Introduction to CLC Main Workbench 20 June 2012

Introduction to CLC Main Workbench 20 June, 2012 Ansuman Chattopadhyay, Ph. D Head, Molecular Biology Information Services Health Sciences Library System University of Pittsburgh ansuman@pitt. edu

Sequence Analysis Software Suits n n Wisconsin GCG Vector. NTI DNA STAR-Laser. Geneious n CLC Main

Why CLC Main ? n n n n Windows Mac Linux DNA, RNA, Protein, Microarray Data Analysis Regular Update HSLS Licensed

CLC Main Access n HSLS CLC Main Registration q n Link: http: //www. hsls. pitt. edu/molbio/clcmain Access via Pitt - Network Connect q Instruction video: http: //goo. gl/JNj. Mt

Topics n n n n CLC Main GUI Import DNA sequence into CLC Import Protein sequence into CLC Design PCR primers Perform restriction enzymes digestions Run in silico agarose gels Protein primary structure analysis Protease digestions

n CLC Main Graphical User Interface (GUI)
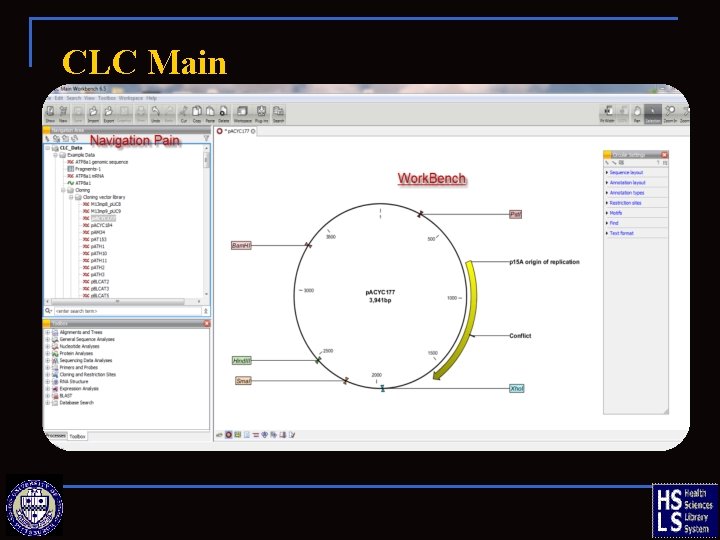
CLC Main

Basic Navigation -DNA -Protein

Import a DNA Sequence

DNA Sequence n Human PLCg 1 q q q Refseq no: NM_002660 FASTA file Raw sequence CLC features: Search, Import, Create new sequence

CLC DNA sequence

Import a Protein Sequence

Protein Sequence n Human PLCg 1 q q Refseq no: NP_002651 Uniprot Accession Number: P 19174 FASTA file Raw sequence CLC features: Search, Import, Create new sequence

CLC protein sequence
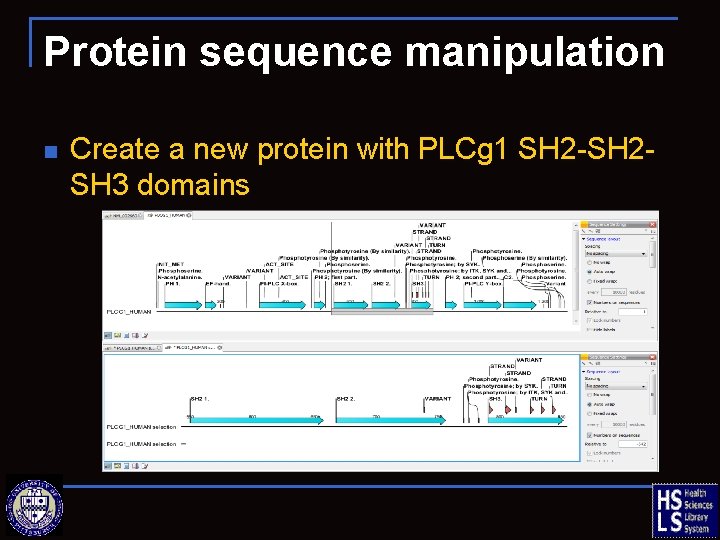
Protein sequence manipulation n Create a new protein with PLCg 1 SH 2 -SH 2 SH 3 domains
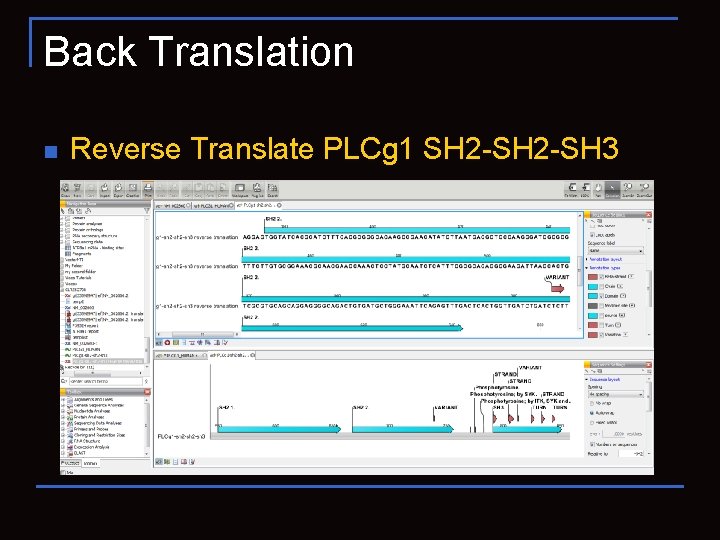
Back Translation n Reverse Translate PLCg 1 SH 2 -SH 3

Perform Restriction Digestion

Restriction Mapping www. biologyreference. com http: //www. hsls. pitt. edu/molbio

Restriction Digestion

Protein Primary Structure Analysis
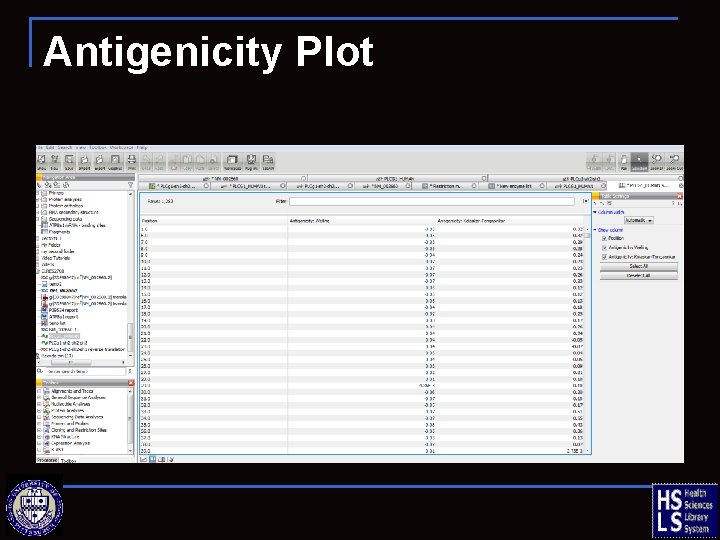
Antigenicity Plot
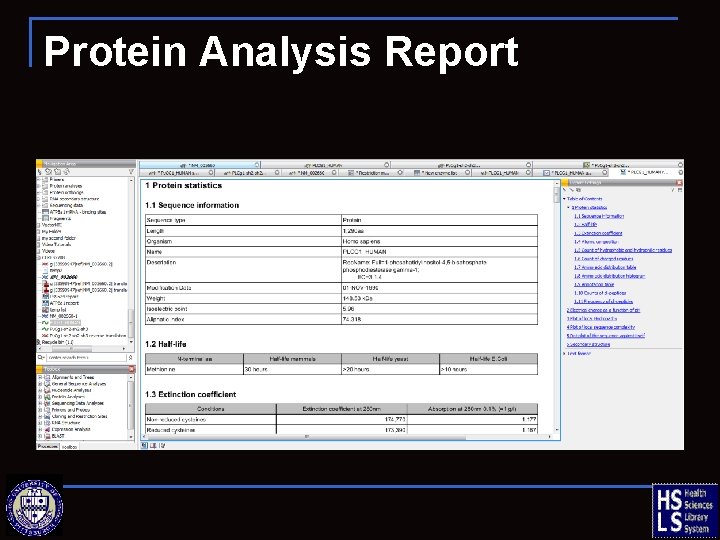
Protein Analysis Report

Protease Digestion

Proteolytic Cleavage

Primer Design

Primer Analysis & Design A little something to get you in the mood… http: //www. hsls. pitt. edu/molbio

Polymerase Chain Reaction (PCR) 1983 -Kary Mullis very simple n n n exponential amplification similar to natural DNA replication The primary reagents, used in PCR are: n n Template DNA–DNA sequence to amplify DNA nucleotides–building blocks for new DNA Taq polymerase–heat stable enzyme catalyzes new DNA Primers–single-stranded DNA, ~20 -50 nucleotides, complimentary to a short region on either side of template DNA http: //www. hsls. pitt. edu/molbio

Things to consider for primer design… n Primer-Dimer formation n Secondary Structures in Primers n Illegitimate Priming in Template DNA due to repeated sequences n Incompatibility with PCR conditions SOURCE: NCBI http: //www. hsls. pitt. edu/molbio
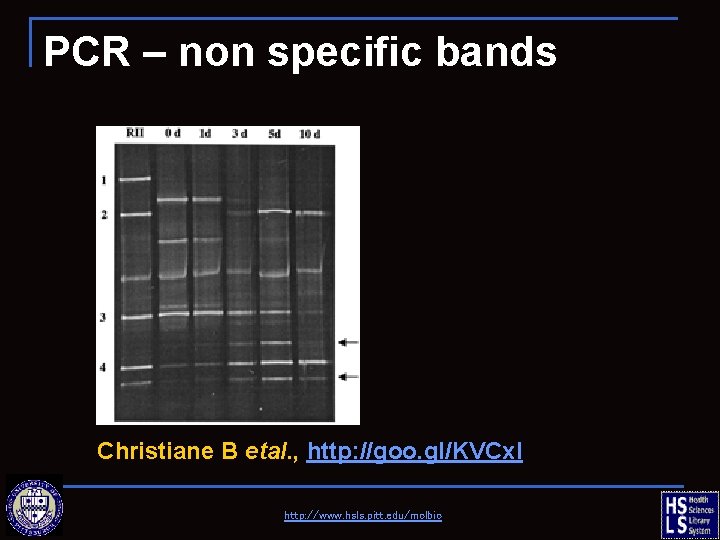
PCR – non specific bands Christiane B etal. , http: //goo. gl/KVCx. I http: //www. hsls. pitt. edu/molbio

n Design PCR Primers to amplify the region covering exons 4 -5 in human PLCg 1 m. RNA sequence http: //www. hsls. pitt. edu/molbio



Design PCR primers to amplify a DNA region covering a protein domain q q PCR amplification of human PLCg 1 SH 3 domain CLC Main Features: n n q Reverse Translate PCR Primer Design Video Tutorials http: //www. hsls. pitt. edu/molbio

In silico cloning

Molecule Construction Clone a fragment from p. BR 322 into p. UC 19 ☼ Donor fragment: p. BR 322, 5’Eco. RI— 3’Ava. I ☼ Recipient fragment: p. UC 19, 5’Sma. I— 3’Eco. RI video tutorials http: //www. hsls. pitt. edu/molbio
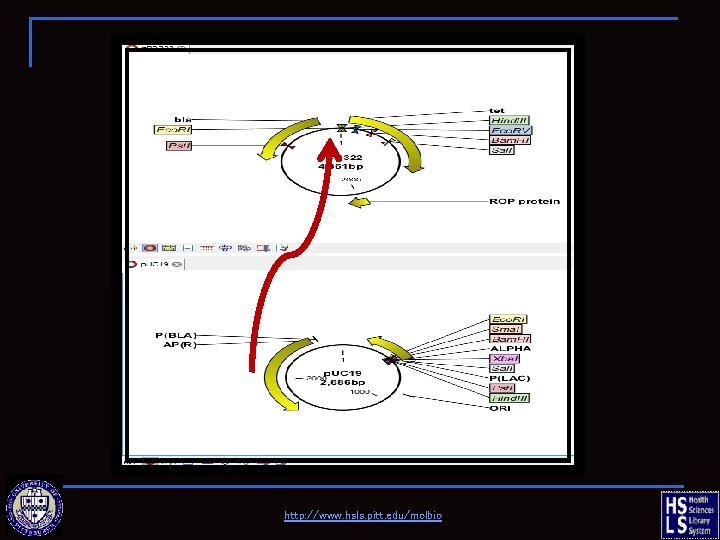
http: //www. hsls. pitt. edu/molbio
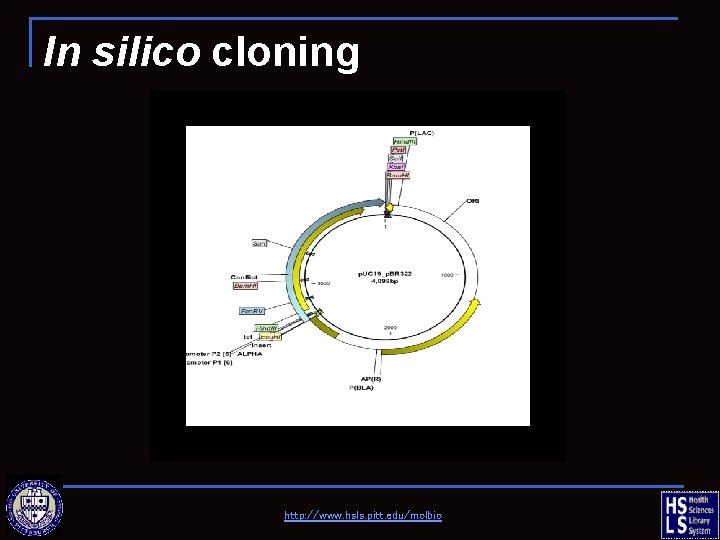
In silico cloning http: //www. hsls. pitt. edu/molbio

Sequence Alignment n Pair-wise Alignment q q n Global Local Multiple Sequence Alignment http: //www. hsls. pitt. edu/molbio

Sequence Alignment http: //www. hsls. pitt. edu/molbio
Pair-wise Sequence Alignment http: //www. hsls. pitt. edu/molbio

Multiple Sequence Alignment http: //www. hsls. pitt. edu/molbio

PLCg 1 Orthologous sequences n PLCg 1: q Mouse: Rat: Cow: Dog: Zebra fish: NP_067255 NP_037319 NP_776850 XP_542998 NP_919388 q Human: NP_002651 q NP_067255, NP_037319, NP_776850, XP_542998, NP_919388, NP_002651 q q http: //www. hsls. pitt. edu/molbio

Thank you! Any questions? Carrie Iwema iwema@pitt. edu 412 -383 -6887 Ansuman Chattopadhyay ansuman@pitt. edu 412 -648 -1297 http: //www. hsls. pitt. edu/molbio
- Slides: 44