Introduction to bioinformatics Lecture 7 Pairwise Sequence Alignment

Introduction to bioinformatics Lecture 7 Pairwise Sequence Alignment (III) and Deriving Amino Acid Substitution Matrices (I)
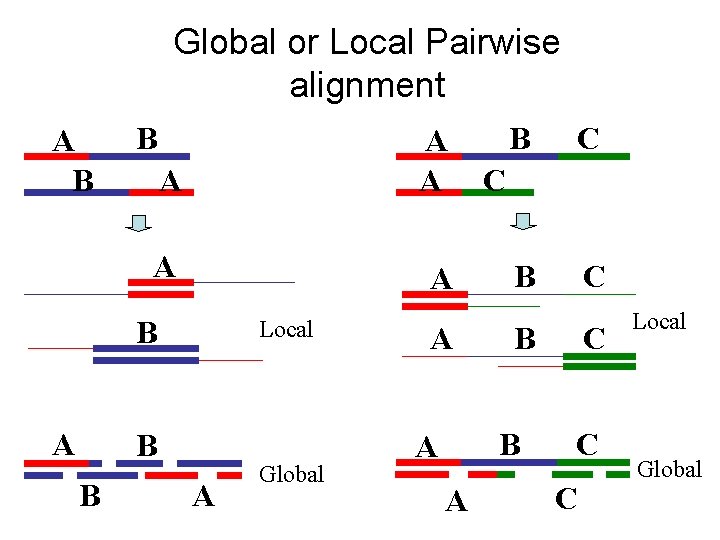
Global or Local Pairwise alignment A B B A A A B A Local B B A Global A B C C B B A A C C C Local Global
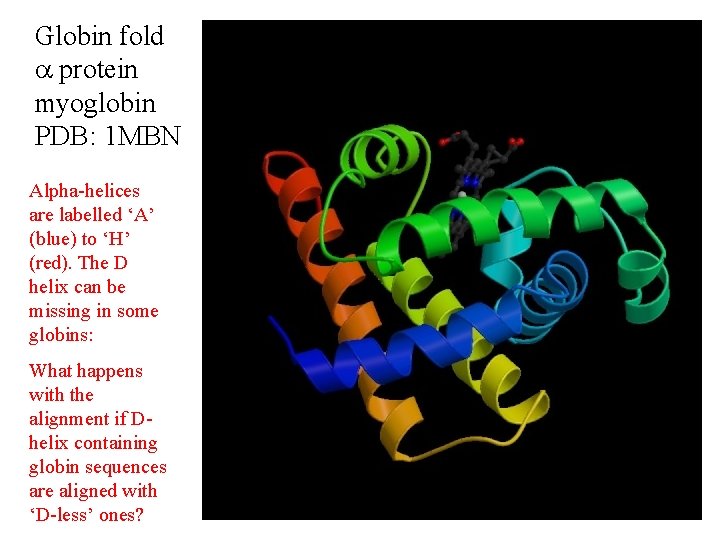
Globin fold protein myoglobin PDB: 1 MBN Alpha-helices are labelled ‘A’ (blue) to ‘H’ (red). The D helix can be missing in some globins: What happens with the alignment if Dhelix containing globin sequences are aligned with ‘D-less’ ones?
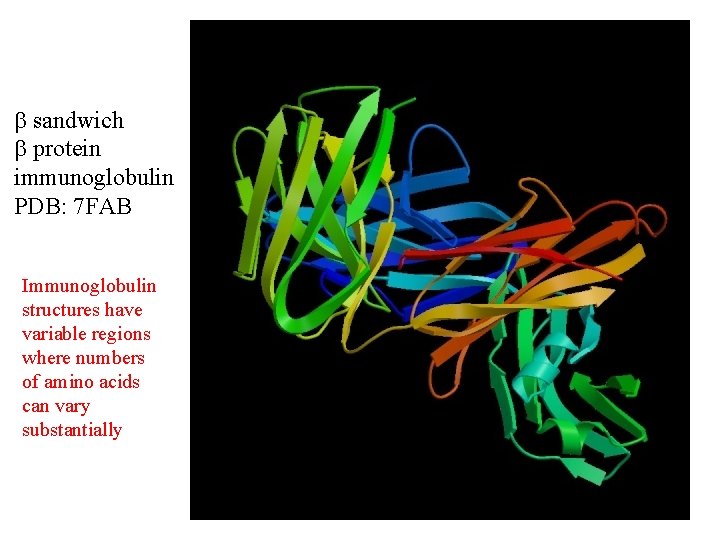
sandwich protein immunoglobulin PDB: 7 FAB Immunoglobulin structures have variable regions where numbers of amino acids can vary substantially
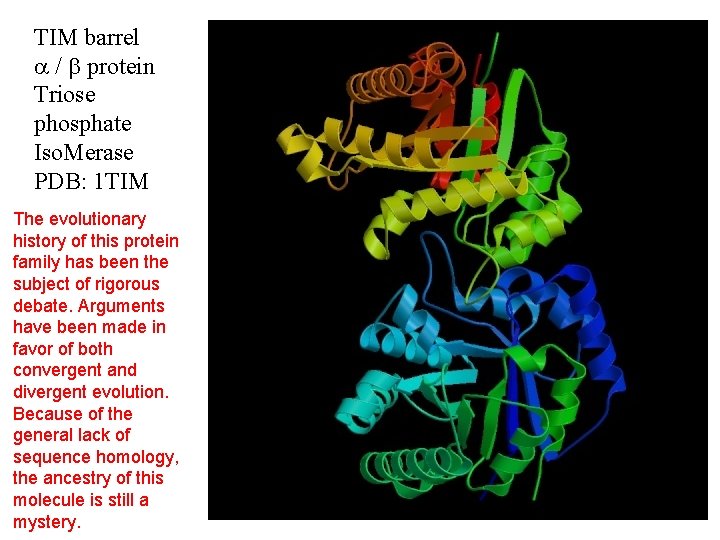
TIM barrel / protein Triose phosphate Iso. Merase PDB: 1 TIM The evolutionary history of this protein family has been the subject of rigorous debate. Arguments have been made in favor of both convergent and divergent evolution. Because of the general lack of sequence homology, the ancestry of this molecule is still a mystery.

Pyruvate kinase Phosphotransferase barrel regulatory domain / barrel catalytic substrate binding domain / nucleotide binding domain

What does all this mean for alignments? • Alignments need to be able to skip secondary structural elements to complete domains (i. e. putting gaps opposite these motifs in the shorter sequence). • Depending on gap penalties chosen, the algorithm might have difficulty with making such long gaps (for example when using high affine gap penalties), resulting in incorrect alignment. • Alignments are only meaningful for homologous sequences (with a common ancestor)

There are three kinds of pairwise alignments n n n Global alignment – align all residues in both sequences; all gaps are penalised Semi-global alignment – align all residues in both sequences; end gaps are not penalised (zero end gap penalties) Local alignment – align one part of each sequence; end gaps are not applicable

Easy global DP recipe for using affine gap penalties (after j-1 Gotoh) Penalty = Pi + gap_length*Pe i-1 n n Max{S 0<x<i-1, j-1 - Pi - (i-x-1)Px} Si, j = si, j + Max Si-1, j-1 Max{Si-1, 0<y<j-1 - Pi - (j-y-1)Px} M[i, j] is optimal alignment (highest scoring alignment until [i, j]) At each cell [i, j] in search matrix, check Max coming from: any cell in preceding row until j-2: add score for cell[i, j] minus appropriate gap penalties; any cell in preceding column until i-2: add score for cell[i, j] minus appropriate gap penalties; or cell[i-1, j-1]: add score for cell[i, j] n Select highest scoring cell in bottom row and rightmost column and do trace-back

Let’s do an example: global alignment Gotoh’s DP algorithm with affine gap penalties (PAM 250, Pi=10, Pe=2) D W V T A L K 0 -12 -14 -16 -18 -20 -22 -24 T -12 0 -17 -14 -13 -17 -19 -22 -3 D -14 -8 -7 -14 -13 2 -2 W -16 -21 9 -13 -19 -18 -2 6 -3 V -18 -20 13 -3 -16 -1 -3 5 L -20 -22 -18 -1 14 K -22 -20 -21 -24 -42 D W V T A L K T 0 -5 0 3 1 1 0 D 4 -7 -2 0 0 -4 0 W -7 17 -6 -5 -6 -2 V -2 -6 4 0 0 L -4 -2 2 1 K 0 -3 -2 0 PAM 250 -22 -42 -1 -14 -12 -41 -18 -16 -14 -12 0 Cell (D 2, T 4) can alternatively come from two cells (same score): ‘high-road’ or ‘low-road’ Row and column ‘ 0’ are filled with 0, -12, -14, -16, … if global alignment is used (for N-terminal endgaps); also extra row and column at the end to calculate the score including C-terminal end-gap penalties. Note that only ‘non-diagonal’ arrows are indicated for clarity (no arrow means that you go back to earlier diagonal cell).

Let’s do another example: semi-global alignment Gotoh’s DP algorithm with affine gap penalties (PAM 250, Pi=10, Pe=2) D W V T A L K T 0 -5 0 3 1 1 0 D 4 -7 -2 0 0 -4 W -7 17 -6 -5 -6 V -2 -6 4 0 L -4 -2 2 K 0 -3 -2 D W V T T 0 -5 0 3 0 D 4 -7 -7 -2 -3 W -7 21 -13 0 2 -2 V -2 -13 25 1 -2 6 -3 L 0 -1 -3 5 K A L K 9 PAM 250 Starting row and column ‘ 0’, and extra column at right or extra row at bottom is not necessary when using semi global alignment (zero endgaps). Rest works as under global alignment.

Easy local DP recipe for using affine gap penalties (after j-1 Gotoh) Penalty = Pi + gap_length*Pe Si, j = Max i-1 n n Si, j + Max{S 0<x<i-1, j-1 - Pi - (i-x-1)Px} Si, j + Si-1, j-1 Si, j + Max {Si-1, 0<y<j-1 - Pi - (j-y-1)Px} 0 M[i, j] is optimal alignment (highest scoring alignment until [i, j]) At each cell [i, j] in search matrix, check Max coming from: any cell in preceding row until j-2: add score for cell[i, j] minus appropriate gap penalties; any cell in preceding column until i-2: add score for cell[i, j] minus appropriate gap penalties; or cell[i-1, j-1]: add score for cell[i, j] n Select highest scoring cell anywhere in matrix and do trace-back until zero-valued cell or start of sequence(s)

Let’s do yet another example: local alignment Gotoh’s DP algorithm with affine gap penalties (PAM 250, Pi=10, Pe=2) D W V T A L K T 0 -5 0 3 1 1 0 D 4 -7 -2 0 0 -4 W -7 17 -6 -5 -6 V -2 -6 4 0 L -4 -2 2 K 0 -3 -2 D W V T T 0 0 0 3 0 D 4 0 0 0 -2 -3 W 0 21 0 0 0 2 -2 V 0 0 25 9 1 -2 6 -3 L 0 0 11 0 -1 -3 5 K 0 0 A L K PAM 250 Extra start/end columns/rows not necessary (no end-gaps). Each negative scoring cell is set to zero. Highest scoring cell may be found anywhere in search matrix after calculating it. Trace highest scoring cell back to first cell with zero value (or the beginning of one or both sequences)

For your first exam D 1: Make sure you understand can carry out 1. the ‘simple’ DP algorithm (for linear gap penalties but with the extension for affine gap penalties) and 2. Gotoh’s algorithm for global, semi-global and local alignment! Gotoh, O. An Improved Algorithm for Matching Biological Sequences. J. Mol. Biol. , 162, pp. 705 -708, 1982.

Introduction to Bioinformatics Deriving Amino Acid Substitution matrices (I)
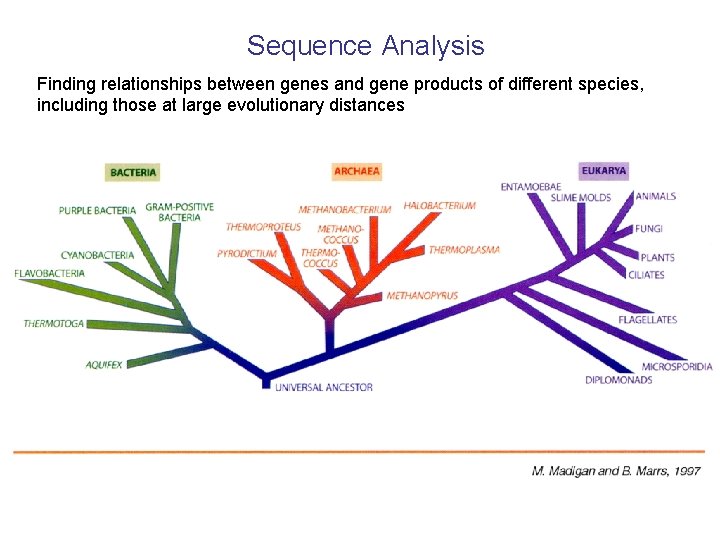
Sequence Analysis Finding relationships between genes and gene products of different species, including those at large evolutionary distances

Archaea Domain Archaea is mostly composed of cells that live in extreme environments. While they are able to live elsewhere, they are usually not found there because outside of extreme environments they are competitively excluded by other organisms. Species of the domain Archaea are not inhibited by antibiotics, lack peptidoglycan in their cell wall (unlike bacteria, which have this sugar/polypeptide compound), and can have branched carbon chains in their membrane lipids of the phospholipid bilayer. It is believed that Archaea are very similar to prokaryotes (e. g. bacteria) that inhabited the earth billions of years ago. It is also believed that eukaryotes evolved from Archaea, because they share many m. RNA sequences, have similar RNA polymerases, and have introns. Therefore, it is believed that the domains Archaea and Bacteria branched from each other very early in history, and membrane infolding* produced eukaryotic cells in the archaean branch approximately 1. 7 billion years ago. There are three main groups of Archaea: extreme halophiles (salt), methanogens (methane producing anaerobes), and hyperthermophiles (e. g. living at temperatures >100º C!). *Membrane infolding is believed to have led to the nucleus of eukaryotic cells, which is a membrane-enveloped cell organelle that holds the cellular DNA. Prokaryotic cells are more primitive and do not have a nucleus.
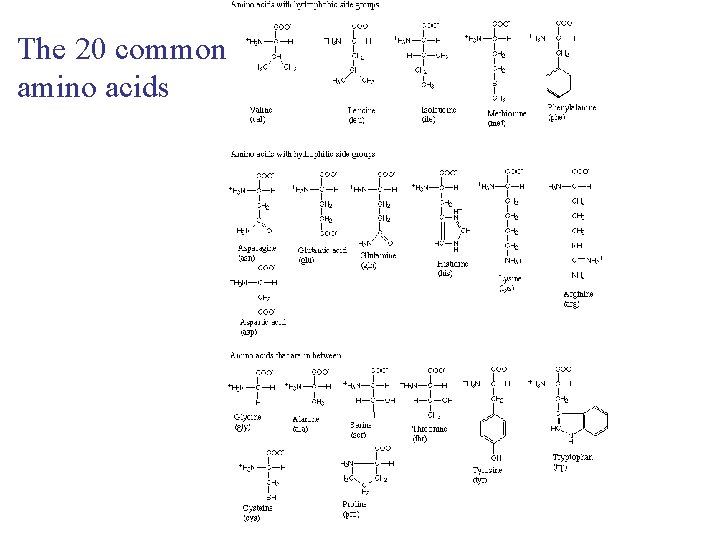
The 20 common amino acids

Example of sequence database entry for Genbank LOCUS DRODPPC 4001 bp INV 15 -MAR-1990 DEFINITION D. melanogaster decapentaplegic gene complex (DPP-C), complete cds. ACCESSION M 30116 KEYWORDS. SOURCE D. melanogaster, c. DNA to m. RNA. ORGANISM Drosophila melanogaster Eurkaryote; mitochondrial eukaryotes; Metazoa; Arthropoda; Tracheata; Insecta; Pterygota; Diptera; Brachycera; Muscomorpha; Ephydroidea; Drosophilidae; Drosophilia. REFER�ENCE 1 (bases 1 to 4001) AUTHORS Padgett, R. W. , St Johnston, R. D. and Gelbart, W. M. TITLE A transcript from a Drosophila pattern gene predicts a protein homologous to the transforming growth factor-beta family JOURNAL Nature 325, 81 -84 (1987) MEDLINE 87090408 COMMENT The initiation codon could be at either 1188 -1190 or 1587 -1589 FEATURES Location/Qualifiers source 1. . 4001 /organism=“Drosophila melanogaster” /db_xref=“taxon: 7227” m. RNA <1. . 3918 /gene=“dpp” /note=“decapentaplegic protein m. RNA” /db_xref=“Fly. Base: FBgn 0000490” gene 1. . 4001 /note=“decapentaplegic” /gene=“dpp” /allele=“” /db_xref=“Fly. Base: FBgn 0000490” CDS 1188. . 2954 /gene=“dpp” /note=“decapentaplegic protein (1188 could be 1587)” /codon_start=1 /db_xref=“Fly. Base: FBgn 0000490” /db_xref=“PID: g 157292” /translation=“MRAWLLLLAVLATFQTIVRVASTEDISQRFIAAIAPVAAHIPLA SASGSGSGRSGSRSVGASTSTALAKAFNPFSEPASFSDSDKSHRSKTNKKPSKSDANR ………… LGYDAYYCHGKCPFPLADHFNSTNAVVQTLVNNMNPGKVPKACCVPTQLDSVAMLYL NDQSTBVVLKNYQEMTBBGCGCR” BASE COUNT 1170 a 1078 c 956 g 797 t ORIGIN 1 gtcgttcaac agcgctgatc gagtttaaat ctataccgaa atgagcggcg gaaagtgagc 61 cacttggcgt gaacccaaag ctttcgagga aaattctcgg acccccatat acaaatatcg 121 gaaaaagtat cgaacagttt cgcga agcgttaaga tcgcccaaag atctccgtgc 181 ggaaacaaag aaattgaggc actattaaga gattgttgtt gtgcgcgagt gtgtgtcttc 241 agctgggtgt gtggaatgtc aactgacggg ttgtaaaggg aaaccctgaa atccgaacgg 301 ccagccaaag caaataaagc tgtgaatacg aattaagtac aacagt tactgaaaca 361 gatacagatt cggattcgaa tagagaaaca gatactggag atgcccccag aaacaattca 421 attgcaaata tagtgcgttg cgcgagtgcc agtggaaaaa tatgtggatt acctgcgaac 481 cgtccgccca aggagccgcc gggtgacagg tgtatccccc aggataccaa cccgagccca 541 gaccgagatc cacatccaga tcccgaccgc agggtgccag tgtgtcatgt gccgcggcat 601 accgca gccacatcta ccgaccaggt gcgcctcgaa tgcggcaaca caattttcaa ……………. 3841 aactgtataa acaaaacgta tgccctataa atatatgaat aactatctac atcgttatgc 3901 gttctaagctcgaat aaatccgtac acgttaatta atctagaatc gtaagaccta 3961 acgcgtaagc tcagcatgtt ggataaatta atagaaacga g //

Example of sequence database entry for SWISSPROT (now UNIPROT) ID AC DT DT DT DE GN OS OC RN RP RM RA RL CC CC CC DR DR DR KW FT FT FT SQ DECA_DROME STANDARD; PRT; 588 AA. P 07713; 01 -APR-1988 (REL. 07, CREATED) 01 -APR-1988 (REL. 07, LAST SEQUENCE UPDATE) 01 -FEB-1995 (REL. 31, LAST ANNOTATION UPDATE) DECAPENTAPLEGIC PROTEIN PRECURSOR (DPP-C PROTEIN). DPP. DROSOPHILA MELANOGASTER (FRUIT FLY). EUKARYOTA; METAZOA; ARTHROPODA; INSECTA; DIPTERA. [1] SEQUENCE FROM N. A. 87090408 PADGETT R. W. , ST JOHNSTON R. D. , GELBART W. M. ; NATURE 325: 81 -84 (1987) [2] CHARACTERIZATION, AND SEQUENCE OF 457 -476. 90258853 PANGANIBAN G. E. F. , RASHKA K. E. , NEITZEL M. D. , HOFFMANN F. M. ; MOL. CELL. BIOL. 10: 2669 -2677(1990). -!- FUNCTION: DPP IS REQUIRED FOR THE PROPER DEVELOPMENT OF THE EMBRYONIC DOORSAL HYPODERM, FOR VIABILITY OF LARVAE AND FOR CELL VIABILITY OF THE EPITHELIAL CELLS IN THE IMAGINAL DISKS. -!- SUBUNIT: HOMODIMER, DISULFIDE-LINKED. -!- SIMILARITY: TO OTHER GROWTH FACTORS OF THE TGF-BETA FAMILY. EMBL; M 30116; DMDPPC. PIR; A 26158. HSSP; P 08112; 1 TFG. FLYBASE; FBGN 0000490; DPP. PROSITE; PS 00250; TGF_BETA. GROWTH FACTOR; DIFFERENTIATION; SIGNAL 1 ? POTENTIAL. PROPEP ? 456 CHAIN 457 588 DECAPENTAPLEGIC PROTEIN. DISULFID 487 553 BY SIMILARITY. DISULFID 516 585 BY SIMILARITY. DISULFID 520 587 BY SIMILARITY. DISULFID 552 INTERCHAIN (BY SIMILARITY). CARBOHYD 120 POTENTIAL. CARBOHYD 342 POTENTIAL. CARBOHYD 377 POTENTIAL. CARBOHYD 529 POTENTIAL. SEQUENCE 588 AA; 65850 MW; 1768420 CN; MRAWLLLLAV LATFQTIVRV ASTEDISQRF IAAIAPVAAH IPLASASGSG SGRSGSRSVG ASTSTAGAKA FNRFSEPASF SDSDKSHRSK TNKKPSKSDA NRQFNEVHKP RTDQLENSKN KSKQLVNKPN HNKMAVKEQR SHHKKSHHHR SHQPKQASAS TESHQSSSIE SIFVEEPTLV LDREVASINV PANAKAIIAE QGPSTYSKEA LIKDKLKPDP STYLVEIKSL LSLFNMKRPP KIDRSKIIIP EPMKKLYAEI MGHELDSVNI PKPGLLTKSA NTVRSFTHKD SKIDDRFPHH HRFRLHFDVK SIPADEKLKA AELQLTRDAL SQQVVASRSS ANRTRYQBLV YDITRVGVRG QREPSYLLLD TKTBRLNSTD TVSLDVQPAV DRWLASPQRN YGLLVEVRTV RSLKPAPHHH VRLRRSADEA HERWQHKQPL LFTYTDDGRH DARSIRDVSG GEGGGKGGRN KRHARRPTRR KNHDDTCRRH SLYVDFSDVG WDDWIVAPLG YDAYYCHGKC PFPLADHRNS TNHAVVQTLV NNMNPGKBPK ACCBPTQLDS VAMLYLNDQS TVVLKNYQEM TVVGCGCR

What to align, nucleotide or amino acid sequences? If you think you have an Open Reading Frame (ORF) then align at protein level – (i) Many mutations within DNA are synonymous, leading to overestimation of sequence divergence if compared at the DNA level. – (ii) Evolutionary relationships can be more finely expressed using a 20× 20 amino acid exchange table than using nucleotide exchanges. – (iii) DNA sequences contain non-coding regions which should be avoided in homology searches. Still an issue when translating into (six) protein sequences through a codon table. – (iv) Searching at protein level: frameshifts can occur, leading to stretches of incorrect amino acids and possibly elongation of sequences due to missed stop codons. But frameshifts normally result in stretches of highly unlikely amino acids: can be used as a signal to trace.









A 2 R -2 6 N 0 0 2 D 0 -1 2 4 PAM 250 matrix C -2 -4 -4 -5 12 Q 0 1 1 2 -5 4 E 0 -1 1 3 -5 2 4 G 1 -3 0 1 -3 -1 0 2 1 -3 1 -2 H -1 2 3 5 6 I -1 -2 -2 -2 -3 -2 5 L -2 -3 -3 -4 -6 -2 -3 -4 -2 2 1 0 -5 1 0 -2 6 K -1 3 M -1 0 -2 -3 -5 -1 -2 -3 -2 2 4 0 6 F -4 -4 -4 -6 -4 -5 -5 -5 -2 1 2 -5 0 5 WR exchange is too large (due to paucity of data) 9 P 1 0 -1 -1 -3 S 1 0 0 -1 0 1 -1 -1 -3 0 -2 -3 1 2 T 1 -1 0 0 -2 -1 0 0 -1 -3 0 1 W -6 0 -1 -1 0 -2 -3 -1 -2 -5 0 -2 2 -4 -7 -8 -5 -7 -7 -3 -5 -2 -3 -4 Y -3 -4 -2 -4 0 -4 -4 -5 0 -1 -1 -4 -2 0 -6 -2 -5 17 7 -5 -3 -3 B 0 -1 2 3 -4 1 2 0 1 -2 -3 1 -2 -5 -1 0 0 -5 -3 -2 2 Z 0 0 1 3 -5 3 3 -1 2 -2 -3 0 -2 -5 0 0 -1 -6 -4 -2 2 3 A R N D Q E H K P S B Z I L 2 -1 -1 -1 0 10 0 -2 -2 -2 -1 -2 G 2 -2 3 V C 4 6 M F 0 -6 -2 T W Y 4 V

PAM model The scores derived through the PAM model are an accurate description of the information content (or the relative entropy) of an alignment (Altschul, 1991). PAM-1 corresponds to about 1 million years of evolution PAM-120 has the largest information content of the PAM matrix series PAM-250 is the traditionally most popular matrix

PAM / MDM / Dayhoff -- summary The late Margaret Dayhoff was a pioneer in protein databasing and comparison. She and her coworkers developed a model of protein evolution which resulted in the development of a set of widely used substitution matrices. These are frequently called Dayhoff, MDM (Mutation Data Matrix), or PAM (Percent Accepted Mutation) matrices: • Derived from global alignments of closely related sequences. • Matrices for greater evolutionary distances are extrapolated from those for lesser ones. • The number associated with the matrix (PAM 40, PAM 100) refers to the evolutionary distance; greater numbers correspond to greater distances. • Several later groups have attempted to extend Dayhoff's methodology or reapply her analysis using later databases with more examples. Extensions: • Jones, Thornton and coworkers used the same methodology as Dayhoff but with modern databases (CABIOS 8: 275 - 1992) • Gonnett and coworkers (Science 256: 1443 - 1992) used a slightly different (but theoretically equivalent) methodology • Henikoff & Henikoff (Proteins 17: 49 - 1993) compared these two newer versions of the PAM matrices with Dayhoff's originals.
- Slides: 32