Introduction to Bioinformatics Lecture 20 Sequencing genomes Nucleic

Introduction to Bioinformatics Lecture 20: Sequencing genomes

Nucleic Acid Basics • Nucleic Acids Are Polymers • Each Monomer Consists of Three Moieties: Nucleotide A Base + A Ribose Sugar + A Phosphate Nucleoside • A Base Can be One of the Five Rings:
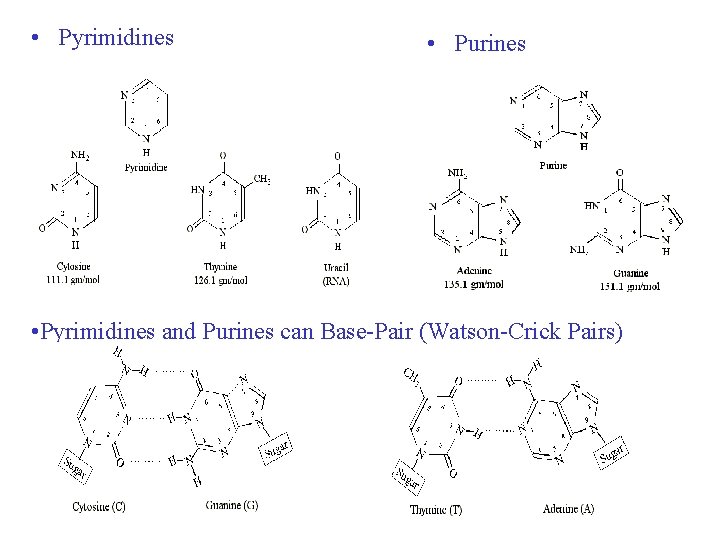
• Pyrimidines • Purines • Pyrimidines and Purines can Base-Pair (Watson-Crick Pairs)
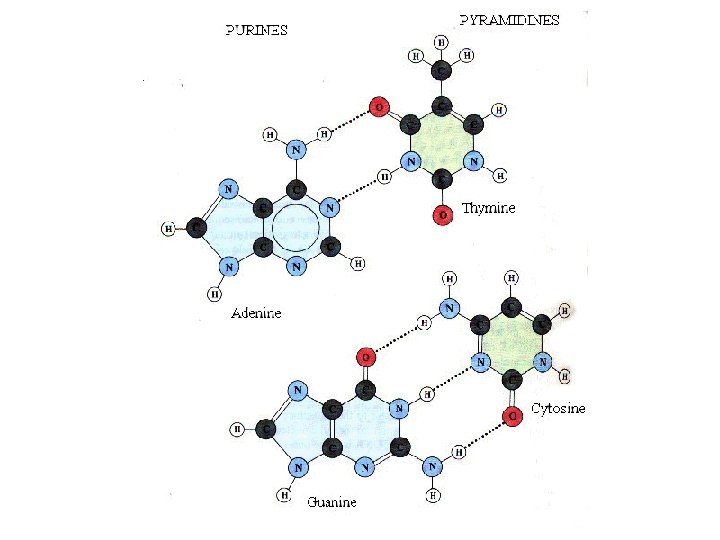
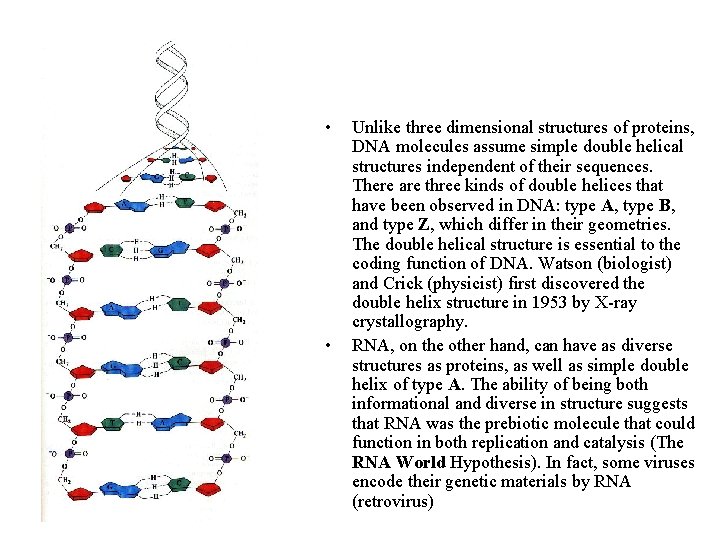
• • Unlike three dimensional structures of proteins, DNA molecules assume simple double helical structures independent of their sequences. There are three kinds of double helices that have been observed in DNA: type A, type B, and type Z, which differ in their geometries. The double helical structure is essential to the coding function of DNA. Watson (biologist) and Crick (physicist) first discovered the double helix structure in 1953 by X-ray crystallography. RNA, on the other hand, can have as diverse structures as proteins, as well as simple double helix of type A. The ability of being both informational and diverse in structure suggests that RNA was the prebiotic molecule that could function in both replication and catalysis (The RNA World Hypothesis). In fact, some viruses encode their genetic materials by RNA (retrovirus)

Forces That Stabilize Nucleic Acid Double Helix • There are two major forces that contribute to stability of helix formation – Hydrogen bonding in base-pairing – Hydrophobic interactions in base stacking 5’ 3’ 3’ 5’ Same strand stacking cross-strand stacking
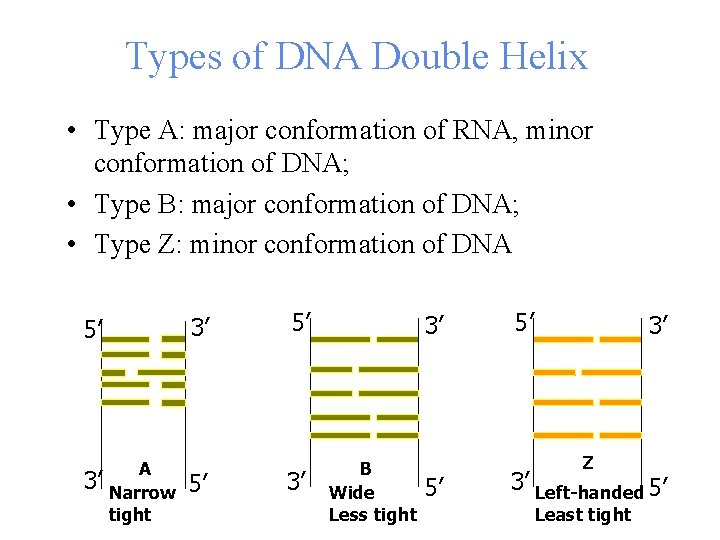
Types of DNA Double Helix • Type A: major conformation of RNA, minor conformation of DNA; • Type B: major conformation of DNA; • Type Z: minor conformation of DNA 3’ 5’ 3’ A Narrow tight 5’ 5’ 3’ 3’ B Wide Less tight 5’ 5’ 3’ Z 3’ Left-handed 5’ Least tight
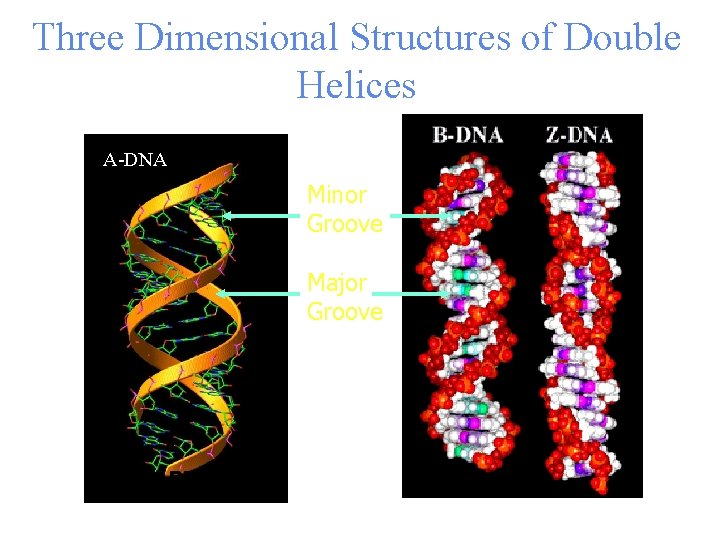
Three Dimensional Structures of Double Helices A-DNA Minor Groove Major Groove A-RNA
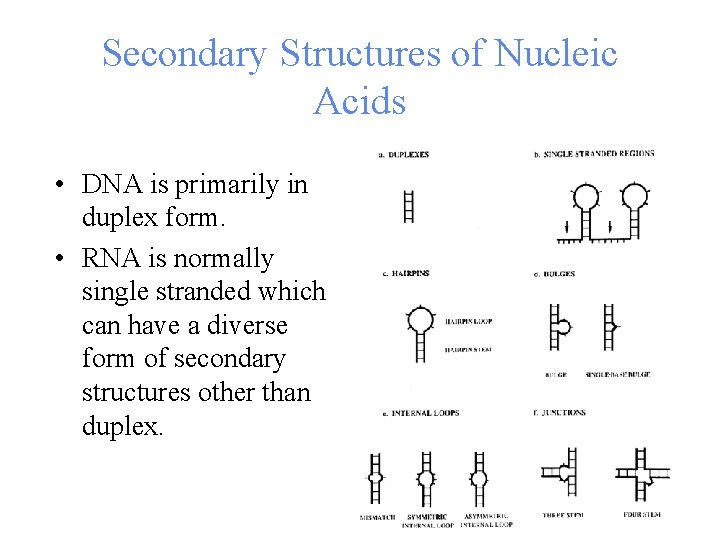
Secondary Structures of Nucleic Acids • DNA is primarily in duplex form. • RNA is normally single stranded which can have a diverse form of secondary structures other than duplex.

More Secondary Structures of Nucleic Acids Pseudoknots: Source: Cornelis W. A. Pleij in Gesteland, R. F. and Atkins, J. F. (1993) THE RNA WORLD. Cold Spring Harbor Laboratory Press.
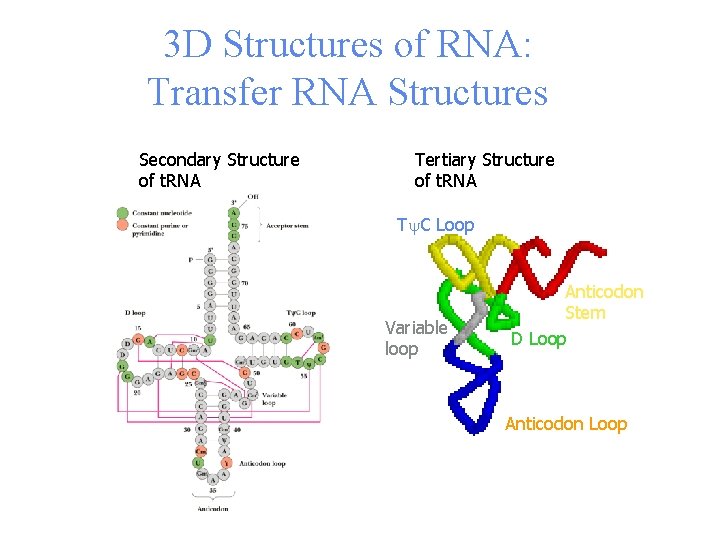
3 D Structures of RNA: Transfer RNA Structures Secondary Structure of t. RNA Tertiary Structure of t. RNA Ty. C Loop Variable loop Anticodon Stem D Loop Anticodon Loop
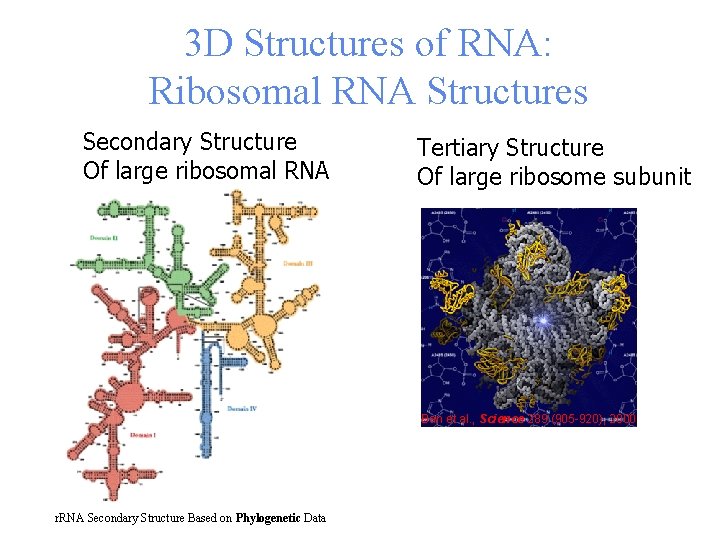
3 D Structures of RNA: Ribosomal RNA Structures Secondary Structure Of large ribosomal RNA Tertiary Structure Of large ribosome subunit Ban et al. , Science 289 (905 -920), 2000 r. RNA Secondary Structure Based on Phylogenetic Data

DNA Sequencing n Chain Termination Method – Sanger, 1977 – single stranded DNA, ~800 b – Method: • Electrophoresis can separate DNA molecules differing 1 bp in length • Dideoxynucleotide (dd. NTP) are used - which stop replication
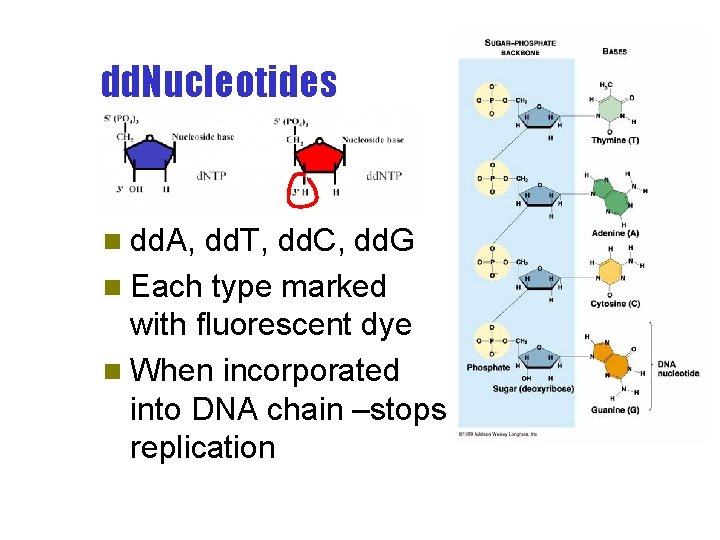
dd. Nucleotides n dd. A, dd. T, dd. C, dd. G n Each type marked with fluorescent dye n When incorporated into DNA chain –stops replication

Chain Termination Method, An Outline n Replication – Obtaining ss. DNA – Add a (universal) primer n Start replication in a soup of A, T, C, G n Continously add tiny amounts of dd. A, dd. T, dd. C, dd. G – gradually stopping all the processes

Chain Termination Method, Reading the Sequence n Running through electrophoresis gel – Four types of dd. NTP have four different fluorescent labels – Automated reading n See: http: //www. dnalc. org/Shockwave/cycseq. html
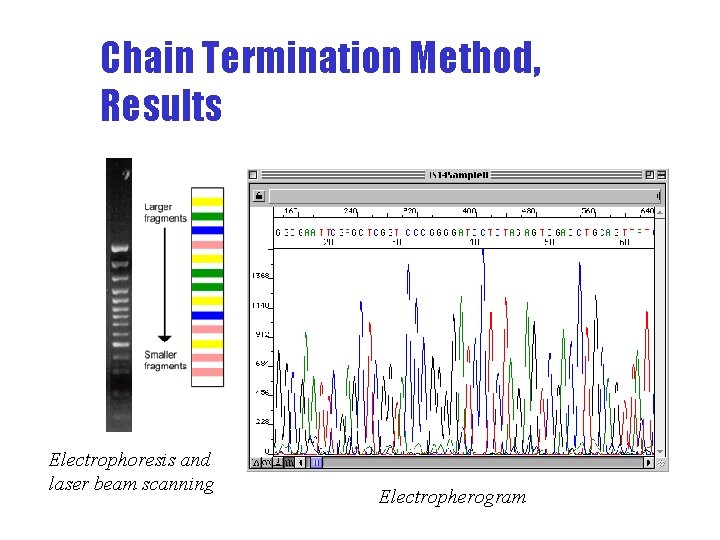
Signal Chain Termination Method, Results Electrophoresis and laser beam scanning time Electropherogram fragment size

Shotgun Method - Overview Cut genome into short fragments n Sequence DNA fragments n Create contigs Contig - continous set of overlapping sequences n Gap!
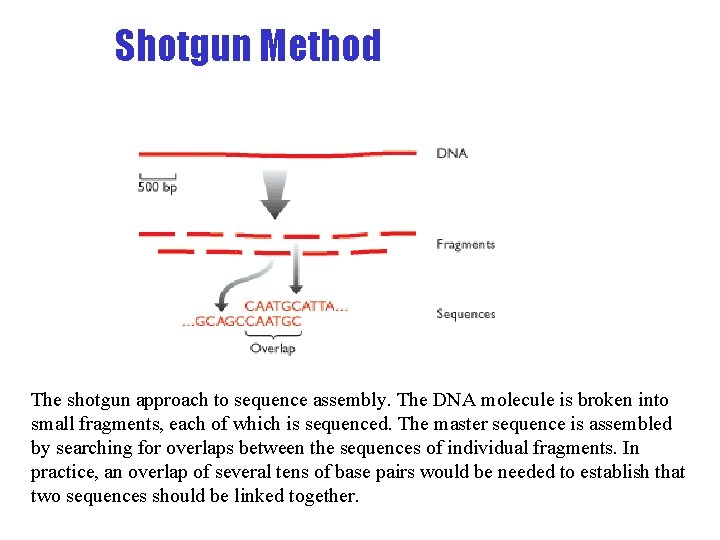
Shotgun Method The shotgun approach to sequence assembly. The DNA molecule is broken into small fragments, each of which is sequenced. The master sequence is assembled by searching for overlaps between the sequences of individual fragments. In practice, an overlap of several tens of base pairs would be needed to establish that two sequences should be linked together.

Shotgun Method – Contig Construction n Two DNA sequences: X=CTATCA Y=AGTAT n How do they overlap? X n Y or Y X Try to apply dynamic programming

Shotgun Method – Contig Construction by Dynamic Programming 2 1
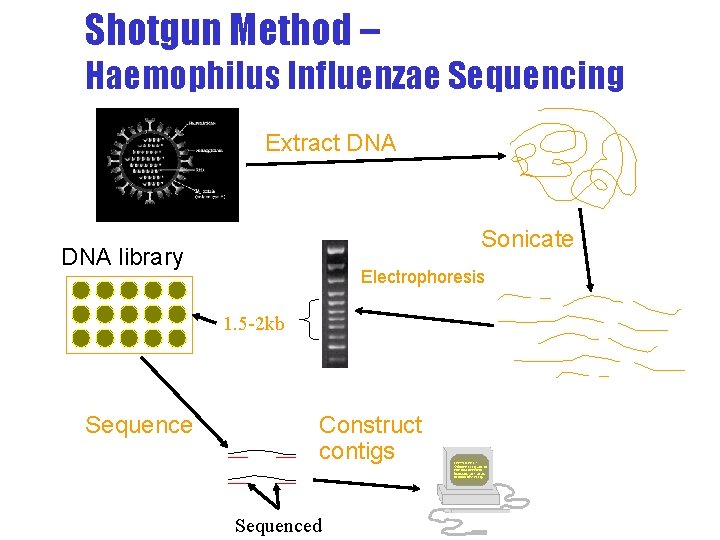
Shotgun Method – Haemophilus Influenzae Sequencing Extract DNA Sonicate DNA library Electrophoresis 1. 5 -2 kb Sequence Construct contigs Sequenced
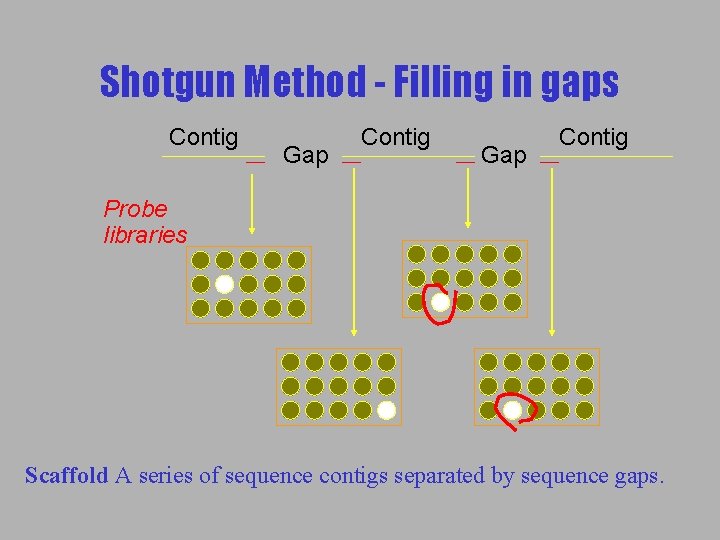
Shotgun Method - Filling in gaps Contig Gap Contig Probe libraries Scaffold A series of sequence contigs separated by sequence gaps.

Shotgun Method - Pros and Cons n Pros – Human labour reduced to minimum n Cons – Computationally demanding – O(n 2) comparisons – High error rate in contig construction • Repeats as the main problem
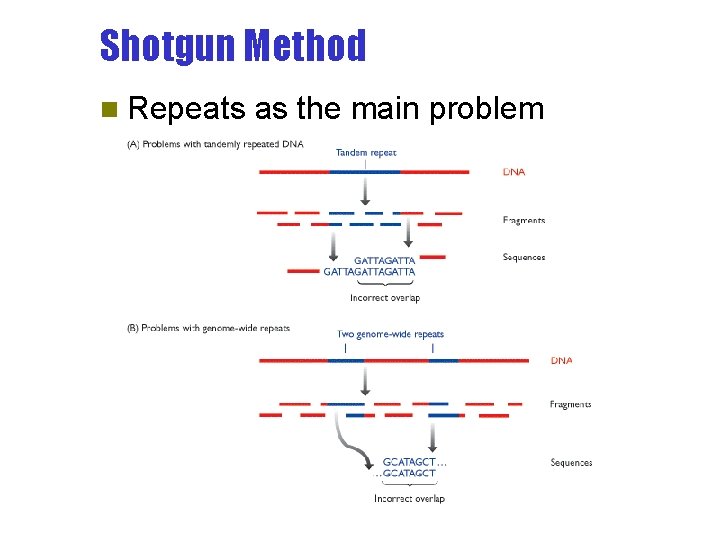
Shotgun Method n Repeats as the main problem

Shotgun vs. Hierarchical Method n Celera vs. Human Genome Project n Hierarchical (top-down) assembly: – The genome is carefully mapped – “Shotgun” into large chunks of 150 kb • Exact location of each chunk is known – Each piece is again “shotgunned” into 2 kb and sequenced

Shotgun vs. Hierarchical Method n Shotgun bottom-up n Hierarchical top-down

New Sequencing Methods n Sequencing By Hybridization – Check which from all possible fragments of length k (k-tuples) hybridize to the sequence TAA AAG AGC ATTCG TAA AAG AGC

Wrapping up Nucleotide, DNA, RNA basics (sequence, structure) n DNA Sequencing n – – Sanger method Shotgun sequencing Hierarchical assembly Contigs, scaffolds, Dynamic Programming
- Slides: 29