. white: edgecolor=yellow wm 1_edits_before/surf/rh. pial: edgecolor=red wm 1_edits_before/surf/rh. inflated: visible=0 wm 1_edits_before/surf/lh. inflated: visible=0 & $ recon-all -autorecon 2 -wm -autorecon 3 -subjid wm 1_edits_before 26
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § White Surface Error: Fill/Delete/Control § White Surface doesn’t capture wm. mgz § Editing: wm. mgz [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -v topo_defect_before/mri/brainmask. mgz topo_defect_before/mri/wm. mgz: colormap=heat: opacity=0. 4 -f topo_defect_before/surf/lh. white: edgecolor=yellow topo_defect_before/surf/lh. pial: edgecolor=red topo_defect_before/surf/rh. white: edgecolor=yellow topo_defect_before/surf/rh. pial: edgecolor=red & $ recon-all -autorecon 2 -wm -autorecon 3 -subjid topo_defect_before 27
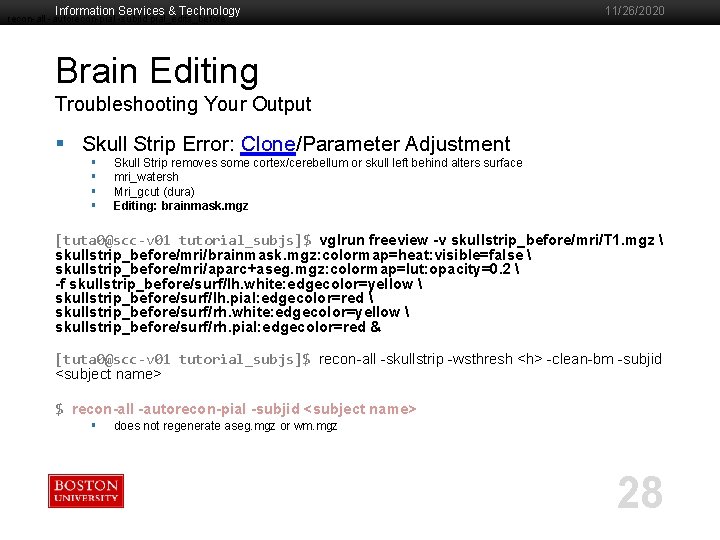
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § Skull Strip Error: Clone/Parameter Adjustment § § Skull Strip removes some cortex/cerebellum or skull left behind alters surface mri_watersh Mri_gcut (dura) Editing: brainmask. mgz [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -v skullstrip_before/mri/T 1. mgz skullstrip_before/mri/brainmask. mgz: colormap=heat: visible=false skullstrip_before/mri/aparc+aseg. mgz: colormap=lut: opacity=0. 2 -f skullstrip_before/surf/lh. white: edgecolor=yellow skullstrip_before/surf/lh. pial: edgecolor=red skullstrip_before/surf/rh. white: edgecolor=yellow skullstrip_before/surf/rh. pial: edgecolor=red & [tuta 0@scc-v 01 tutorial_subjs]$ recon-all -skullstrip -wsthresh <h> -clean-bm -subjid <subject name> $ recon-all -autorecon-pial -subjid <subject name> § does not regenerate aseg. mgz or wm. mgz 28

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § Intensity Normalization Error: Control Points § Intensity normalization can fail if wm intensity cant be determined § Control point is a user selected location of wm boundary that should be normalized to an intensity of 110 § Editing: subjid/tmp/control. dat [tuta 0@scc-v 01 tutorial_subjs]$ vglrun freeview -v cp_before/mri/brainmask. mgz -f cp_before/surf/lh. white: edgecolor=yellow cp_before/surf/lh. pial: edgecolor=red cp_before/surf/rh. white: edgecolor=yellow cp_before/surf/rh. pial: edgecolor=red & $ recon-all -autorecon 2 -cp -autorecon 3 -subjid cp_before 29

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § § Pial Surface Error: Erase Voxels Segmentation Error: Fill Voxels White Surface Error: Fill/Delete/Control Skull Strip Error: Clone/Parameter Adjustment § Intensity Normalization Error: Control Points § Less common: Talairach, Aseg 30

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Brain Editing Troubleshooting Your Output § Tips: § Look at all 3 views scrolling back and forth through slices § 30 minutes or less § Leave it if unsure § Less is more § Copy your subject and only edit copy § Don’t erase pial when editing brainmask 31

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Group Analysis § Thickness, Functional, PET, MEG/EEG, Diffusion § Basic Processing Stages: 1. 2. 3. 4. 5. 6. Subject & Measurement Assemble Data § Resample into common space § Smooth § Concatenate Data into one file General Linear Model § Specify covariates/demographics (design matrix or FSGD) § Specify contrast (the query of the model) Fit the Model Multiple Comparisons Correction Visualize 32

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Group Analysis - Anatomical § Surface-based thickness study § 40 Subjects from Randy Buckner’s lab [tuta 0@scc-v 01 tutorial_subjs]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/ tutorial_subjs/group_analysis_tutorial [tuta 0@scc-v 01 tutorial_subjs]$ cd $SUBJECTS_DIR/glm 33
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Assemble the Data Group Analysis - Anatomical § mris_preproc § Prepares surface-based data for high-level analysis § Resamples surface or volume source data to a common subject (usually an average subject) § Concatenates them into one file to be used by a number of programs (eg, mri_glmfit). [tuta 0@scc-v 01 glm]$ mris_preproc --fsgd gender_age. fsgd --target fsaverage --hemi lh --meas thickness --out lh. gender_age. thickness. 00. mgh 34

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Assemble the Data Group Analysis - Anatomical § mri_surf 2 surf § Resample surface to another space, smooth a surface § recon-all -s subjid -all -qcache for thickness, curv, sulc, area, and jacobian_white [tuta 0@scc-v 01 glm]$ mri_surf 2 surf --s fsaverage --sval lh. gender_age. thickness. 00. mgh --fwhm 10 --cortex --tval lh. gender_age. thickness. 10 B. mgh 35
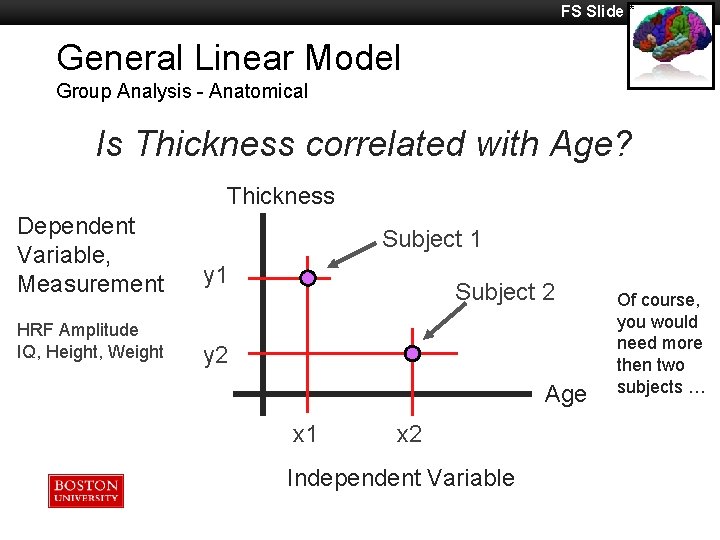
FS Slide * General Linear Model Group Analysis - Anatomical Is Thickness correlated with Age? Thickness Dependent Variable, Measurement y 1 HRF Amplitude IQ, Height, Weight y 2 Subject 1 Subject 2 Age x 1 x 2 Independent Variable Of course, you would need more then two subjects …

FS Slide * General Linear Model Group Analysis - Anatomical Intercept: b (=Offset) Slope: m y 1 System of Linear Equations y 1 = 1*b + x 1*m y 2 = 1*b + x 2*m Matrix Formulation y 2 Age x 1 x 2 } } X = Design Matrix b = Regression Coefficients = Parameter estimates = “betas” = beta. mgh (mri_glmfit output) b 1 x 1 y 1 = * m 1 x 2 y 2 -One row per subject -x values are dependent variable (age) -Column of 1’s is the ‘offset’ term (to multiply by b) Y = X*b b= b m Two parameters Thickness

Information Services & Technology 11/26/2020 recon-all -autorecon-pial -subjid pial_edits_before General Linear Model Group Analysis - Anatomical Do groups differ in Intercept? Does b 1=b 2? Does b 1 -b 2 = 0? C = [+1 -1 0 0], g = C*b Do groups differ in Slope? Does m 1=m 2? Does m 1 -m 2=0? C = [0 0 +1 -1], g = C*b Is average slope different than 0? Does (m 1+m 2)/2 = 0? C = [0 0 0. 5], g = C*b Y = X*b+n b= Thickness b 1 b 2 m 1 m 2 Intercept: b 1 Slope: m 2 Age 38 Intercept: b 2

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 General Linear Model Group Analysis - Anatomical § mri_glmfit § Performs general linear model (GLM) analysis in the volume/surface § Options include simulation for correction for multiple comparisons, weighted LMS, variance smoothing, PCA/SVD analysis of residuals, per-voxel design matrices, and 'self' regressors § Performs both the estimation and inference [tuta 0@scc-v 01 glm]$ cp -r. . /glm ~/ && cd ~/glm [tuta 0@scc-v 01 glm]$ mri_glmfit --y lh. gender_age. thickness. 10. mgh --fsgd gender_age. fsgd dods --C lh-Avg-thickness-age-Cor. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir 39

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 General Linear Model Group Analysis - Anatomical § mri_glmfit § Output: lh. gender_age. glmdir § beta. mgh -- all parameter estimates (surface overlay) § dof. dat -- degrees of freedom (text) § fwhm. dat -- average FWHM of residual (text) § lh-Avg-thickness-age-Cor -- contrast subdirectory § mask. mgh -- binary mask (surface overlay) § mri_glmfit. log -- log file (text, send this with bug reports) § rstd. mgh -- residual standard deviation (surface overlay) § rvar. mgh -- residual variance (surface overlay) § sar 1. mgh -- residual spatial AR 1 (surface overlay) § surface -- the subject and hemisphere used for this analysis (text) § Xg. dat -- design matrix (text) § X. mat -- design matrix (MATLAB format) § y. fsgd -- copy of input FSGD file (text) 40

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 General Linear Model Group Analysis - Anatomical § mri_glmfit § Output: lh. gender_age. glmdir/lh-Avg-thickness-age-Cor § C. dat -- original contrast matrix (text) § cnr. mgh -- contrast-to-noise ratio (surface overlay) § efficiency. dat -- statistical efficiency for the contrast (text) § F. mgh -- F ratio of contrast (surface overlay) § gamma. mgh -- contrast effect size (surface overlay) § gammavar. mgh --contrast variance (surface overlay) § maxvox. dat -- voxel with the maximum statistic (text) § pcc. mgh -- partial (pearson) correlation coefficient (surface overlay) § sig. mgh -- significance, -log 10(pvalue), uncorrected (surface overlay) § z. mgh -- z-stat that corresponds to the significance (surface overlay) 41

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 General Linear Model Group Analysis - Anatomical § mri_glmfit [tuta 0@scc-v 01 glm]$ vglrun freeview -f $SUBJECTS_DIR/fsaverage/surf/lh. inflated: annot=aparc. annot : annot_outline=1: overlay=lh. gender_age. glmdir/lh-Avgthickness-age-Cor/sig. mgh: overlay_threshold=4, 5 -viewport 3 d -layout 1 & [tuta 0@scc-v 01 glm]$ vglrun freeview -f $SUBJECTS_DIR/fsaverage/surf/lh. inflated: annot=aparc. annot: an not_outline=1: overlay=lh. gender_age. glmdir/lh-Avg-thickness-age -Cor/F. mgh: overlay_threshold=20, 50 -viewport 3 d -layout 1 & 42
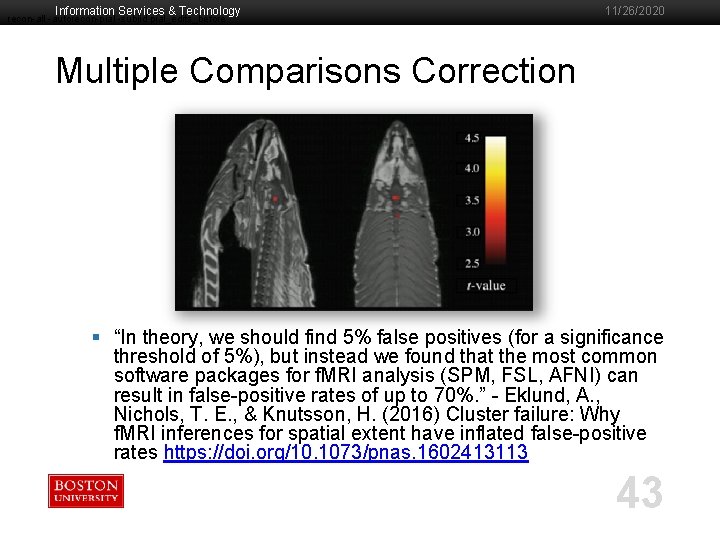
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multiple Comparisons Correction § “In theory, we should find 5% false positives (for a significance threshold of 5%), but instead we found that the most common software packages for f. MRI analysis (SPM, FSL, AFNI) can result in false-positive rates of up to 70%. ” - Eklund, A. , Nichols, T. E. , & Knutsson, H. (2016) Cluster failure: Why f. MRI inferences for spatial extent have inflated false-positive rates https: //doi. org/10. 1073/pnas. 1602413113 43
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multiple Comparisons Correction § mri_glmfit-sim [tuta 0@scc-v 01 glm]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/ tutorial_subjs/group_analysis_tutorial_perm [tuta 0@scc-v 01 glm]$ mri_glmfit-sim --glmdir lh. gender_age. glmdir --perm 1000 4. 0 abs --cwp 0. 05 --2 spaces --bg 1 44
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multiple Comparisons Correction § Cluster Summary [tuta 0@scc-v 01 glm]$ cp -r $SUBJECTS_DIR/glm ~/glm_perm && cd ~/glm_perm [tuta 0@scc-v 01 glm_perm]$ less lh. gender_age. glmdir/lh-Avg-thickness-age. Cor/perm. th 40. abs. sig. cluster. summary [tuta 0@scc-v 01 glm_perm]$ vglrun freeview -f $SUBJECTS_DIR/fsaverage/surf/lh. inflated: overlay=lh. gender_age. glmdir/lh-Avg -thickness-age. Cor/perm. th 40. abs. sig. cluster. mgh: overlay_threshold=2, 5: annot=lh. gender_age. glmdir/lh-Avg-thickness-age-Cor/perm. th 40. abs. sig. ocn. annot -viewport 3 d layout 1 & 45

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multiple Comparisons Correction § mri_glmfit-sim § Output: lh. gender_age. glmdir/lh-Avg-thickness-age-Cor § perm. th 40. abs. pdf. dat -- probability distribution function of clusterwise correction § perm. th 40. abs. sig. cluster. mgh -- cluster-wise corrected map (overlay) § perm. th 40. abs. sig. cluster. summary -- summary of clusters (text) § perm. th 40. abs. sig. masked. mgh -- uncorrected sig values masked by the clusters that survive correction § perm. th 40. abs. sig. ocn. annot -- output cluster number (annotation of clusters) § perm. th 40. abs. sig. ocn. mgh -- output cluster number (segmentation showing where each numbered cluster is) § perm. th 40. abs. sig. voxel. max. dat -- maximum voxel-wise significance § perm. th 40. abs. sig. voxel. mgh -- voxel-wise map corrected for multiple comparisons at a voxel (rather than cluster) level § perm. th 40. abs. sig. y. ocn. dat -- the average value of each subject in each cluster 46

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multiple Comparisons Correction § mri_glmfit [tuta 0@scc-v 01 glm]$ less lh. gender_age. glmdir/lh-Avgthickness-age-Cor/perm. th 40. abs. sig. cluster. summary 47
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 ROI Analysis - Anatomical § Examining segmentation, parcellation, and color lookup table [tuta 0@scc-v 01 glm]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs/group_an alysis_tutorial [tuta 0@scc-v 01 glm]$ cp -r $SUBJECTS_DIR/004 ~/004 && cd ~ [tuta 0@scc-v 01 ~]$ vglrun freeview -v 004/mri/orig. mgz 004/mri/aparc+aseg. mgz: colormap=lut: opacity=0. 4 -f 004/surf/lh. white: annot=aparc. annot & [tuta 0@scc-v 01 ~]$ less $FREESURFER_HOME/Free. Surfer. Color. LUT. txt 48
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 ROI Analysis - Anatomical § Labeling § Draw label on fsaverage and map to subject with mri_label 2 label [tuta 0@scc-v 01 ~]$ mri_label 2 label --srcsubject fsaverage --srclabel $SUBJECTS_DIR/fsaverage/label/lh. BA 45_exvivo. label --trgsubject 004 --trglabel 004/label/lh. BA 45_exvivo. label --hemi lh --regmethod surface § Load Label on Volume [tuta 0@scc-v 01 ~]$ freeview -v 004/mri/orig. mgz v File > Load ROI > lh. BA 45_exvivo. label § Load Label on Surface § Highlight surface v Label > Load > lh. BA 45_exvivo. label 49

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 ROI Analysis - Anatomical § aseg. stats & aparc. stats § Table generated by recon-all containing volume and thickness statistics for labeled regions for each subject [tuta 0@scc-v 01 ~]$ cd $SUBJECTS_DIR/004/stats [tuta 0@scc-v 01 stats]$ less aseg. stats [tuta 0@scc-v 01 stats]$ less lh. aparc. stats 50
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 ROI Analysis - Anatomical § Group Stats § Create a file of aseg. stats across subjects [tuta 0@scc-v 01 ~]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs/g roup_analysis_tutorial && cd ~ [tuta 0@scc-v 01 stats]$ asegstats 2 table --subjects 004 021 040 067 080 092 --segno 11 17 18 --tablefile aseg. vol. table § --segno selects Table IDs from Free. Surfer. Color. LUT. txt 51
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 ROI Analysis - Anatomical § Select desired measurements from aseg. stats [tuta 0@scc-v 01 ~]$ asegstats 2 table --subjects 004 021 040 067 080 092 --segno 11 17 18 --meas mean --tablefile aseg. mean-intensity. table § Select which segmentation atlas to retrieve stats from [tuta 0@scc-v 01 stats]$ asegstats 2 table --subjects 004 021 040 067 080 092 --segno 3007 3021 3022 4022 --stats wmparc. stats --tablefile wmparc. vol. table 52

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 ROI Analysis - Anatomical § Create a file of aparc. stats across subjects [tuta 0@scc-v 01 ~]$ aparcstats 2 table --hemi lh --subjects 004 021 040 067 080 092 --tablefile lh. aparc. area. table § Select which summary measure and parcellation atlas to retrieve stats from [tuta 0@scc-v 01 stats]$ aparcstats 2 table --hemi lh --subjects 004 021 040 067 080 092 --meas thickness --parc aparc. a 2009 s --tablefile lh. aparc. a 2009. thickness. table 53

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multimodal Registration § Multimodal integration - registering T 1 with functional or diffusion scans § Data from Functional Biomedical Research Network (f. BIRN) Manual Registration: [tuta 0@scc-v 01 ~]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs [tuta 0@scc-v 01 ~]$ cd $SUBJECTS_DIR/multimodal/fmri/fbirn-101 [tuta 0@scc-v 01 ~]$ vglrun freeview -v template. nii $SUBJECTS_DIR/fbirn-anat-101. v 6/mri/orig. mgz: visible=0 -f $SUBJECTS_DIR/fbirn-anat-101. v 6/surf/lh. white $SUBJECTS_DIR/fbirn-anat-101. v 6/surf/rh. white -viewport cor -transform-volume § Registration matrix can be saved as a. lta file or can save volume in new space 54

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Multimodal Registration Automatic Registration: [tuta 0@scc-v 01 ~]$ bbregister --mov template. nii --bold --s fbirn-anat-101. v 6 -lta ~/register. lta [tuta 0@scc-v 01 ~]$ cat register. dat. mincost [tuta 0@scc-v 01 ~]$ vglrun freeview -v $SUBJECTS_DIR/fbirn-anat-101. v 6/mri/orig. mgz: visible=0 template. nii: reg=register. lta -f $SUBJECTS_DIR/fbirn-anat-101. v 6/surf/lh. white $SUBJECTS_DIR/fbirn-anat-101. v 6/surf/rh. white -viewport cor -transform-volume -or[tuta 0@scc-v 01 ~]$ vglrun tkregisterfv --mov template. nii --reg /usr 4/tutorial/tuta 0/register. lta --surfs 55

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 FS-FAST f. MRI Analysis § Explain § Functional Data Directory vs Anatomical Data Directory vs SUBJECTS_DIR 56

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 FS-FAST f. MRI Analysis Tutorial Data: Understanding, Downloading, Setting up § Tutorial Data Description: § Working-memory paradigm with distractors § f. BIRN, n=18, 1 run/subject, 3 T § Functional Paradigm § Block design § 3 phases: Encode (16 s), Distractor (16 s Emotional/Neutral), Probe (16 sec) § 5 conditions (scrambled is modelled as baseline rather than condition) 57

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 FS-FAST f. MRI Analysis Tutorial Data: Understanding, Downloading, Setting up § Functional Data § § § Original data was 8 runs per subject This data is only 1 run per subject 142 time points TR = 2 seconds One run for rest data One B 0 map for each subject [tuta 0@scc-v 01 ~]$ mri_info $TUTORIAL_DATA/fsfastfunctional/sess 01. noproc/bold/001/f. nii. gz § Anatomical Data § Free. Surfer analysis has been run for all 18 subjects (reconall) 58
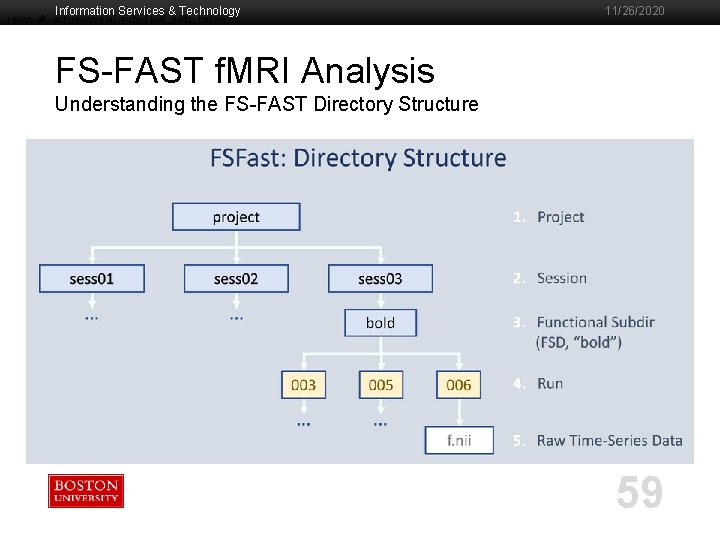
Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 FS-FAST f. MRI Analysis Understanding the FS-FAST Directory Structure 59

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 FS-FAST f. MRI Analysis Understanding the FS-FAST Directory Structure § Project Directory (fsfast-functional) [tuta 0@scc-v 01 ~]$ cd $TUTORIAL_DATA/fsfast-functional [tuta 0@scc-v 01 fsfast-functional]$ ls § Session Directory § sess 01. noproc - what the directory should look like prior to beginning analysis § Directory Structure, Raw Data in functional subdirectories (FSD), ‘subjectname’ file, paradigm files for each run [tuta 0@scc-v 01 fsfast-functional]$ cd $TUTORIAL_DATA/fsfast-functional/sess 01. noproc [tuta 0@scc-v 01 sess 01. noproc]$ ls § SUBJECTS_DIR & ‘subjectname' § SUBJECTS_DIR must point to Anatomical Data § ‘subjectname’ file must include name of session in SUBJECTS_DIR [tuta 0@scc-v 01 sess 01. noproc]$ export SUBJECTS_DIR=$TUTORIAL_DATA/fsfast-tutorial. subjects [tuta 0@scc-v 01 sess 01. noproc]$ cat subjectname [tuta 0@scc-v 01 sess 01. noproc]$ ls $SUBJECTS_DIR 60

Information Services & Technology 11/26/2020 recon-all -autorecon-pial -subjid pial_edits_before FS-FAST f. MRI Analysis Understanding the FS-FAST Directory Structure § Functional Subdirectories (FSD) [tuta 0@scc-v 01 [tuta 0@scc-v 01 sess 01. noproc]$ ls rest ls bold mri_info --dim bold/001/f. nii. gz mri_info --res bold/001/f. nii. gz vglrun freeview -v bold/001/f. nii. gz -timecourse § Paradigm Files § Text files with the stimulus schedule [tuta 0@scc-v 01 sess 01. noproc]$ cat bold/001/workmen. par Seconds Condition (#) 22. 000 1 38. 000 2 Duration 16. 000 Weight 1. 0000 Condition (name) Encode Edistractor § Subject list § ‘sessidlist’ 61

Information Services & Technology 11/26/2020 recon-all -autorecon-pial -subjid pial_edits_before FS-FAST f. MRI Analysis Understanding the FS-FAST Directory Structure § Functional Subdirectories (FSD) [tuta 0@scc-v 01 [tuta 0@scc-v 01 sess 01. noproc]$ ls rest ls bold mri_info --dim bold/001/f. nii. gz mri_info --res bold/001/f. nii. gz vglrun freeview -v bold/001/f. nii. gz -timecourse § Paradigm Files § Text files with the stimulus schedule [tuta 0@scc-v 01 sess 01. noproc]$ cat bold/001/workmen. par Seconds Condition (#) 22. 000 1 38. 000 2 Duration 16. 000 Weight 1. 0000 Condition (name) Encode Edistractor § Subject list § ‘sessidlist’ 62

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Individual f. MRI Integration § Check Registration [tuta 0@scc-v 01 ~]$ export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs [tuta 0@scc-v 01 ~]$ cd $SUBJECTS_DIR/multimodal/fmri/fbirn-101 [tuta 0@scc-v 01 ~]$ vglrun freeview -v $SUBJECTS_DIR/fbirn-anat-101. v 6/mri/orig. mgz: visible=0 template. nii: reg=register. lta -f $SUBJECTS_DIR/fbirn-anat-101. v 6/surf/lh. white $SUBJECTS_DIR/fbirn-anat-101. v 6/surf/rh. white -viewport cor -transform-volume [tuta 0@scc-v 01 ~]$ cd $SUBJECTS_DIR/multimodal/fmri/fbirn-101 63

Information Services & Technology recon-all -autorecon-pial -subjid pial_edits_before 11/26/2020 Individual f. MRI Integration § View Significance Map [tuta 0@scc-v 01 ~]$ vglrun freeview -v $SUBJECTS_DIR/fbirn-anat-101. v 6/mri/orig. mgz $SUBJECTS_DIR/fbirn-anat 101. v 6/mri/aparc+aseg. mgz: colormap=lut: opacity=0. 2 sig. nii: reg=register. lta: colormap=heat: heatscale=2, 2, 4 -colorscale -viewport coronal 64
Information Services & Technology 11/26/2020 Questions § All SCC questions welcome. Those I can’t answer I will make every effort to get you an answer for later. § Email Addresses: § Dustin Clark– clarkdc@bu. edu § General help using the SCC – help@scc. bu. edu 65
Information Services & Technology 11/26/2020 Tutorial Survey § Please open a web browser and go to: § http: //scv. bu. edu/survey/tutorial_evaluation. html § to fill out the tutorial survey. Thanks for coming. § Tutorials slides available on the web from: http: //www. bu. edu/tech/support/research/trainingconsulting/live-tutorials/ § My Contact Information: aarondf@bu. edu or (617) 353 -8255 66

Information Services & Technology 11/26/2020 Additional Free. Surfer Web Resources § FS Voxel Edit § MRI Coordinate Systems 67

Information Services & Technology 11/26/2020 Additional BU RCS Web Resources § Research Computing Support Pages http: //www. bu. edu/tech/support/research/ § Technical Summary of SCC Resources http: //www. bu. edu/tech/support/research/computingresources/tech-summary/ § SCC Updates – Latest SCC News http: //www. bu. edu/tech/support/research/whatshappening/updates/ § Code Examples for Popular Software Packages http: //scv. bu. edu/examples/ 68
Information Services & Technology 11/26/2020 Image Sources § https: //surfer. nmr. mgh. harvard. edu/ § https: //tecnologia. umcomo. com. br/artigo/como-fazerprint-screen-no-mac-15383. html § https: //www. health. harvard. edu/pain/headache-whento-worry-what-to-do § http: //i. liketightpants. net/and/absolute-beginners-unixfor-art-students-part-1 § https: //www. xnat. org/ § https: //mytherapynyc. com/how-to-relieve-stress-bydeep-breathing/ 69
- Slides: 69