Introduction to Bioinformatics Similarity searches in sequence databases

Introduction to Bioinformatics Similarity searches in sequence databases

Rappel : alignement de séquences avec gap 2
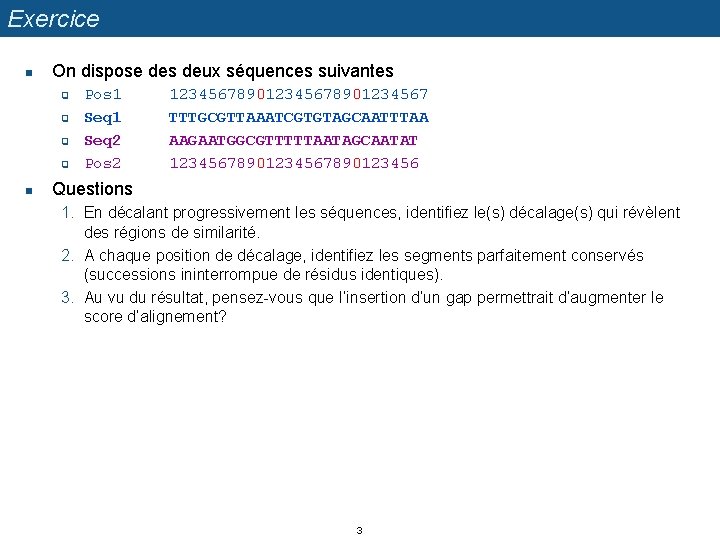
Exercice n On dispose des deux séquences suivantes q q n Pos 1 Seq 2 Pos 2 12345678901234567 TTTGCGTTAAATCGTGTAGCAATTTAA AAGAATGGCGTTTTTAATAGCAATAT 1234567890123456 Questions 1. En décalant progressivement les séquences, identifiez le(s) décalage(s) qui révèlent des régions de similarité. 2. A chaque position de décalage, identifiez les segments parfaitement conservés (successions ininterrompue de résidus identiques). 3. Au vu du résultat, pensez-vous que l’insertion d’un gap permettrait d’augmenter le score d’alignement? 3
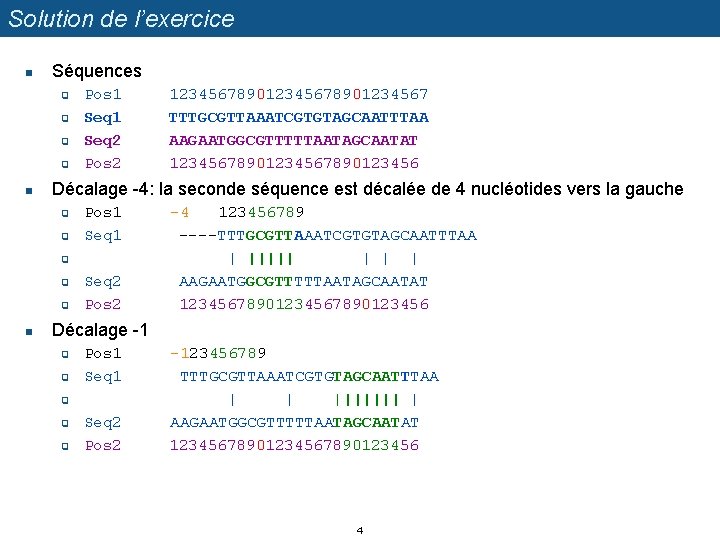
Solution de l’exercice n Séquences q q n Pos 1 Seq 2 Pos 2 Décalage -4: la seconde séquence est décalée de 4 nucléotides vers la gauche q q Pos 1 Seq 1 q q q n 12345678901234567 TTTGCGTTAAATCGTGTAGCAATTTAA AAGAATGGCGTTTTTAATAGCAATAT 1234567890123456 Seq 2 Pos 2 -4 123456789 ----TTTGCGTTAAATCGTGTAGCAATTTAA | ||||| | AAGAATGGCGTTTTTAATAGCAATAT 1234567890123456 Décalage -1 q q Pos 1 Seq 1 q q q Seq 2 Pos 2 -123456789 TTTGCGTTAAATCGTGTAGCAATTTAA | | ||||||| | AAGAATGGCGTTTTTAATAGCAATAT 1234567890123456 4

Alignement avec « gaps » (brèches) n n n Les alignements sans gaps sont rarement pertinents, car les divergences entre séquences incluent souvent des insertions et délétions. Les gaps permettent de mettre en évidence les régions de similarités multiples. ----TTTGCGTT--AAATCGTGTAGCAATTTAA s=substitution; |=identité 1111 s|s|||||11 s||22222|||||||s|22 1=gap dans la 1ère séquence AAGAATGGCGTTTTTAA-----TAGCAATAT-2=gap dans la 2 de séquence Gaps, insertions et délétions q Les “gaps” (brèches) reflètent soit une insertion dans l’une des séquences, soit une délétion dans l’autre. q Sur seule base de l’alignement d’une paire de séquences, on ne peut pas déterminer si un gap correspond à une délétion ou une insertion. q On utilise le terme indel pour désigner cet événement évolutif de nature indéterminée (insertion ou délétion) qui a donné lieu à un gap observé dans un alignement. 5

Bioinformatics Sequence search algorithms

Matching a sequence against a database n Example of utilization q n n n We obtained the sequence of a protein of unknown function, and we would like to search Uni. Prot for all similar proteins, in order to emit hypotheses about the possible function (function prediction by similarity). Approach: we will align our query sequence to each entry in Uni. Prot. Problem of size : in Dec 2005, there were 2. 500. 000 entries in Uni. Prot (Swiss. Prot + TREMBL) It is possible to apply dynamical programming, but it takes a lot of time or requires high computation power. 7

Fast algorithms for database matching n n n Fast. A BLAST (Basic Local Alignment Search Tool) In short q q q These algorithms are ~50 times faster than Smith-Waterman They cannot guarantee the optimal solution However, a comparison with results obtained by dynamical programming has shown that Fast. A answer is close to the optimum 8

Fast. A strategy n n Fast. A builds an index with the positions of all the small words (k-tuples) found in the query sequence. The program then detects diagonals of k-tuples between the query and the database sequences. When a significant diagonal is detected, the two sequences are aligned with Smith-Waterman algorithm. The size of words (k) influences the behaviour of the program: when k increases, the search is faster but one might miss some matches. 1 2 3 pos 1234567890123456789012 seq MVDFYYLPGSSMVDVFDFYAKAVGVELNKKLL 3 -tuples MVD VDF DFY FYY YYL. . . position index k-tuple MVD VDF DFY. . . positions 1, 12 2 3, 17. . . 9

Principle of the Fast. A strategy n Left q n Center q q q n Comparison of k-mer positions between query and database sequences. Highest-density regions ("init regions") are identified. The best one is highlighted with a star. Regions with a score below a given threshold appear in dotted lines, for illustation purpose. Right q Low-scoring regions are filtered out, and the remaining regions are joined. Source: Mount (2000) 10

BLAST strategies (Altschul et al. , 1990; 1997) n n n Version 1 (1990): gapless BLAST q Prior indexing of all k-mers (words) in the database (formatdb). q When search submitted, builds a dictionary of k-mers found in the query sequence. q Uses a substitution matrix (e. g. BLOSUM) to calculate a score between these words and each possible word of the same length. q Only retains word pairs with sufficient score (threshold on word pair scores). q Each time a word pair from the dictionary passes the threshold (hit), extends it in both directions (without gaps), to obtain a High-scoring Segment Pair (HSP). q The program returns sequences with significant high-scoring segment pairs. Valeurs par défaut des tailles de mots q Pour les séquences d’acides nucléiques k=11 (cf illustration) q Pour les séquences peptidiques, k=3 Exercice n Calculer la probabilité de trouver un HSP de taille 11 à une position donnée, dans une séquences nucléique composée de nucléotides équiprobables. GGTAGCAAATGTCCTGTACTGTACATGGTCAAACTGGTGAAT ||||||||||| TGTATCAAATGTCCTGTGTGAATGGTAGATGGTCAAACTGGTCAAT 11

BLAST strategy (gapless version, Altschul et al. , 1990) n n n Version 1 (1990) q Builds a dictionary of k-tuples (small words) found in the query sequence. q Uses a substitution matrix (e. g. BLOSUM) to calculate a score between these words and each possible word of the same length. q Only retains the words with a high score. q Each time a pair of words from the dictionary are found (hits) in the database sequence, extends the hit in both direction (without gaps), to obtain a High-scoring Segment Pair (HSP). q The program returns sequences with significant high-scoring segment pairs. High scoring pairs (HSP) q For nucleic acid sequences, default k=11 (illustration) q For peptidic sequences, default k=3 Exercise: compute the probability to find a 11 bp HSP starting at a given position, in a nucleic acid sequence with equiprobable nucleotides. GGTAGCAAATGTCCTGTACTGTACATGGTCAAACTGGTGAAT ||||||||||| TGTATCAAATGTCCTGTGTGAATGGTAGATGGTCAAACTGGTCAAT 12

BLAST - Elongation de l’alignement Score d’identité = 1 Score de substitution = -1 The quick brown fox jumps over the lazy dog The quiet brown cat purrs when she sees him 13

BLAST - Elongation de l’alignement Score d’identité = 1 Score de substitution = -1 The quick brown fox jump The quiet brown cat purr 123 45654 56789 876 5654 <= SCORE 14

BLAST – Elongation of the alignment Identity score = 1 Substitution score = -1 The 123 000 quick quiet 45654 00012 brown 56789 10000 fox cat 876 123 jump purr 5654 4345 <= SCORE <=(SCORE(max)-SCORE) Elongation stops if (SCORE(max)-SCORE) > prefedined threshold 15

BLAST - Elongation de l’alignement HSP (High Scoring Pair): The quick brown ||||||| The quiet brown The 123 000 quick quiet 45654 00012 brown 56789 10000 fox cat 876 123 jump purr 5654 4345 <= SCORE <=(SCORE(max)-SCORE) Ecourter l’alignement jusqu’au dernier Score(max) 16

BLAST - Elongation de l’alignement n Elongation de l’alignement de deux côtés à partir des mots du dictionnaire q q L’élongation s’arrête si le score diminue en-deçà d’une limite prédéfinie par rapport au dernier maximum. L’alignement est écourté jusqu’au dernier score maximal • (S) => HSP (High Scoring Pair) 17
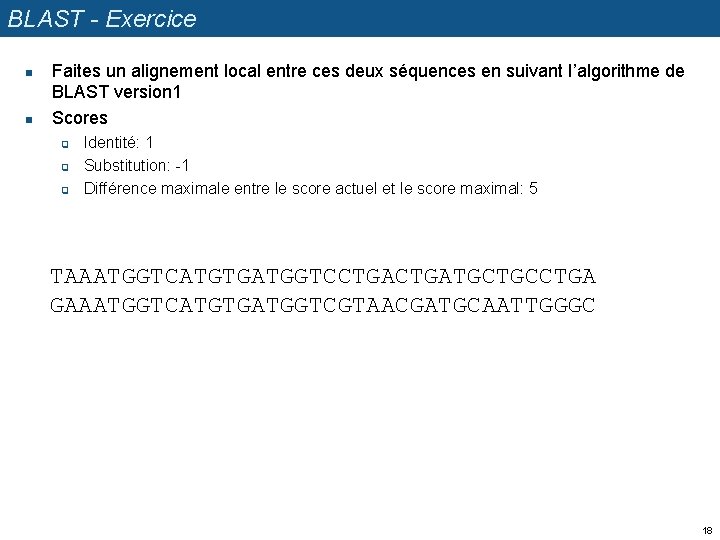
BLAST - Exercice n n Faites un alignement local entre ces deux séquences en suivant l’algorithme de BLAST version 1 Scores q q q Identité: 1 Substitution: -1 Différence maximale entre le score actuel et le score maximal: 5 TAAATGGTCATGTGATGGTCCTGATGCTGCCTGA GAAATGGTCATGTGATGGTCGTAACGATGCAATTGGGC 18
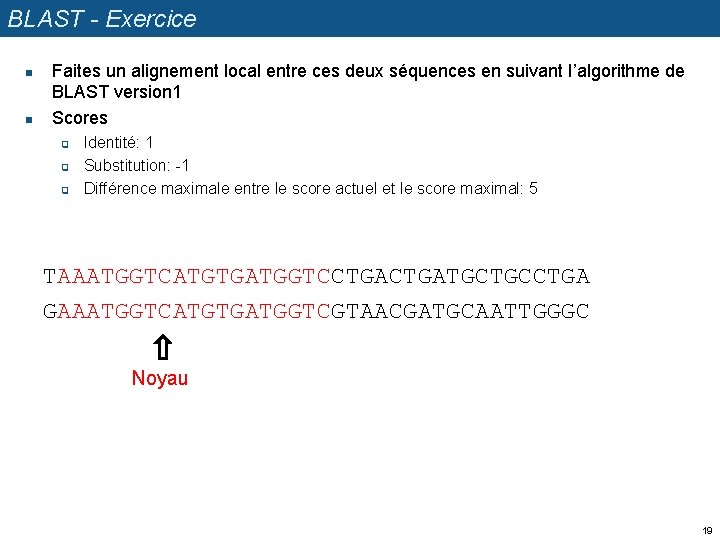
BLAST - Exercice n n Faites un alignement local entre ces deux séquences en suivant l’algorithme de BLAST version 1 Scores q q q Identité: 1 Substitution: -1 Différence maximale entre le score actuel et le score maximal: 5 TAAATGGTCATGTGATGGTCCTGATGCTGCCTGA GAAATGGTCATGTGATGGTCGTAACGATGCAATTGGGC Noyau 19

BLAST - Exercice n n Faites un alignement local entre ces deux séquences en suivant l’algorithme de BLAST version 1 Scores q q q Identité: 1 Substitution: -1 Différence maximale entre le score actuel et le score maximal: 5 TAAATGGTCATGTGATGGTCCTGATGCTGCCTGA Seq 1 GAAATGGTCATGTGATGGTCGTAACGATGCAATTGGGC Seq 2 1234567891111111211111 Score 01234567898989098765 20
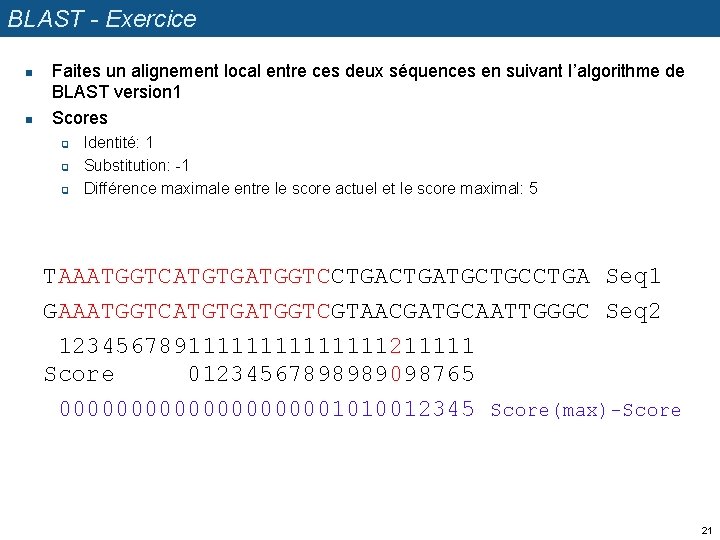
BLAST - Exercice n n Faites un alignement local entre ces deux séquences en suivant l’algorithme de BLAST version 1 Scores q q q Identité: 1 Substitution: -1 Différence maximale entre le score actuel et le score maximal: 5 TAAATGGTCATGTGATGGTCCTGATGCTGCCTGA Seq 1 GAAATGGTCATGTGATGGTCGTAACGATGCAATTGGGC Seq 2 1234567891111111211111 Score 01234567898989098765 00000000001010012345 Score(max)-Score 21
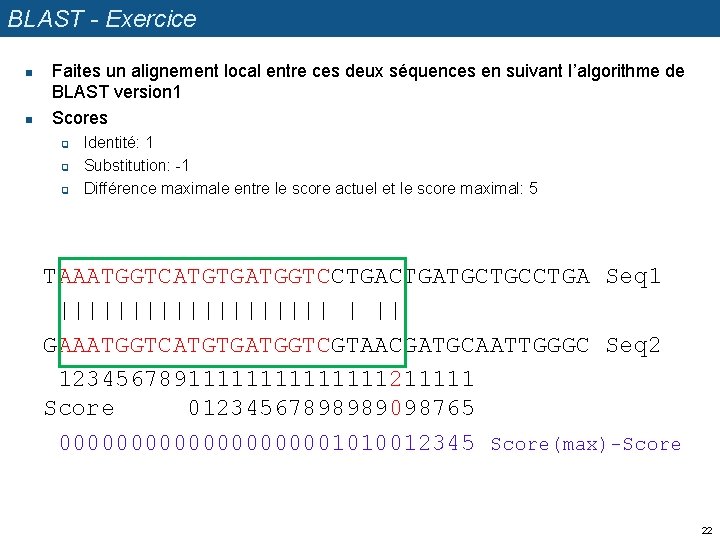
BLAST - Exercice n n Faites un alignement local entre ces deux séquences en suivant l’algorithme de BLAST version 1 Scores q q q Identité: 1 Substitution: -1 Différence maximale entre le score actuel et le score maximal: 5 TAAATGGTCATGTGATGGTCCTGATGCTGCCTGA Seq 1 |||||||||| | || GAAATGGTCATGTGATGGTCGTAACGATGCAATTGGGC Seq 2 1234567891111111211111 Score 01234567898989098765 00000000001010012345 Score(max)-Score 22

BLAST strategies (Altschul et al. , 1990; 1997) n n n Version 1 (1990): gapless BLAST q Prior indexing of all k-mers (words) in the database (formatdb). q When search submitted, builds a dictionary of k-mers found in the query sequence. q Uses a substitution matrix (e. g. BLOSUM) to calculate a score between these words and each possible word of the same length. q Only retains word pairs with sufficient score (threshold on word pair scores). q Each time a word pair from the dictionary passes the threshold (hit), extends it in both directions (without gaps), to obtain a High-scoring Segment Pair (HSP). q The program returns sequences with significant high-scoring segment pairs. Version 2 (1997): gapped BLAST q Use smaller words, but only extend when there are two hits on the same diagonal. q Extension includes gaps (dynamical programming). q The extension costs more time, but the number of times it is done is reduced, because the extension requires a pair of hits. Exercise n Compute the probability to find a hit pair starting at a given position, with a spacing comprised between 0 and 30, in a nucleic acid sequence with equiprobable nucleotides. 23

BLAST strategies (Altschul et al. , 1990; 1997) n n n Version 1 (1990): gapless BLAST q Prior indexing of all k-mers (words) in the database (formatdb). q When search submitted, builds a dictionary of k-mers found in the query sequence. q Uses a substitution matrix (e. g. BLOSUM) to calculate a score between these words and each possible word of the same length. q Only retains word pairs with sufficient score (threshold on word pair scores). q Each time a word pair from the dictionary passes the threshold (hit), extends it in both directions (without gaps), to obtain a High-scoring Segment Pair (HSP). q The program returns sequences with significant high-scoring segment pairs. Version 2 (1997): gapped BLAST q Use smaller words, but only extend when there are two hits on the same diagonal. q Extension includes gaps (dynamical programming). q The extension costs more time, but the number of times it is done is reduced, because the extension requires a pair of hits. PSI-BLAST (in the 1997 article as well) q A second step after the proper BLAST process. q Once the gapped BLAST has returned a set of sequences, these sequences are aligned and used to build a profile motif. q The database is then scanned with this profile motif to collect additional similarities. q The process can be iterated several times • collect sequences > build a profile -> collect sequences -> build a profile. . . 24

Some traps for BLAST searches n Spurious domains q n Low complexity regions (repetitive sequences). q n Some domains are found in many proteins. This does not mean that these proteins have the same function. The width of the alignment should thus be analyzed to assess whether the alignment covers most of the sequence length, or only a small segment. return multiple matches with no apparent functional relationship. Cloning vectors q Some entries in the database contain a fragment of the cloning vector. This can return many apparent matches, where the matching region is restricted to the cloning vector. 25

Bioinformatics Statistics of sequence similarities

Matching statistics - raw score The raw score is computed by summing the scores (obtained from the substitution matrix) for each pair of residues (r 1, i and r 2, i) over the length of the alignment (L). n R L A S V E T D M P L T L R Q H T L T S L Q T T L K A H L G T H 27
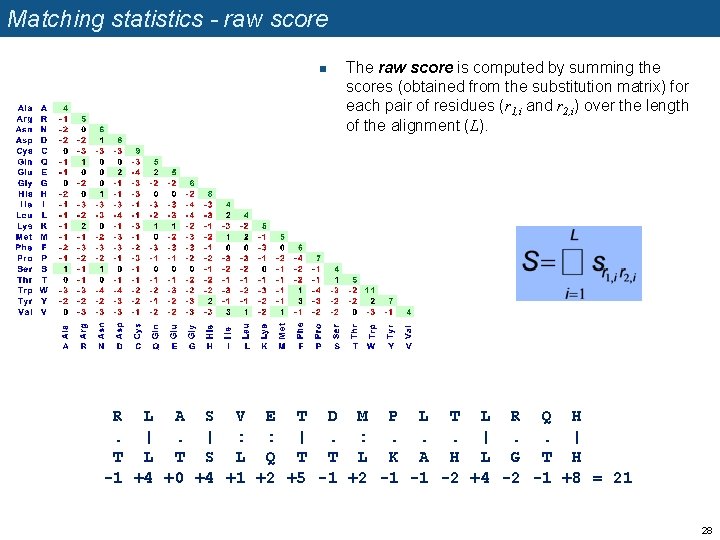
Matching statistics - raw score n The raw score is computed by summing the scores (obtained from the substitution matrix) for each pair of residues (r 1, i and r 2, i) over the length of the alignment (L). R L A S V E T D M P L T L R Q H. |. | : : |. : . . . | T L T S L Q T T L K A H L G T H -1 +4 +0 +4 +1 +2 +5 -1 +2 -1 -1 -2 +4 -2 -1 +8 = 21 28

MSP-wise P-value and bit score n The P-value of a matching segment pair (MSP) with score S is the probability to observe a score of at least S by chance. q q n Karlin and Altschul (1990) defined a way to calculate the P-value of an MSP. The P-value follows an exponential distribution, with two parameters : lambda and K. These two parameters depend on the substitution matrix chosen. They have thus to be estimated for each substitution matrix separately. The analytic way to determine the parameters lambda and L is only valid for gapless alignments. For alignment with gaps, Altschul et al (1997) propose to estimate these parameters on the basis of empirical observations Bit score of an MSP q q Karlin and Altschul (1990) also propose to convert the raw score S into a bit score S’. This facilitates the interpretation of the score, because the P-value can be directly calculated from the bit score, irrespective of the substitution matrix used for the alignment. . 29

Matching statistics- the E-value n n n n Let us imagine that we align a random word with another random word. The score is likely to be generally low. However, if this is repeated billions of times, some high scores will occasionally occur by chance. In a database scan, each word of the query sequence is compared to each word of the database. For a query sequence of size m and a database of size n, the search space (number of word pair comparisons) is thus N=nm. Fast. A and BLAST estimate, for each score, the number of matches that would be expected by chance, given the size of the database. This is called the E-value. The E-value is the product of the nominal P-value (i. e. the risk of false positives for a single comparison) by the size of the search space. For a given score S, the expected number of random matches thus increases with the size of the database. 30

Threshold on E-value n n The lower is the E-value, the more significant is the match. High E-value ( > 1) indicate that the match should not be trusted too much. One essential parameter of Fast. A and BLAST is the threshold on E-value. Very low values (e. g. : 1 e-21) q q q n Indicate that the match is very unlikely to result from chance. It is thus likely that it results from a common ancestry between the aligned sequences. In such case, we can thus admit the hypothesis of homology. Beware q q q On the BLAST Web server at NCBI, the default threshold value is 10 This means that each query would return ~10 matches by chance alone. If this default value is used, we already know that the answer is likely to contain ~10 false positive. 31

Matching statistics - database-wise P-value (=Family-Wise Error Rate) n n n From the E-value (E), one can estimate the probability to observe at least X matches by chance in random sequences. This is a simple application of the Poisson distribution : calculate the probability to observe X occurrences of an event whilst expecting E. A particular case is the probability to observe at least one match by chance q P(X>=1) This probability is generally called Family -Wise Error Rate (FWER). In the case of similarity searches, one can call it database-wise P-value. This P-value represents the probability to find at least one spurious match in the whole database search, with a score greater or equal to S. 32
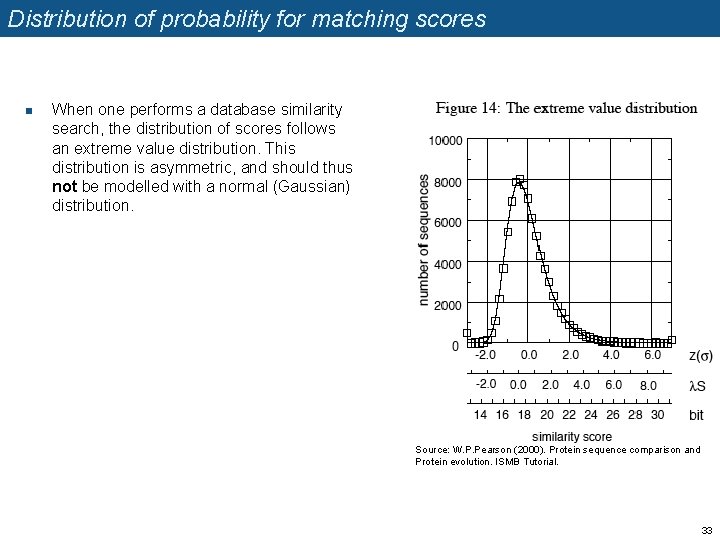
Distribution of probability for matching scores n When one performs a database similarity search, the distribution of scores follows an extreme value distribution. This distribution is asymmetric, and should thus not be modelled with a normal (Gaussian) distribution. Source: W. P. Pearson (2000). Protein sequence comparison and Protein evolution. ISMB Tutorial. 33

Interpreting similarity search results
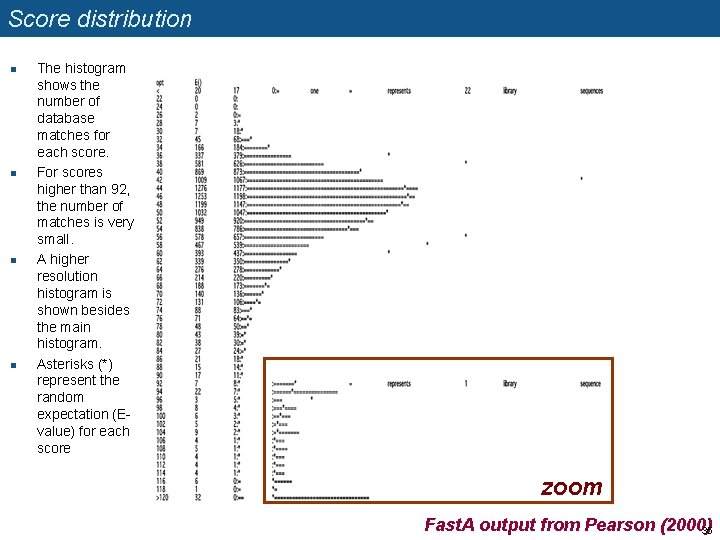
Score distribution n n The histogram shows the number of database matches for each score. For scores higher than 92, the number of matches is very small. A higher resolution histogram is shown besides the main histogram. Asterisks (*) represent the random expectation (Evalue) for each score zoom Fast. A output from Pearson (2000) 35
![BLAST result n BLASTP 2. 2. 6 [Apr-09 -2003] n Reference: Altschul, Stephen F. BLAST result n BLASTP 2. 2. 6 [Apr-09 -2003] n Reference: Altschul, Stephen F.](http://slidetodoc.com/presentation_image_h2/af013969cae7e003d9c190f400f72295/image-36.jpg)
BLAST result n BLASTP 2. 2. 6 [Apr-09 -2003] n Reference: Altschul, Stephen F. , Thomas L. Madden, Alejandro A. Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25: 3389 -3402. Query= met. L gi|16131778|ref|NP_418375. 1| aspartokinase II and homoserine dehydrogenase II; bifunctional: aspartokinase II (N-terminal); homoserine dehydrogenase II (C-terminal) [Escherichia coli K 12] (810 letters) n n Database: /Users/jvanheld/rsatools/data/genomes/Escherichia_coli_K 12/genome/NC_000913. faa 4242 sequences; 1, 351, 322 total letters The text shows the result of a BLAST search, Query: the E. coli protein Met. L, a bifunctional enzyme combining aspartokinase and homoserine dehydrogenase activities. Database: all proteins from Escherichia coli K 12. The BLAST result file starts with a summary of q q Searching. . done Score E (bits) Value Sequences producing significant alignments: gi|16131778|ref|NP_418375. 1| gi|16127996|ref|NP_414543. 1| gi|16131850|ref|NP_418448. 1| gi|16128228|ref|NP_414777. 1| aspartokinase II and homoserine deh. . . bifunctional: aspartokinase I (N-te. . . aspartokinase III, lysine sensitive. . . gamma-glutamate kinase [Escherichia. . . 1596 344 122 31 the parameters used for the search The matching sequences and the score of each match. 0. 0 2 e-95 7 e-29 0. 28 >gi|16131778|ref|NP_418375. 1| aspartokinase II and homoserine dehydrogenase II; bifunctional: aspartokinase II (N-terminal); homoserine dehydrogenase II (C-terminal) [Escherichia coli K 12] Length = 810 Score = 1596 bits (4132), Expect = 0. 0 Identities = 810/810 (100%), Positives = 810/810 (100%) 36
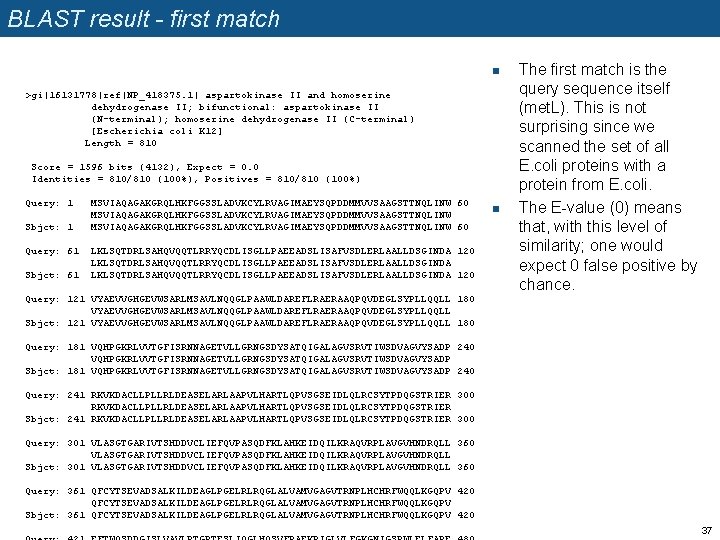
BLAST result - first match n >gi|16131778|ref|NP_418375. 1| aspartokinase II and homoserine dehydrogenase II; bifunctional: aspartokinase II (N-terminal); homoserine dehydrogenase II (C-terminal) [Escherichia coli K 12] Length = 810 Score = 1596 bits (4132), Expect = 0. 0 Identities = 810/810 (100%), Positives = 810/810 (100%) Query: 1 Sbjct: 1 Query: 61 Sbjct: 61 MSVIAQAGAKGRQLHKFGGSSLADVKCYLRVAGIMAEYSQPDDMMVVSAAGSTTNQLINW 60 LKLSQTDRLSAHQVQQTLRRYQCDLISGLLPAEEADSLISAFVSDLERLAALLDSGINDA 120 n The first match is the query sequence itself (met. L). This is not surprising since we scanned the set of all E. coli proteins with a protein from E. coli. The E-value (0) means that, with this level of similarity; one would expect 0 false positive by chance. Query: 121 VYAEVVGHGEVWSARLMSAVLNQQGLPAAWLDAREFLRAERAAQPQVDEGLSYPLLQQLL 180 VYAEVVGHGEVWSARLMSAVLNQQGLPAAWLDAREFLRAERAAQPQVDEGLSYPLLQQLL Sbjct: 121 VYAEVVGHGEVWSARLMSAVLNQQGLPAAWLDAREFLRAERAAQPQVDEGLSYPLLQQLL 180 Query: 181 VQHPGKRLVVTGFISRNNAGETVLLGRNGSDYSATQIGALAGVSRVTIWSDVAGVYSADP 240 VQHPGKRLVVTGFISRNNAGETVLLGRNGSDYSATQIGALAGVSRVTIWSDVAGVYSADP Sbjct: 181 VQHPGKRLVVTGFISRNNAGETVLLGRNGSDYSATQIGALAGVSRVTIWSDVAGVYSADP 240 Query: 241 RKVKDACLLPLLRLDEASELARLAAPVLHARTLQPVSGSEIDLQLRCSYTPDQGSTRIER 300 RKVKDACLLPLLRLDEASELARLAAPVLHARTLQPVSGSEIDLQLRCSYTPDQGSTRIER Sbjct: 241 RKVKDACLLPLLRLDEASELARLAAPVLHARTLQPVSGSEIDLQLRCSYTPDQGSTRIER 300 Query: 301 VLASGTGARIVTSHDDVCLIEFQVPASQDFKLAHKEIDQILKRAQVRPLAVGVHNDRQLL 360 VLASGTGARIVTSHDDVCLIEFQVPASQDFKLAHKEIDQILKRAQVRPLAVGVHNDRQLL Sbjct: 301 VLASGTGARIVTSHDDVCLIEFQVPASQDFKLAHKEIDQILKRAQVRPLAVGVHNDRQLL 360 Query: 361 QFCYTSEVADSALKILDEAGLPGELRLRQGLALVAMVGAGVTRNPLHCHRFWQQLKGQPV 420 QFCYTSEVADSALKILDEAGLPGELRLRQGLALVAMVGAGVTRNPLHCHRFWQQLKGQPV Sbjct: 361 QFCYTSEVADSALKILDEAGLPGELRLRQGLALVAMVGAGVTRNPLHCHRFWQQLKGQPV 420 37

BLAST result - second match n >gi|16127996|ref|NP_414543. 1| bifunctional: aspartokinase I (N-terminal); homoserine dehydrogenase I (C-terminal) [Escherichia coli K 12] Length = 820 Score = 344 bits (882), Expect = 2 e-95 Identities = 247/821 (30%), Positives = 410/821 (49%), Gaps = 44/821 (5%) Query: 16 Sbjct: 5 Query: 75 Sbjct: 65 n KFGGSSLADVKCYLRVAGIMAEYSQPDDMM-VVSAAGSTTNQLINWLKLSQTDRLSAHQV 74 KFGG+S+A+ + +LRVA I+ ++ + V+SA TN L+ ++ + + KFGGTSVANAERFLRVADILESNARQGQVATVLSAPAKITNHLVAMIEKTISGQDALPNI 64 QQTLRRYQCDLISGLLPAEEADSL--ISAFVSDLERLAALLDSGIN------DAVYAEVV 126 R + +L++GL A+ L + FV + GI+ D++ A ++ SDAERIF-AELLTGLAAAQPGFPLAQLKTFVDQEFAQIKHVLHGISLLGQCPDSINAALI 123 n Query: 127 GHGEVWSARLMSAVLNQQGLPAAWLDAREFLRAER---AAQPQVDEGLSYPLLQQLLVQH 183 GE S +M+ VL +G +D E L A + + E ++ H Sbjct: 124 CRGEKMSIAIMAGVLEARGHNVTVIDPVEKLLAVGHYLESTVDIAESTRRIAASRIPADH 183 Query: 184 PGKRLVVTGFISRNNAGETVLLGRNGSDYSATQIGALAGVSRVTIWSDVAGVYSADPRKV 243 +++ GF + N GE V+LGRNGSDYSA + A IW+DV GVY+ DPR+V Sbjct: 184 ---MVLMAGFTAGNEKGELVVLGRNGSDYSAAVLAACLRADCCEIWTDVDGVYTCDPRQV 240 Query: 244 KDACLLPLLRLDEASELARLAAPVLHARTLQPVSGSEIDLQLRCSYTPDQ-----GSTRI 298 DA LL + EA EL+ A VLH RT+ P++ +I ++ + P G++R Sbjct: 241 PDARLLKSMSYQEAMELSYFGAKVLHPRTITPIAQFQIPCLIKNTGNPQAPGTLIGASRD 300 n The second match is another bifunctional protein, product of the gene thr. A. This protein contains the same two domains as met. A (aspartokinase and homoserine dehydrogenase). The alignment covers almost the complete sequences (820 aa), with 30% identities and 49% similarity. The E-value is very low (2 e-95), indicating that thr. A and met. L are likely to be true homologs. Query: 299 ERVLASGTGARIVTSHDDVCLIEFQVPASQDFKLAHKEIDQILKRAQVRPLAVGVHNDRQ 358 E L + +++ + P + + + RA++ + Sbjct: 301 EDELP----VKGISNLNNMAMFSVSGPGMKGMVGMAARVFAAMSRARISVVLITQSSSEY 356 Query: 359 LLQFCYTSEVADSALKILDEA-------GLPGELRLRQGLALVAMVGAGVTRNPLHCHRF 411 + FC A + + E GL L + + LA++++VG G+ +F Sbjct: 357 SISFCVPQSDCVRAERAMQEEFYLELKEGLLEPLAVTERLAIISVVGDGMRTLRGISAKF 416 Query: 412 WQQLKGQPVEFTW--QSDDGISLVAVLRTGPTESLIQGLHQSVFRAEKRIGLVLFGKGNI 469 38

BLAST result - third match The third match is the product of the gene lys. C: aspartokinase III. n This protein contains the aspartokinase domain, but not the homoserine dehydrogenase. n Consequently, the alignment only extends over the first half of the query protein (453 aa). n On this segment, there is a good level of identity (26%) and similarity (42%). n The E-value is very low (7 e– 29), indicating that the two domains are likely to be true homologs. n >gi|16131850|ref|NP_418448. 1| aspartokinase III, lysine sensitive; aspartokinase III, lysine-sensitive [Escherichia coli K 12] Length = 449 Score = 122 bits (307), Expect = 7 e-29 Identities = 121/452 (26%), Positives = 194/452 (42%), Gaps = 25/452 (5%) Query: 16 Sbjct: 8 Query: 75 Sbjct: 64 KFGGSSLADVKCYLRVAGIMAEYSQPDDMMVVSAAGSTTNQLINWLK-LSQTDRLSAHQV 74 KFGG+S+AD R A I+ + ++V+SA+ TN L+ + L +R + KFGGTSVADFDAMNRSADIVLSDANVR-LVVLSASAGITNLLVALAEGLEPGERF---EK 63 QQTLRRYQCDLISGLLPAEEADSLISAFVSDLERLAALLDSGINDAVYAEVVGHGEVWSA 134 +R Q ++ L I + ++ LA + A+ E+V HGE+ S LDAIRNIQFAILERLRYPNVIREEIERLLENITVLAEAAALATSPALTDELVSHGELMST 123 Query: 135 RLMSAVLNQQGLPAAWLDAREFLRA-ERAAQPQVDEGLSYPLLQQLLVQHPGKRLVVT-G 192 L +L ++ + A W D R+ +R +R + + D L L+ + LV+T G Sbjct: 124 LLFVEILRERDVQAQWFDVRKVMRTNDRFGRAEPDIAALAELAALQLLPRLNEGLVITQG 183 Query: 193 FISRNNAGETVLLGRNGSDYSATQIGALAGVSRVTIWSDVAGVYSADPRKVKDACLLPLL 252 FI N G T LGR GSDY+A + SRV IW+DV G+Y+ DPR V A + + Sbjct: 184 FIGSENKGRTTTLGRGGSDYTAALLAEALHASRVDIWTDVPGIYTTDPRVVSAAKRIDEI 243 Query: 253 RLDEASELARLAAPVLHARTLQPVSGSEIDLQLRCSYTPDQGSTRI-----ERVLA 303 EA+E+A A VLH TL P S+I + + S P G T + R LA Sbjct: 244 AFAEAAEMATFGAKVLHPATLLPAVRSDIPVFVGSSKDPRAGGTLVCNKTENPPLFRALA 303 Query: 304 SGTGARIVTSHDDVCLIEFQVPASQDFKLAHKEIDQILKRAQVRPLAVGVHNDRQLLQFC 363 ++T H L A LA I L +A+ L Sbjct: 304 LRRNQTLLTLHSLNMLHSRGFLAEVFGILARHNISVDLITTSEVSVAL-------TLDTT 356 Query: 364 YTSEVADSAL--KILDEAGLPGELRLRQGLALVAMVGAGVTRNPLHCHRFWQQLKGQPVE 421 ++ D+ L +L E + + +GLALVA++G +++ + L+ + Sbjct: 357 GSTSTGDTLLTQSLLMELSALCRVEVEEGLALVALIGNDLSKACGVGKEVFGVLEPFNIR 416 Query: 422 FTWQSDDGISLVAVLRTGPTESLIQGLHQSVF 453 39

BLAST result - fourth match The fourth match is a gamma-glutamate kinase, product of pro. B. n The match has the same level of identity (30%) and similarity (51%) as the second match (thr. A). n However, this match only extends over 56 aa, whereas the alignment between thr. A and met. L extends over 821 aa. n The significance of the match is thus much lower: the E-value is quite high (0. 28) suggesting that the similarity could be an artefact. n This clearly illustrates the fact that the important parameter to evaluate the significance of an alignment is the Evalue, not the percentage of similarity ! n >gi|16128228|ref|NP_414777. 1| gamma-glutamate kinase [Escherichia coli K 12] Length = 367 Score = 31. 2 bits (69), Expect = 0. 28 Identities = 17/56 (30%), Positives = 29/56 (51%) Query: 194 ISRNNAGETVLLGRNGSDYSATQIGALAGVSRVTIWSDVAGVYSADPRKVKDACLL 249 I+ N+A T + +D + LAG ++ + +D G+Y+ADPR A L+ Sbjct: 133 INENDAVATAEIKVGDNDNLSALAAILAGADKLLLLTDQKGLYTADPRSNPQAELI 188 40

BLAST result - summary n Database: /Users/jvanheld/rsatools/data/genomes/Escherichia_coli_K 12/genome/NC_000913. faa Posted date: Sep 8, 2004 12: 13 PM Number of letters in database: 1, 351, 322 Number of sequences in database: 4242 The last part of the BLAST result gives some statistics about the search: q q Lambda 0. 320 Gapped Lambda 0. 267 K H 0. 136 K H 0. 0410 0. 397 q Number of hits Number of sequences in the DB … 0. 140 Matrix: BLOSUM 62 Gap Penalties: Existence: 11, Extension: 1 Number of Hits to DB: 2, 199, 628 Number of Sequences: 4242 Number of extensions: 96525 Number of successful extensions: 290 Number of sequences better than 1. 0: 4 Number of HSP's better than 1. 0 without gapping: 4 Number of HSP's successfully gapped in prelim test: 0 Number of HSP's that attempted gapping in prelim test: 279 Number of HSP's gapped (non-prelim): 5 length of query: 810 length of database: 1, 351, 322 effective HSP length: 92 effective length of query: 718 effective length of database: 961, 058 effective search space: 690039644 effective search space used: 690039644 T: 11 A: 40 X 1: 16 ( 7. 4 bits) X 2: 38 (14. 6 bits) X 3: 64 (24. 7 bits) S 1: 41 (21. 8 bits) S 2: 65 (29. 6 bits) 41

BLAST – examples of results
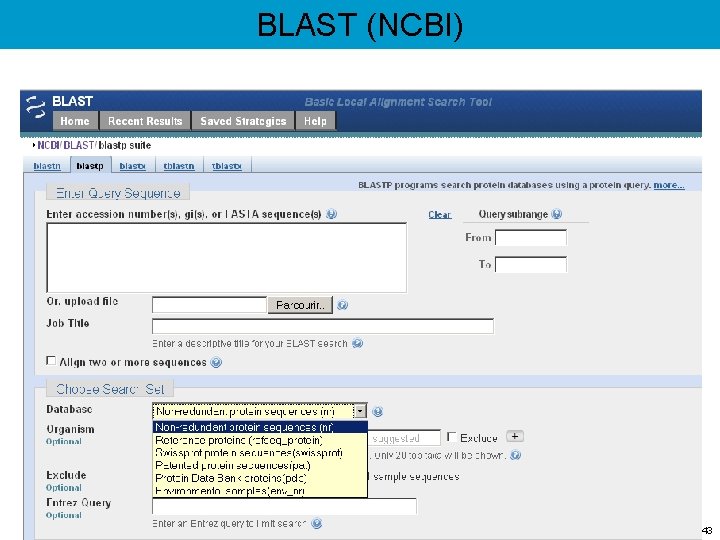
BLAST (NCBI) 43

BLAST (NCBI) 44
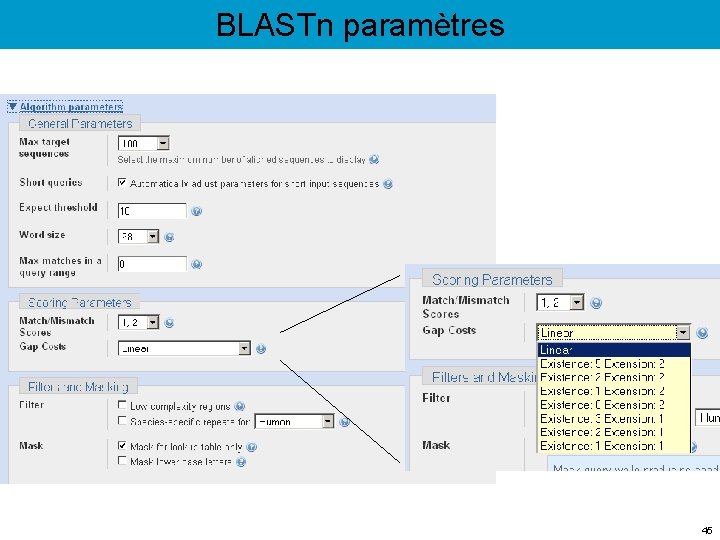
BLASTn paramètres 45

BLASTp paramètres 46
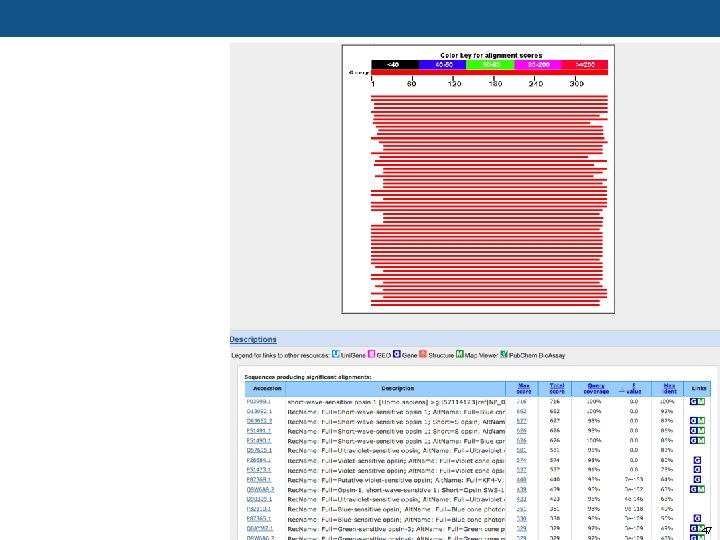
47

Checking expected values with random sequences (“negative control”)
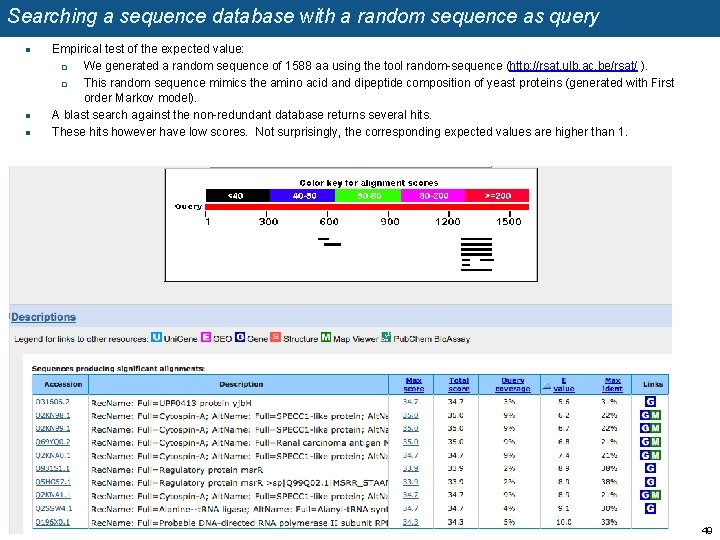
Searching a sequence database with a random sequence as query n n n Empirical test of the expected value: q We generated a random sequence of 1588 aa using the tool random-sequence (http: //rsat. ulb. ac. be/rsat/ ). q This random sequence mimics the amino acid and dipeptide composition of yeast proteins (generated with First order Markov model). A blast search against the non-redundant database returns several hits. These hits however have low scores. Not surprisingly, the corresponding expected values are higher than 1. 49

blastp with random sequences n n pblast of 10 random sequences against the non-redundant database. q Between 1 and 15 matches per trial. q Was this to be expected ? This corresponds pretty well to the expectation q On the NCBI BLAST web server, the default threshold on expect has been set to 10. q We thus expect, for each request, an average of 10 matches by chance. q We indeed observe this order of magnitude when submitting random sequences. 50

The modalities of BLAST

DNA versus protein searches n When the query is a coding DNA sequence, it is recommended to apply searches with the translated rather than raw DNA sequences q q q This allows to introduce a substitution matrix (PAM, BLOSUM, . . . ), which better reflects the evolutionary changes. It has been shown that some distant relationships can be detected with translated searches, but escape detection with the DNA search. It is easier to filter out low complexity regions from proteins than from DNA sequences. 52
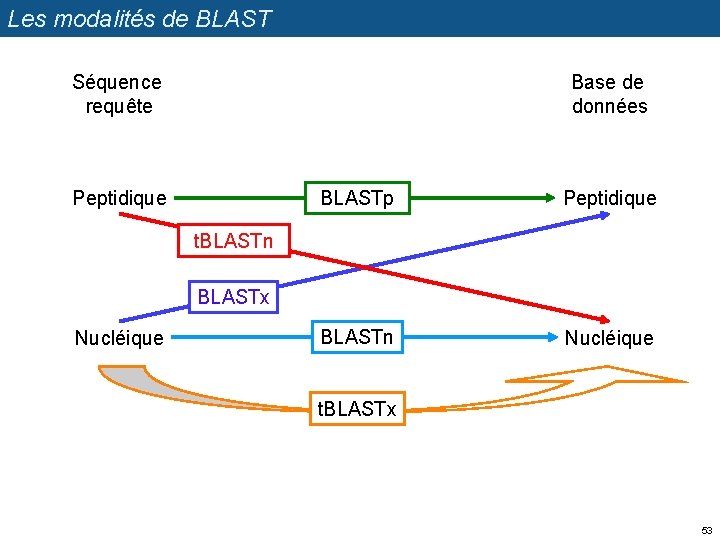
Les modalités de BLAST Séquence requête Base de données Peptidique BLASTp Peptidique BLASTn Nucléique t. BLASTn BLASTx Nucléique t. BLASTx 53
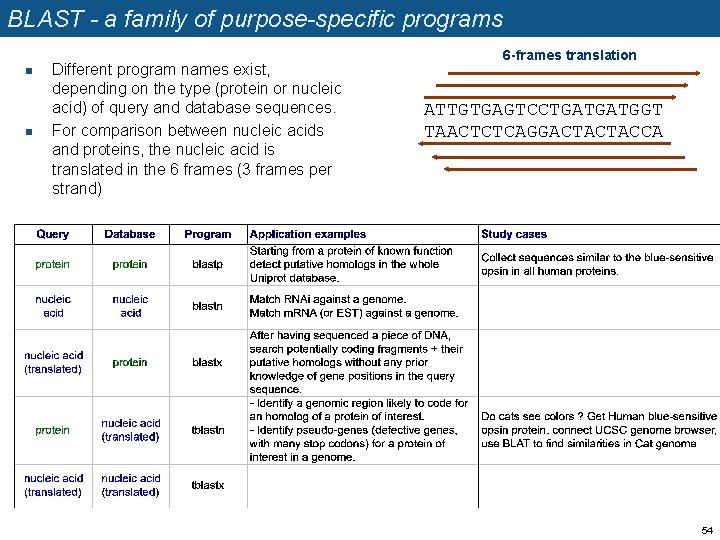
BLAST - a family of purpose-specific programs n n Different program names exist, depending on the type (protein or nucleic acid) of query and database sequences. For comparison between nucleic acids and proteins, the nucleic acid is translated in the 6 frames (3 frames per strand) 6 -frames translation ATTGTGAGTCCTGATGATGGT TAACTCTCAGGACTACTACCA 54
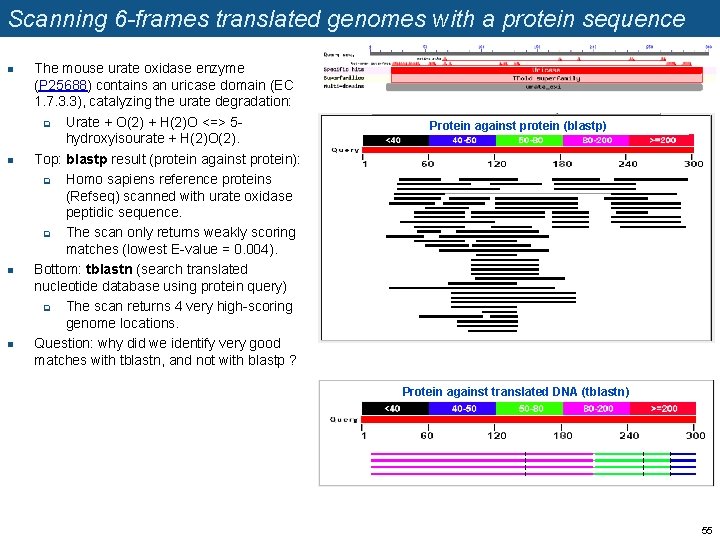
Scanning 6 -frames translated genomes with a protein sequence n n The mouse urate oxidase enzyme (P 25688) contains an uricase domain (EC 1. 7. 3. 3), catalyzing the urate degradation: q Urate + O(2) + H(2)O <=> 5 hydroxyisourate + H(2)O(2). Top: blastp result (protein against protein): q Homo sapiens reference proteins (Refseq) scanned with urate oxidase peptidic sequence. q The scan only returns weakly scoring matches (lowest E-value = 0. 004). Bottom: tblastn (search translated nucleotide database using protein query) q The scan returns 4 very high-scoring genome locations. Question: why did we identify very good matches with tblastn, and not with blastp ? Protein against protein (blastp) Protein against translated DNA (tblastn) 55
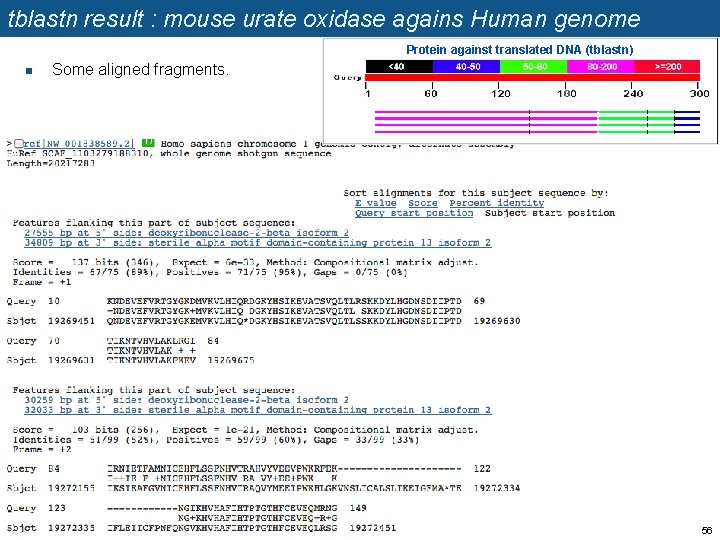
tblastn result : mouse urate oxidase agains Human genome Protein against translated DNA (tblastn) n Some aligned fragments. >ref|NW_001838589. 2| Download subject sequence NW_001838589 spanning the HSP Homo sapiens chromosome 1 genomic contig, alternate assembly Hu. Ref SCAF_1103279188310, whole genome shotgun sequence Length=20217283 Sort alignments for this subject sequence by: E value Score Percent identity Query start position Subject start position Features flanking this part of subject sequence: 27555 bp at 5' side: deoxyribonuclease-2 -beta isoform 2 34809 bp at 3' side: sterile alpha motif domain-containing protein 13 isoform 2 Score = 137 bits (346), Expect = 6 e-33, Method: Compositional matrix adjust. Identities = 67/75 (89%), Positives = 71/75 (95%), Gaps = 0/75 (0%) Frame = +1 Query 10 KNDEVEFVRTGYGKDMVKVLHIQRDGKYHSIKEVATSVQLTLRSKKDYLHGDNSDIIPTD 69 +NDEVEFVRTGYGK+MVKVLHIQ DGKYHSIKEVATSVQLTL SKKDYLHGDNSDIIPTD Sbjct 19269451 QNDEVEFVRTGYGKEMVKVLHIQ*DGKYHSIKEVATSVQLTLSSKKDYLHGDNSDIIPTD 19269630 Query 70 TIKNTVHVLAKLRGI 84 TIKNTVHVLAK + + Sbjct 19269631 TIKNTVHVLAKFKEV 19269675 56

tblastn example – do cat see colors ? n n n Approach: match the peptidic sequence of the Human Short-wave-sensitive opsin (blue) against the complete cat genome (6 -frames translated). Tool: BLAT tool at UCSC genome browser (http: //genome. ucsc. edu/) Result: 3 matches in cat genome q q q Short wave (blue) sensitive opsin rhodopsin Long wave (red) sensitive opsin. Partial match with 1 exon, but sufficient to “fish out” the cat OPN 1 LW gene. 57

Blastx example: scanning a genomic sequence with a protein sequence
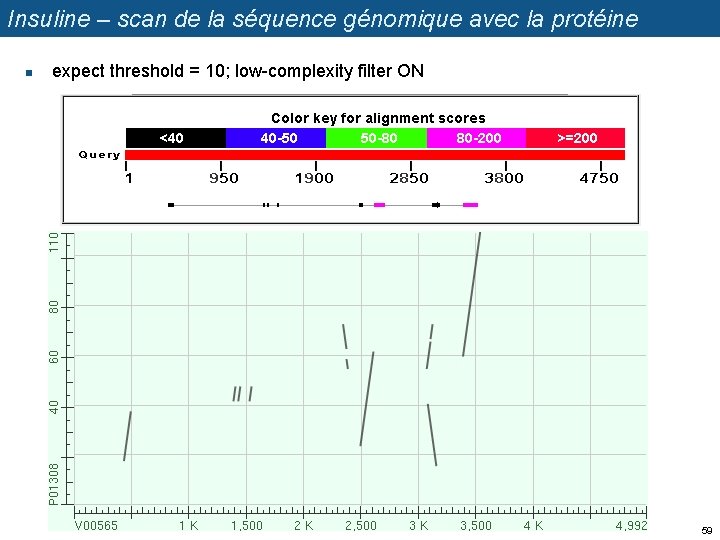
Insuline – scan de la séquence génomique avec la protéine n expect threshold = 10; low-complexity filter ON 59
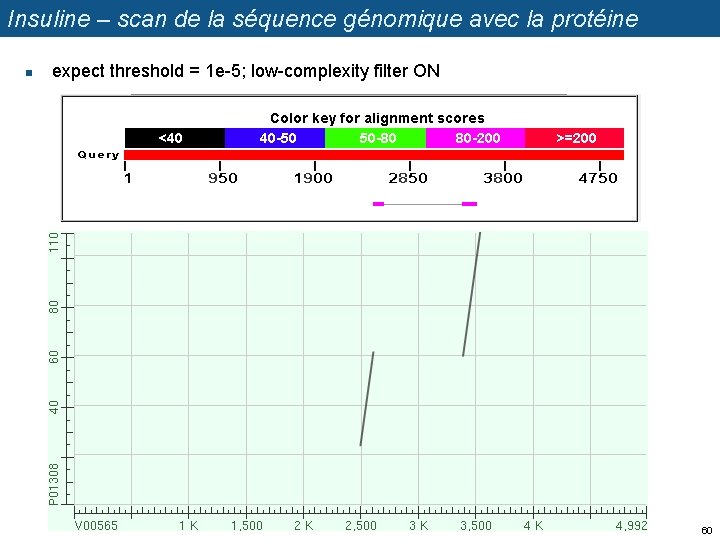
Insuline – scan de la séquence génomique avec la protéine n expect threshold = 1 e-5; low-complexity filter ON 60
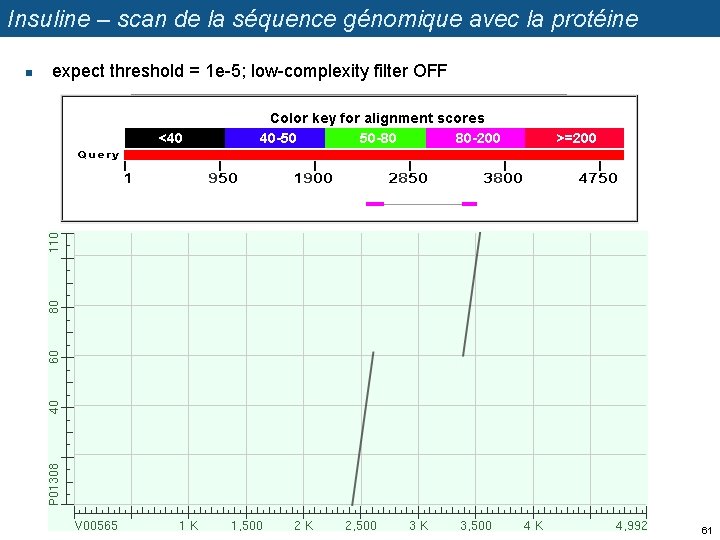
Insuline – scan de la séquence génomique avec la protéine n expect threshold = 1 e-5; low-complexity filter OFF 61
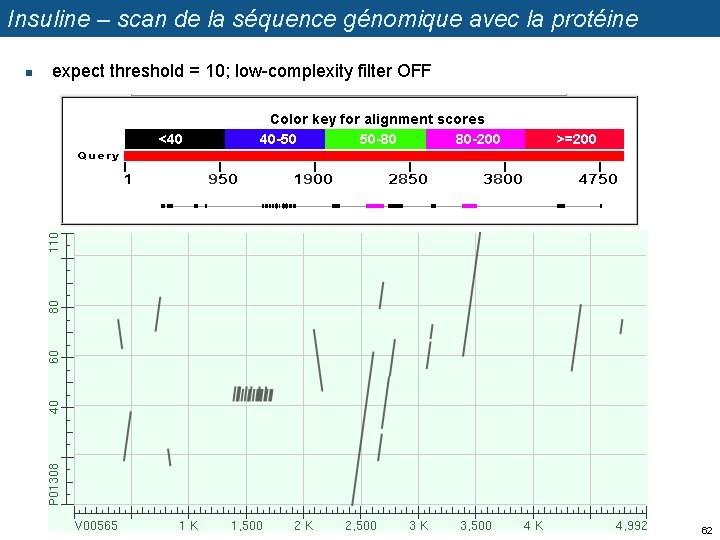
Insuline – scan de la séquence génomique avec la protéine n expect threshold = 10; low-complexity filter OFF 62
- Slides: 62