Introduction to Bioinformatics Dr Lokesh Gambhir Department Of

Introduction to Bioinformatics Dr. Lokesh Gambhir Department Of Life Sciences Shri Guru Ram Rai Institute of Technology & Sciences (SGRRITS)

What is bioinformatics? There are many different answers to this. One basic definition is that it is the use of computational methods to analyse biological data. By the end of this course, you will • have knowledge of the many data resources available at the NCBI and EBI, • understand some of the basic principles behind aligning sequences, • understand some key points about different sequence alignment programs, • have experience running some web-based bioinformatics programs, • understand the information returned by some sequence database searching programs, • appreciate some of the practical approaches available for automating bioinformatics. Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 2

Databases There are many freely available data resources. A large number are hosted by large national and international institutions such as the American center, the National Center for Biotechnology Information (NCBI) and the European Bioinformatics Centre (EBI). Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 3
Few Concepts to remember DNA Protein/Structure Pattern recognition Folding problem and structure prediction The Twilight Zone Orthologs Paraolgs DNA sequencing Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun ? 4

What can be discovered about a gene by a database search? • A little or a lot, depending on the gene – Evolutionary information: homologous genes, taxonomic distributions, allele frequencies, synteny, etc. – Genomic information: chromosomal location, introns, UTRs, regulatory regions, shared domains, etc. – Structural information: associated protein structures, fold types, structural domains – Expression information: expression specific to particular tissues, developmental stages, phenotypes, diseases, etc. – Functional information: enzymatic/molecular function, pathway/cellular role, localization, role in diseases Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 5

Searching sequence databases • Start from sequence, find information about it • Many kinds of input sequences – Could be amino acid or nucleotide sequence – Genomic or m. RNA/c. DNA or protein sequence – Complete or fragmentary sequences • Exact matches are rare (even uninteresting in many cases), so often goal is to retrieve a set of similar sequences. – Both small (mutations) and large (required for function) differences within “similar” can be interesting. Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 6

What might we want to know about a sequence? • Is this sequence similar to any known genes? How close is the best match? Significance? • What do we know about that gene? – Genomic (chromosomal location, allelic information, regulatory regions, etc. ) – Structural (known structure? structural domains? etc. ) – Functional (molecular, cellular & disease) • Evolutionary information: – Is this gene found in other organisms? – What is its taxonomic tree? Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 7
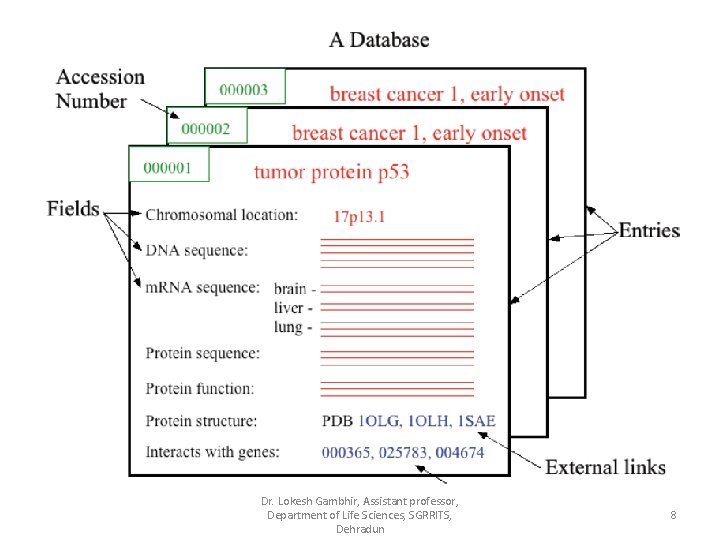
Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 8
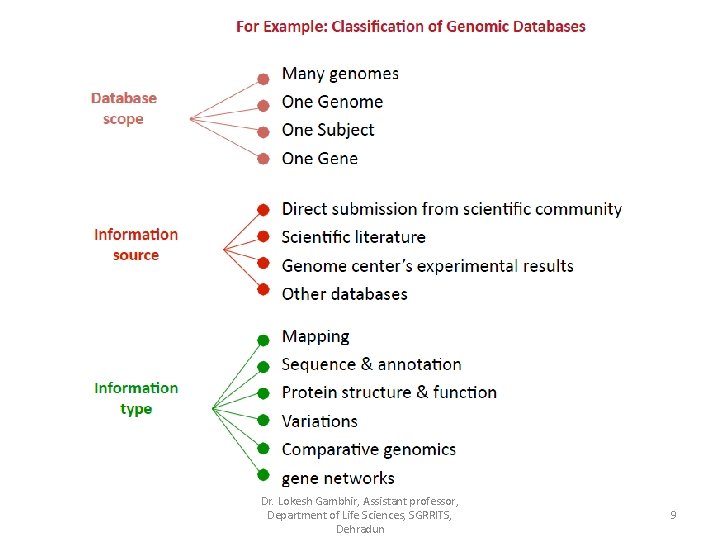
Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 9
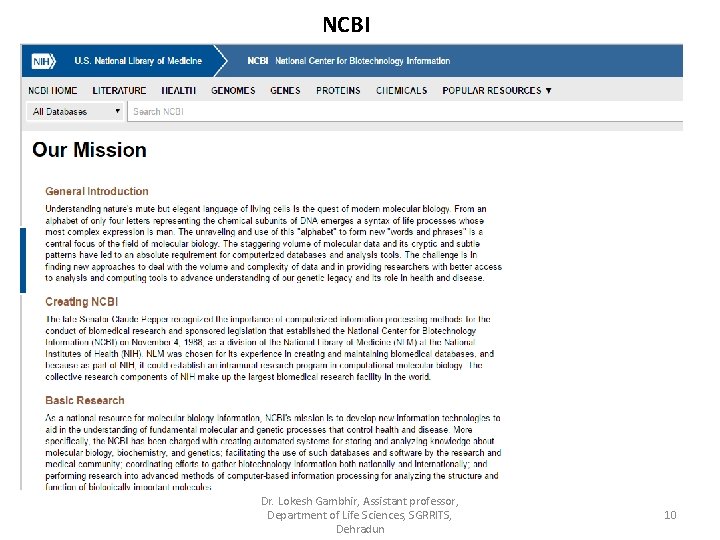
NCBI Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 10

To carry out its diverse responsibilities, NCBI: • Conducts research on fundamental biomedical problems at the molecular level using mathematical and computational methods. • Maintains collaborations with several NIH institutes, academia, industry, and other governmental agencies • Fosters scientific communication by sponsoring meetings, workshops, and lecture series • Supports training on basic and applied research in computational biology for postdoctoral fellows through the NIH Intramural Research Program • Engages members of the international scientific community in informatics research and training through the Scientific Visitors Program • Develops, distributes, supports, and coordinates access to a variety of databases and software for the scientific and medical communities • Develops and promotes standards for databases, data deposition and exchange, and biological nomenclature Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 11

Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 12

Entrez The Entrez Global Query Cross-Database Search System is a federated search engine, or web portal that allows users to search many discrete health sciences databases at the National Center for Biotechnology Information (NCBI) website. Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 13
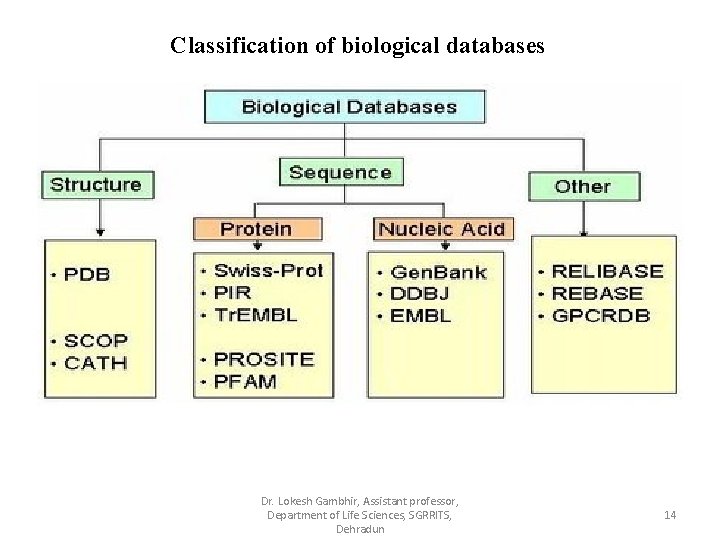
Classification of biological databases Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 14

Characteristics of entries in the primary nucleotide repositories • The large nucleotide databases are not hand-curated: the quality of the information is largely dependent on the people submitting the sequence. • Records can be updated by the original submitter, or by a third party if the submitter granted them permission and notified the relevant institute (not common). • There are redudant entries in these databases. • Entries can contradict one another. • Predicted or known proteins coded for by the sequence are linked to via their accession number in the Uniprot knowledgebase. • Information from any species, including sequences of unknown origin, can be deposited in the database. Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 15

Gen. Bank® is the NIH genetic sequence database, an annotated collection of all publicly available DNA sequences. • Collaboration between NCBI (National Center for Biotechnology Information), EMBL (The European Molecular Biology Laboratory ), EBI (European Bioinformatics Institute), DDBJ (DNA Data Bank of Japan). Each record in Gen. Bank is in a “Gen. Bank flat file format”. • Each record contains information about a sequence type (DNA/protein/RNA……) • source/organism, reference, …… • features • functions of a region on the sequence • The sequence Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 16
Gen. Bank http: //www. ncbi. nlm. nih. gov/genbank/ Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 17

Flat File Format of Gen. Bank Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 18
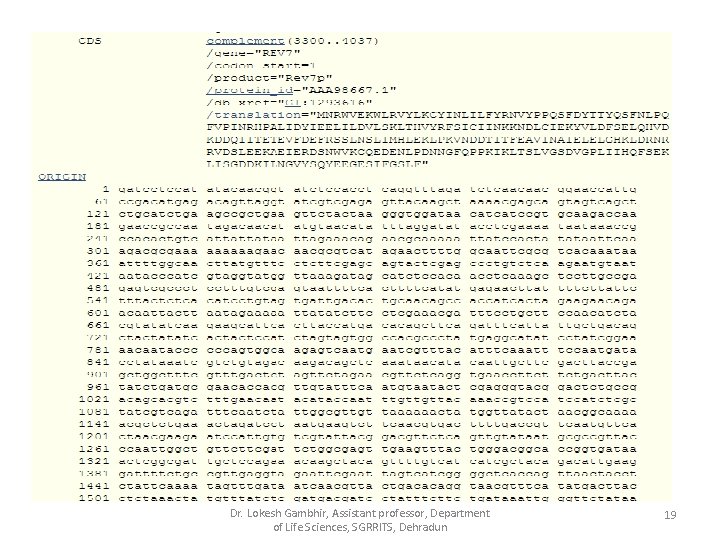
Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 19

Abraxane is a chemotherapeutic drug. How will you determine the molecular target of the drug ? Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 20

EMBL-EBI • The roots of the EMBL-EBI lie in the world's first nucleotide sequence database • The EMBL Nucleotide Sequence Data Library (now EMBL Bank, part of the European Nucleotide Archive), which was established in 1980 at the European Molecular Biology Laboratory in Heidelberg, Germany. • The original goal was to establish a central database of DNA sequences, rather than have scientists submit sequences to journals. • Data retrieval is done by employing SRS which connects the primary DNA-Protein databases along with secondary and specialised database • MEDLINE is used for reference application Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 21

Uni. Prot/SWISS-Prot The mission of Uni. Prot is to provide the scientific community with a comprehensive, high quality and freely accessible resource of protein sequence and functional information. Uni. Prot is comprised of four components, each optimised for different uses: ØThe Uni. Prot Knowledgebase (Uni. Prot. KB) is the central access point for extensive curated protein information, including function, classification, and crossreference. Uni. Prot. KB comprises two sections: • Uni. Prot. KB/Swiss-Prot which is manually annotated and is reviewed • Uni. Prot. KB/Tr. EMBL which is automatically annotated and is not reviewed. ØThe Uni. Prot Reference Clusters (Uni. Ref) databases provide clustered sets of sequences from the Uni. Prot. KB and selected Uni. Prot Archive records to obtain complete coverage of sequence space at several resolutions while hiding redundant sequences. ØThe Uni. Prot Archive (Uni. Parc) is a comprehensive repository, used to keep track of sequences and their identifiers. Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 22
Uni. Prot/SWISS-Prot Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 23
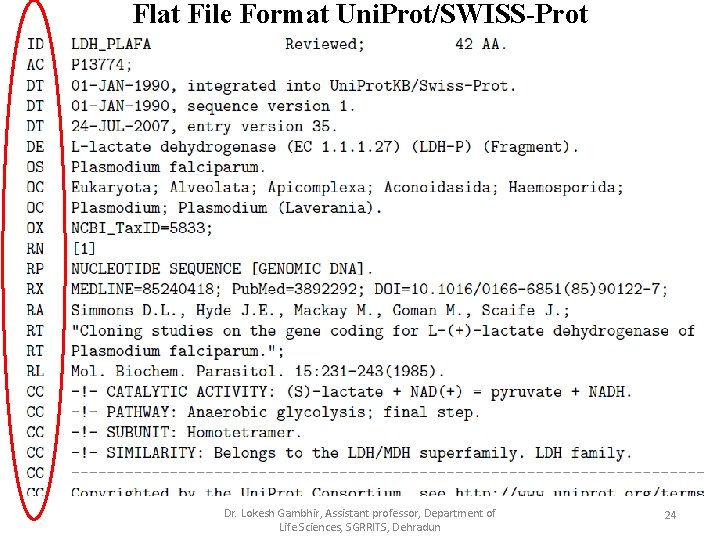
Flat File Format Uni. Prot/SWISS-Prot Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 24

Dr. Lokesh Gambhir, Assistant professor, Department of Life Sciences, SGRRITS, Dehradun 25
- Slides: 25