Introduction Science of genetics can be divided into

Introduction Science of genetics can be divided into 4 subdisciplines: • Transmission genetics: how are genes passed from one generation to the next? Study object: individual • Molecular genetics: how relates DNA to phenotype? Study object: cell • Population genetics: how behave traits in populations? Study object: population • Quantitative genetics: what are rules for polygenic inheritance? Study object: population In this course we will study • Complex transmission genetics and we will use statistical methods for the analysis of genetic experiments • Population and quantitative genetics • Extra-nuclear genetics Laboratoriumtechniek

Laws of Probability 1. Rule of multiplication = The probability that independent events will occur simultaneously is the product of their individual probabilities 2. Rule of addition = The probability of an event that can occur in two or more independent ways is the sum of the separate probabilities of the different ways Question: In a Mendelian cross between pea plants that are heterozygous for flower color (Pp), what is the probability that the offspring will be homozygous recessive? Question: In a Mendelian cross between pea plants that are heterozygous for flower color (Pp), what is the probability of the offspring being a heterozygote? Laboratoriumtechniek

Laws of Probability 3. Conditional probability The probability of an outcome that depends on a specific condition related to that outcome Question: In the F 2 of a monohybrid cross with plants with purple and white flowers: what is the probability that a plant with purple flowers is heterozygous? Pa probability of being heterozygous = 1/2 Pb probability of the condition (purple) = 3/4 Pc = Pa/Pb = 2/3 Laboratoriumtechniek

Laws of Probability 4. The binomial theorem • The probability of two alternative outcomes during a number of trials. Question: In a family of four, what is the probability of having two boys and two girls? N Binomial Expanded binomial 1 2 3 4 5 (x + y)1 = 1 x + 1 y (x + y)2 = 1 x 2 + 2 xy + 1 y 2 (x + y)3 = 1 x 3 + 3 x 2 y + 3 xy 2 + 1 y 3 (x + y)4 = 1 x 4 + 4 x 3 y + 6 x 2 y 2 + 4 xy 3 + 1 y 4 (x + y)5 = 1 x 5 y 0 + 5 x 4 y 1 + 10 x 3 y 2 + 10 x 2 y 3 + 5 x 1 y 4 + 1 x 0 y 5 N=4, x=0. 5, y=0. 5 P(4 boys)= 1 x 4 = 1/16 P(2 boys and 2 girls)= 6 x 2 y 2 = 6/16 Laboratoriumtechniek

Laws of Probability • The general formula for the binomial expansion is: Probability = N! ·(px)(q. N-x) x!(N-x)! N=total number of events p is the probability of event P q=probability of the alternative event x is the number of times event P occurs ! means factorial. The total number of events (N) is 4 children. The number of times (x) that event P (male child) occurs is 2. The probabilities p and q are both 0. 5. Thus: 4! ·(0. 52) = 24/4× 1/16 =6/16=3/8 (2!) Of all families with 4 children 3 out of 8 are predicted to have two boys and two girls. Laboratoriumtechniek

Testing hypotheses using the 2 -statistic Mendel raised 705 purple-flowered and 224 white-flowered plants in the F 2 generation. Hypothesis: Is this a ratio of 3: 1? 1. Calculate expected values based on the result total (929): 3 : 1 = 696. 75 : 232. 25 2. Calculate the differences between observed (O) and expected (E) 3. Standardise the differences: |O – E |2 2 = E 4. Calculate degrees of freedom (DF) = number of differences – 1 5. Lookup in 2 table at DF 6. If p<0. 05 then reject hypothesis O E |O–E| 705 696. 75 8. 25 224 232. 25 8. 25 929 (8. 25 -0. 5)2 2 = + 696. 75 232. 25 The correction for continuity of 0. 5 is only applied when DF=1 2 = 0. 086 + 0. 259 = 0. 345 At DF=1 0. 345 lies between 0. 016 and 0. 455 with 0. 900>p>0. 500, so accept hypothesis Laboratoriumtechniek

Testing hypotheses using the 2 -statistic Mendel raised the F 2 generation of a dihybrid cross and found yellow/round 315, green/round 108, yellow/wrinkled 101 and green/wrinkled 32. Hypothesis: Is this a ratio of 9: 3: 3: 1? • class round/yellow round/green wrinkled/yellow wrinkled/green Total observed 315 108 101 32 556 expected 312. 75 104. 25 34. 75 556. 00 deviation +2. 25 +3. 75 -3. 25 -2. 75 +0. 00 • Standardise the differences: |O – E |2 2 = E = 0. 470 • Calculate degrees of freedom (DF) = number of differences – 1 = 3 • Lookup in 2 table at DF: P>0. 9 • If P<0. 05 then reject hypothesis. Laboratoriumtechniek
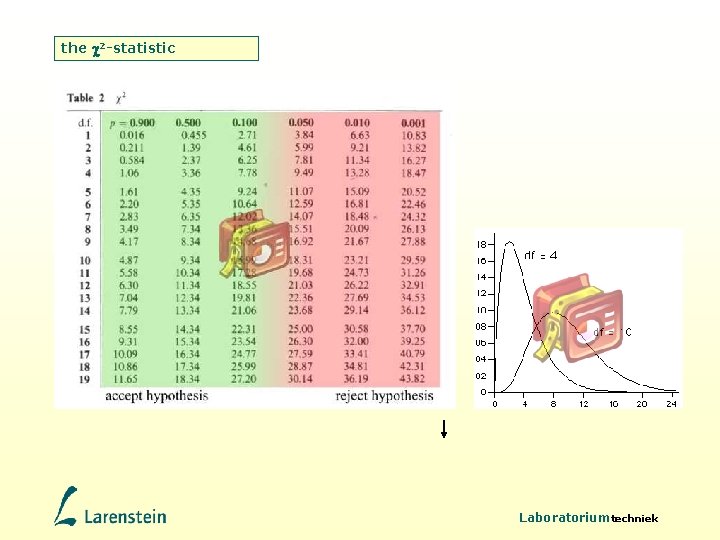
the 2 -statistic Laboratoriumtechniek

Population genetics Questions studied in Population genetics: • how much genetic variation is found in natural populations and what processes control the amount of variation? • What processes are responsible for producing genetic divergence between populations? • How do population characteristics like breeding system, fecundity, and age structure influence the gene pool? Laboratoriumtechniek

Genetic structure of populations To study the genetic structure of a Mendelian population one must describe the gene pool by calculating the genotypic and allelic frequencies • Genotypic frequencies tiger moth: f(BB)= 452/497=0. 909 (top 2) f(Bb)= 43/497= 0. 087 (middle 4) f(bb)= 2/497= 0. 004 (bottom 1) • Allelic frequencies: tiger moth: p=f(B)=(2× 452+43)/(2× 497)=0. 953 q=f(b)=(2× 2+43)/(2× 497)=0. 047 or p=f(B)=f(BB)+½f(Bb)=0. 953 q=f(b)=f(bb)+½f(Bb)=0. 047 Laboratoriumtechniek

Genetic structure of populations Allelic frequencies with multiple alleles A 1, A 2 and A 3 (2×count of A 1 A 1)+(A 1 A 2)+(A 1 A 3) p=f(A 1)= (2×total number of individuals) (2×count of A 2 A 2)+(A 2 A 1)+(A 2 A 3) q=f(A 2)= (2×total number of individuals) (2×count of A 3 A 3)+(A 3 A 1)+(A 3 A 2) r=f(A 3)= (2×total number of individuals) Laboratoriumtechniek

Genetic structure of populations Allelic frequencies at an X-linked locus: (2×XAXA females)+(XA Xa females)+(XA Y males) p=f(XA)= (2×number of females)+(number of males) (2×Xa. Xa females)+(XA Xa females)+(Xa Y males) q=f(Xa)= (2×number of females)+(number of males) If the numbers of males and females are equal: 2 1 1 3 2 3 p=f(XA)= — [f(XAXA) + — f(XAXa)] + — f(XAY) q=f(Xa)= — [f(Xa. Xa) + — f(Xa. XA)] + — f(Xa. Y) Laboratoriumtechniek

The Hardy-Weinberg Law says that: • In an infinitely large, randomly mating population, free from mutation, migration, and natural selection, • the frequencies of the alleles do noy change over time, • genotypic frequencies remain in the proportions p 2 (frequency of AA), 2 pq (frequency of Aa) and q 2 (frequency of aa), p 2 + 2 pq + q 2 = 1 Assumptions: If population size is limited: chance deviations cause genetic drift Random mating not necessary for all traits: humans mate preferentially for height, IQ, skin colour, etc. but random for blood type. Laboratoriumtechniek

The Hardy-Weinberg Law Female gametes When is a population in genetic equilibrium? 1. The allele frequencies will not change from generation to generation 2. The genotypic frequencies will be in the proportions p 2, 2 pq and q 2 after one generation of random mating Male gametes A(p) a(q) A(p) AA(p 2) Aa(pq) a(q) Aa(pq) aa(q 2) Why? Each generation of zygotes produces A and a in proportions p and q. Female gametes If p=0. 6 and q=0. 4 then AA=0. 36 (p 2), Aa=2× 0. 24 (2 pq), and aa=0. 16 (q 2) Total=1. 00 Male gametes A(0. 6) a(0. 4) A(0. 6) AA(0. 36) Aa(0. 24) a(0. 4) Aa(0. 24) aa(0. 16) Laboratoriumtechniek 14
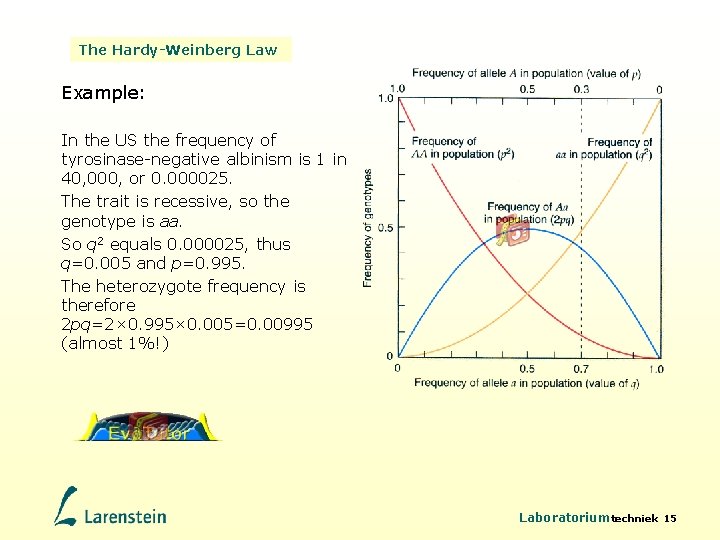
The Hardy-Weinberg Law Example: In the US the frequency of tyrosinase-negative albinism is 1 in 40, 000, or 0. 000025. The trait is recessive, so the genotype is aa. So q 2 equals 0. 000025, thus q=0. 005 and p=0. 995. The heterozygote frequency is therefore 2 pq=2× 0. 995× 0. 005=0. 00995 (almost 1%!) Laboratoriumtechniek 15

The Hardy-Weinberg Law Extension for more than 2 alleles. • 2 alleles: p 2 + 2 pq + q 2 = (p + q)2 • 3 alleles: (p + q + r)2 = p 2 + q 2 + r 2 + 2 pq + 2 pr + 2 qr • 4 alleles: (p + q + r + s)2 • etc. Example: allozym-polymorphism in Mytilus edulis Allele frequency LAP 98 LAP 96 LAP 94 p=0. 52 q=0. 31 r=0. 17 LAP=leucine aminopeptidase Assignment: calculate the genotypic frequencies Laboratoriumtechniek

The Hardy-Weinberg Law Testing for Hardy-Weinberg equilibrium using 2 Example tiger moth: observed genotypic frequencies: f(BB)= 452/497=0. 909 f(Bb)= 43/497= 0. 087 f(bb)= 2/497= 0. 004 Allelic frequencies: p=f(B)=0. 953 q=f(b)=0. 047 expected f(BB)= p 2 × N = f(Bb)= 2 pq × N = f(bb)= q 2 × N = observed |O–E| 451. 4 452 0. 6 44. 5 43 1. 5 2 0. 9 1. 1 The number of degrees of freedom is 1, because p and q are not independent, so the correction for continuity is applied 2 (1. 5 -0. 5)2 (0. 9 -0. 5)2 (0. 6 -0. 5) 2 = + + = 0. 168 451. 4 44. 5 1. 1 The change P for this result is >0. 5, so there is no significant deviation from the Hardy-Weinberg equilibrium. Laboratoriumtechniek 17

The Hardy-Weinberg Law can be used to estimate allelic frequencies Example: In Arizona (US) live the Hopi Indians. In this tribe 26 cases of albinism were observed in a population of 6000. So q 2 = 26/6000 = 0. 0043 and q= 0. 065. Therefore p=0. 935. The frequency of heterozygotes is 2 pq = 2× 0. 935× 0. 065 = 0. 122, thus one out of eight Hopi’s carries an allele for albinism. Photograph taken in 1900 Is this calculation allowed? Laboratoriumtechniek 18
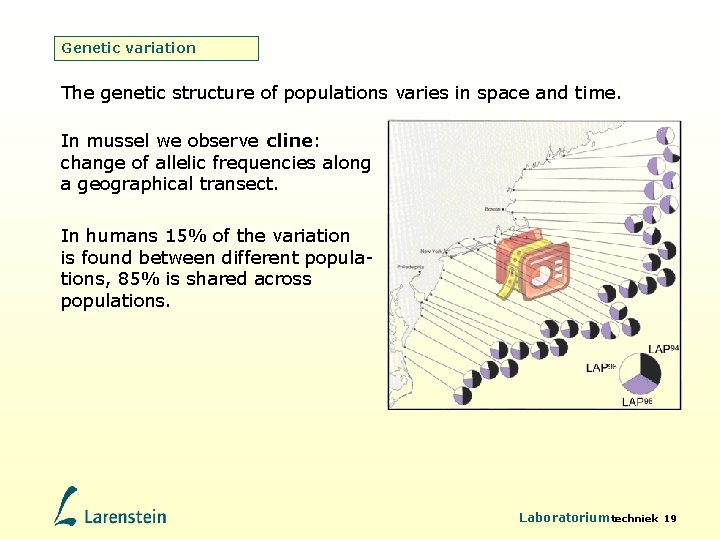
Genetic variation The genetic structure of populations varies in space and time. In mussel we observe cline: change of allelic frequencies along a geographical transect. In humans 15% of the variation is found between different populations, 85% is shared across populations. Laboratoriumtechniek 19

Genetic variation How is genetic variation measured? • The genetic basis of most traits is too complex to assign genotypes to individuals. Only a few traits behave in a Mendelian fashion. • Cytological variation (chromosome morphology and banding pattern) was observed in salivary glands of fruit flies. • Use of starch gel electrophoresis to study protein polymorphism made it possible to determine genotypes of many individuals at many loci. • By measuring variation at the DNA- or RNA-level. Laboratoriumtechniek
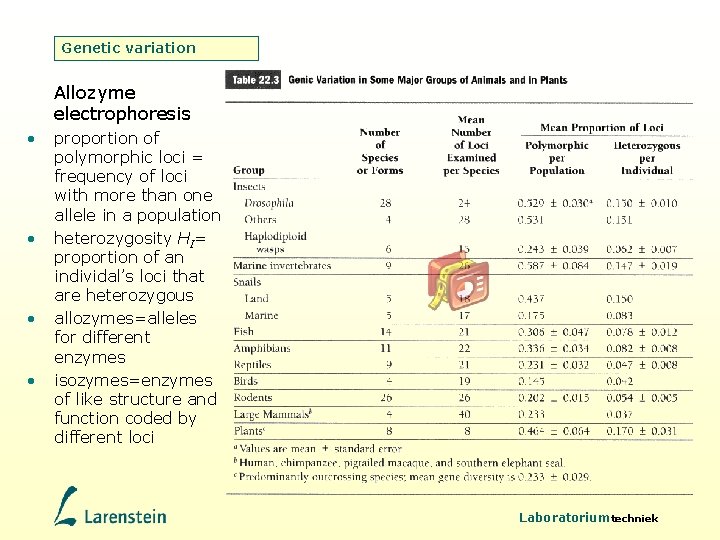
Genetic variation Allozyme electrophoresis • • proportion of polymorphic loci = frequency of loci with more than one allele in a population heterozygosity HI= proportion of an individal’s loci that are heterozygous allozymes=alleles for different enzymes isozymes=enzymes of like structure and function coded by different loci Laboratoriumtechniek
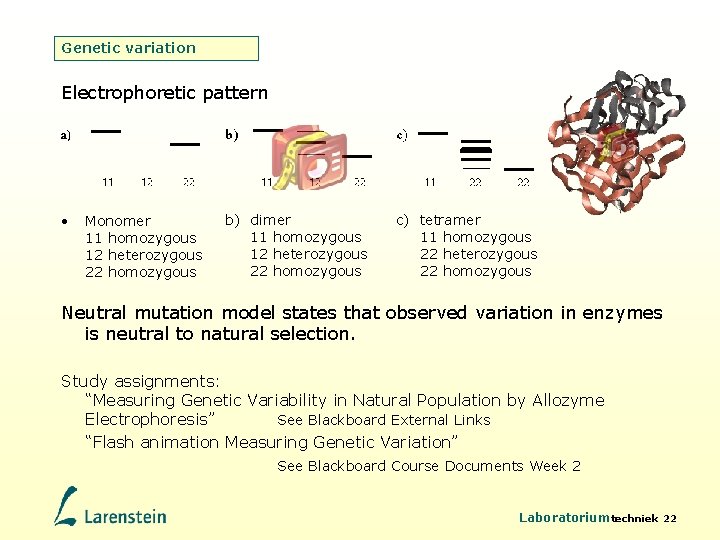
Genetic variation Electrophoretic pattern • Monomer 11 homozygous 12 heterozygous 22 homozygous b) dimer 11 homozygous 12 heterozygous 22 homozygous c) tetramer 11 homozygous 22 heterozygous 22 homozygous Neutral mutation model states that observed variation in enzymes is neutral to natural selection. Study assignments: “Measuring Genetic Variability in Natural Population by Allozyme Electrophoresis” See Blackboard External Links “Flash animation Measuring Genetic Variation” See Blackboard Course Documents Week 2 Laboratoriumtechniek 22
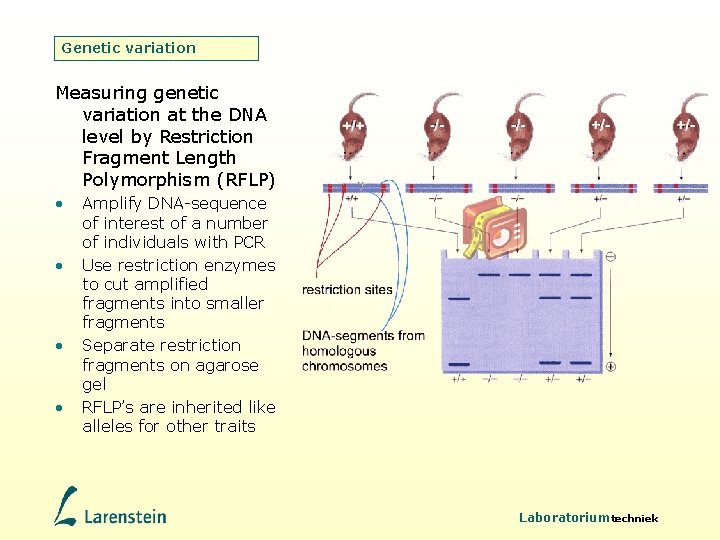
Genetic variation Measuring genetic variation at the DNA level by Restriction Fragment Length Polymorphism (RFLP) • • Amplify DNA-sequence of interest of a number of individuals with PCR Use restriction enzymes to cut amplified fragments into smaller fragments Separate restriction fragments on agarose gel RFLP’s are inherited like alleles for other traits Laboratoriumtechniek

Genetic variation • Individual nucleotide heterozygosity HI in eukaryotes varies from 0. 001 to 0. 02 • In the human genome nucleotide heterozygosity HI is estimated 0. 0008, which means that an individual is heterozygous at about 1 in every 1250 nucleotides • The highest diversity occurs at sites that do not change the amino acid sequence, which are called synonymous changes • Non-synonymous changes may affect a protein’s function and are subjected to natural selection and have been eleminated from the population • In many regions of the DNA there are short, identical segments, called Short Tandem Repeats (STR) or microsatellites, the number of which varies from person to person. They are used in forensics, parent testing, etc. Exercises: 22. 1 -22. 14 with a * Laboratoriumtechniek

Forces that change gene frequencies • The simplest way to calculate the population heterozygosity HT for a single locus is as: HT = 1 – pi 2 i where pi is the frequency of the i-th of k alleles. [Note that p 1, p 2, p 3 etc. may correspond to what you would normally think of as p, q, r, s etc. ]. • If we want the gene diversity over several loci we need double summation and subscripting as follows: HT = 1 – pij 2 i j • 2 alleles: if p=q=0, 5 then HT = 1–(0, 25+0, 25) = 0, 5 • 10 alleles: if p 1=p 2=p 3 … p 10=0, 1 then HT = 1–(10× 0, 01) = 0, 9 Laboratoriumtechniek 25

Forces that change gene frequencies For many populations the conditions for Hardy-Weinberg equilibrium do not hold, because: • Mutation occurs • They are small • Natural selection favours/dismisses alleles • Mating may be non-random • Migration takes place Laboratoriumtechniek 26

Forces that change gene frequencies Mutations are heritable random changes in the DNA of a locus • New alleles occur only as a result of mutation • Mutations provide raw genetic material on which natural selection acts • Kinds of mutations: - base substitutions: Original Nucleotide Base transitions A «» G and C «» T are substitutions A T C more common than transversions A 4. 4 6. 5 Mutant A «» T, G «» T, C «» G and A «» C Nucleotide T 4. 7 21. 0 - Deletions and insertions C 5. 0 8. 2 - Chromosomal mutations G 9. 4 3. 3 4. 2 (inversions and translocations) see Flash animation Translocations on Blackboard G 20. 7 7. 2 5. 3 - Laboratoriumtechniek 27

Forces that change gene frequencies Estimates of mutation rates • Using sequence analysis – 1/1. 000 mutations per base pair per cell division – Varies 1000 -fold within genome: high at mutational hotspots • Usingle base mutations with phenotypic effect – 1/100. 000 per gamete (humans, mice, Drosophila) • By keeping DNA non-expressed for many generations – Mutation rate = 1 per zygote, mostly mildly deleterious and recessive • Using amino acid chang – 4. 2 amino acid altering mutations per ind. per generation – 1. 6 of these are deleterious • Mutation rates in males > females Laboratoriumtechniek

Forces that change gene frequencies • A mutation can be neutral, detrimental, or beneficial, which depends on the specific environment e. g. DDT-resistance in insects • Forward mutation rate u is higher than reverse mutation rate v • Forward mutation from A to a lowers p and the reverse mutation form a to A increases p. At equilibrium: v u p = and q = u + v Laboratoriumtechniek

Forces that change gene frequencies • How fast do mutations change allele frequencies? Example: p=0. 9 and q=0. 1 u=5× 10– 5 and v=2× 10– 5 After one generation: p=vq–up=(2× 10– 5× 0. 1)–(5× 10– 5× 0. 9)=– 0. 000043 At equilibrium p=0. 286 This means that a change from 0. 50 to 0. 49 takes 680 generations A change from 0. 30 to 0. 29 takes 17, 200 generations! Laboratoriumtechniek 30

Forces that change gene frequencies Random genetic drift Laboratoriumtechniek 31
Forces that change gene frequencies Random genetic drift Laboratoriumtechniek 32

Forces that change gene frequencies Random genetic drift is caused by sampling error when gametes are drawn randomly from a small population. Example: A small population with 12 individuals p=0. 5 and q=0. 5 has genotypes 3 AA, 6 Aa and 3 aa. Suppose an accident kills 25% of the population, which all happen to be aa. This chance is. . . . In the next generation p becomes. . . and q becomes. . . . Laboratoriumtechniek 33

Forces that change gene frequencies Bottlenecks and founder effects • When populations remain small over many generations drift plays an important role in allelic frequencies. Scattered populations in isolated habitats each undergo drift. • Founder effect occurs when a population is established with a small number of breeding individuals. Example: Tristan da Cunha was settled in 1817 by William Glass and wife. A few more settlers joined them. In 1855 population size was 100 with 26% of all genes from the Glasses. In 1961 after a volcanic eruption the whole population of 300 was evacuated to England. 14% of their genes were still Glass’s. Laboratoriumtechniek 34
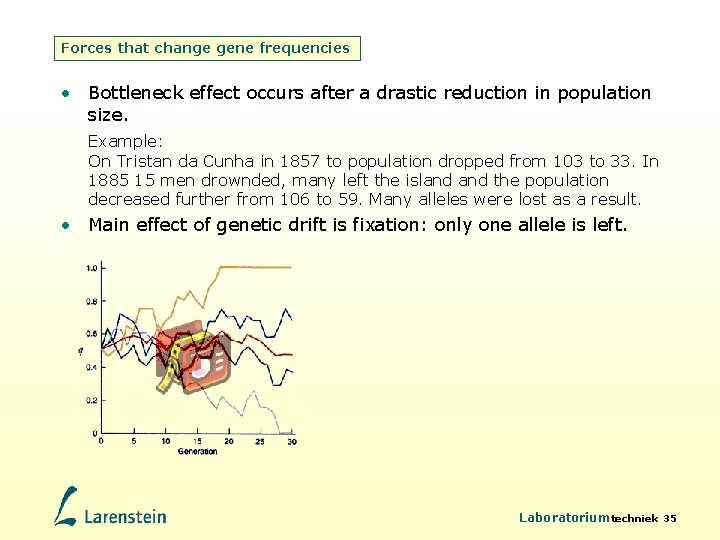
Forces that change gene frequencies • Bottleneck effect occurs after a drastic reduction in population size. Example: On Tristan da Cunha in 1857 to population dropped from 103 to 33. In 1885 15 men drownded, many left the island the population decreased further from 106 to 59. Many alleles were lost as a result. • Main effect of genetic drift is fixation: only one allele is left. Laboratoriumtechniek 35

Forces that change gene frequencies • Genetic drift is random, individual populations do not change in the same direction. This divergence increases over generations. • This is the basis of the Neutral theory of molecular evolution: a new mutation that is neutral with respect to natural selection will most likely be lost as a result of genetic drift. Occasionally the mutation will drift to fixation. As mutation is a recurring event, a gene will accumulate differences over time by chance alone. In this way the genes of two related lineages can be compared and used to estimate the date since they last shared a common ancestor. Following this method humans and chimpanzees shared a common ancestor 6 million years ago. What is your opinion about this method? Laboratoriumtechniek 36

Forces that change gene frequencies • Can we predict Genetic drift? Depends on effective population size = number of individuals that contribute gametes to the next generation 4×Nf×Nm pq pq Ne = , variance = , standard error s= Nf+Nm 2 Ne √ In which Nf=number of females, Nm= number of males Example: In a population of 70 females where only 2 males fertilise all the females, the females each contribute 1/2× 1/70= 0. 007 and each male 1/ × 1/ = 0. 25 of the gametes 2 2 Ne = (4× 70× 2)/(70+2) = 7. 8, which means that drift occurs in about the same rate as in a population with 4 females and 4 males! If p=0. 8 then sp= √pq/2 Ne = 0. 1 Laboratoriumtechniek 37

Forces that change gene frequencies Genghis Khan a Prolific Lover, DNA Data Implies Hillary Mayell for National Geographic News February 14, 2003 Genghis Khan, the fearsome Mongolian warrior of the 13 th century, may have done more than rule the largest empire in the world; according to a recently published genetic study, he may have helped populate it too. An international group of geneticists studying Y-chromosome data have found that nearly 8 percent of the men living in the region of the former Mongol empire carry y-chromosomes that are nearly identical. That translates to 0. 5 percent of the male population in the world, or roughly 16 million descendants living today. Laboratoriumtechniek 38
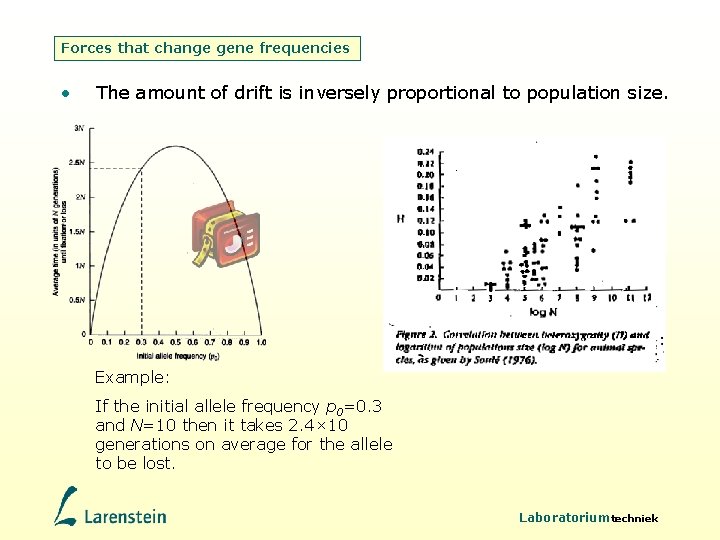
Forces that change gene frequencies • The amount of drift is inversely proportional to population size. Example: If the initial allele frequency p 0=0. 3 and N=10 then it takes 2. 4× 10 generations on average for the allele to be lost. Laboratoriumtechniek

Forces that change gene frequencies • Balance between mutation and random genetic drift. Mutation introduces new variation, genetic drift removes it. • The Infinite alleles model states that every mutation generates a novel allele. In a large gene of 10, 000 base pairs the chance that two mutations generate the same allele is very small. (300 amino acid protein ‡ 900 nucleotides ‡ 4900 = 10542 possible sequences) • In this situation the forces of mutation and genetic drift balance each other. The chance of drawing two alleles and having them be different is 4 Ne×u HT = 1+4 Ne×u HT is equal to the heterozygosity in the total population. Laboratoriumtechniek 40

Forces that change gene frequencies Migration • Immigration introduces new alleles into populations: gene flow • Gene flow changes allelic frequencies in the recipient population • Example: px = f(A)x = 0. 8 in population x and py = f(A)y = 0. 3 in population y In each generation some individuals migrate from x to y. After migration population y consists of two groups of individuals: a proportion m of migrants with px = 0. 8 and 1–m residents with py = 0. 3. The frequency of A in y after migration is p’y = mpx + (1–m)py p= m(px–py) • Through exchange of alleles different populations remain similar, divergence is reduced Example: Monarch butterfly Laboratoriumtechniek 41

Forces that change gene frequencies Natural selection • In contrast to mutation, genetic drift and migration, natural selection results in adaptation. Adaptation is the proces by which traits evolve that make organisms more suited to their environment. • Natural selection is the dominant force in evolution. The concept comes from Darwin (who had no clue about transmission genetics). • Natural selection can be defined as differential reproduction of genotypes. It simply means that individuals with certain alleles produce more offspring than others. Therefore those alleles increase in frequency in the next generation. • Example Industrial melanism, described around 1900. Laboratoriumtechniek 42
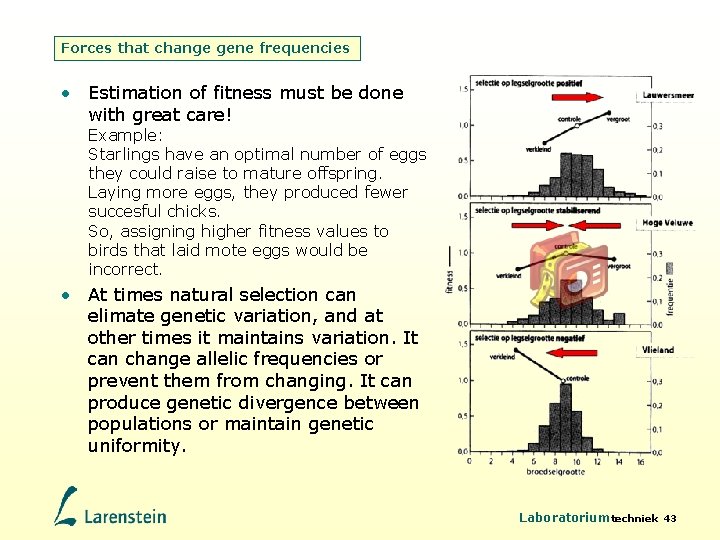
Forces that change gene frequencies • Estimation of fitness must be done with great care! Example: Starlings have an optimal number of eggs they could raise to mature offspring. Laying more eggs, they produced fewer succesful chicks. So, assigning higher fitness values to birds that laid mote eggs would be incorrect. • At times natural selection can elimate genetic variation, and at other times it maintains variation. It can change allelic frequencies or prevent them from changing. It can produce genetic divergence between populations or maintain genetic uniformity. Laboratoriumtechniek 43

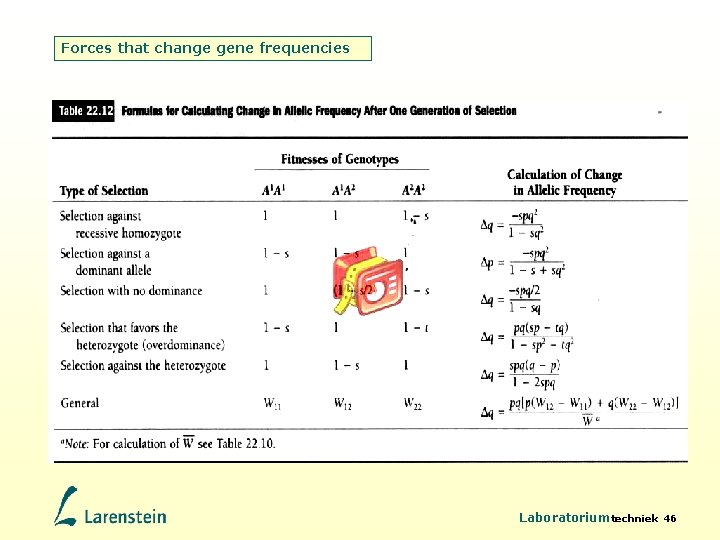
Forces that change gene frequencies • Natural selection is measured in terms of Darwinian fitness W, which is defined as the relative reproductive ability of a genotype. • The genotype that produces the most offspring has fitness W=1. • Selection coefficient s is the relative intensity of selection against a genotype = 1–W Example: Laboratoriumtechniek 44

Forces that change gene frequencies Possible outcomes of natural selection • W 11= W 12= W 22. No selection, allelic frequencies do not change. • W 11= W 12<1 and W 22= 1. Selection operates against the dominant allele. • W 11= W 12= 1 and W 22<1. Selection operates against the recessive allele. • W 11< W 12< 1 and W 22=1. Selection operates without effects of dominance, the heterozygote has intermediate fitness. • W 11 and W 22<1 and W 12= 1. Selection is favouring the heterozygote. • W 12< W 11 and W 22= 1. Selection operates is favouring both homozygotes. Laboratoriumtechniek 45
Forces that change gene frequencies Laboratoriumtechniek 46
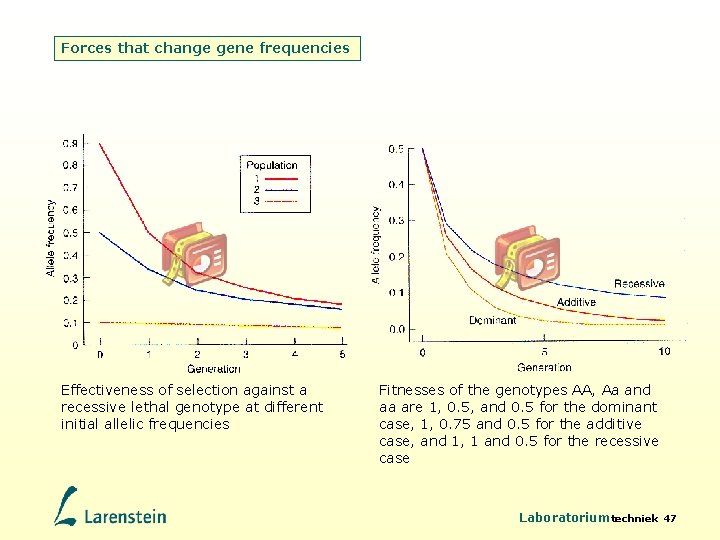
Forces that change gene frequencies Effectiveness of selection against a recessive lethal genotype at different initial allelic frequencies Fitnesses of the genotypes AA, Aa and aa are 1, 0. 5, and 0. 5 for the dominant case, 1, 0. 75 and 0. 5 for the additive case, and 1, 1 and 0. 5 for the recessive case Laboratoriumtechniek 47

Forces that change gene frequencies • Heterozygote superiority = heterosis = overdominance • In this case both alleles are maintained in the population at an equilibrium, depending on the relative fitnesses of the homozygotes. If the selection coefficient of AA is s and the selection coefficient of aa is t, then at equilibrium: peq=f(A)=t/(t+s) and qeq=f(a)=s/(t+s) • Example: Sickle-cell anemia is common in areas with malaria. Hb-A/Hb-A has normal blood Hb-A/Hb-S has sickle-cell trait Hb-S/Hb-S has sickle-cell disease Heterozygotes are relatively resistent to the malaria parasite. Laboratoriumtechniek 48

Forces that change gene frequencies Balance between mutation and natural selection • Natural selection reduces the frequency of deleterious alleles: q=–spq 2 • Mutation produces new alleles that can be harmful: q=up • At equilibrium both forces are in balance: spq 2 =up • So at equilibrium: qeq= (u/s) • If the recessive homozygote is lethal: qeq= u • Example: mutation rate u=10– 6 and s=0. 1 at equilibrium the frequency of the allele will be qeq= u/s=0. 0032 • Most deleterious alleles remain within the population at low frequency because of equilibrium between mutation and natural selection Laboratoriumtechniek 49

Forces that change gene frequencies Assortative mating • Many populations do not mate randomly for some traits • Positive assortative mating occurs when similar phenotypes mate preferentially • Negative assortative mating occurs when dissimilar phenotypes mate preferentially • Assortative mating does not alter the allelic frequencies, but it may influence the genotypic frequencies Laboratoriumtechniek 50

Forces that change gene frequencies Inbreeding • Inbreeding involves preferential mating between relatives • Inbreeding is measured as F, the coefficient of inbreeding. The greater F, the greater the reduction in heterozygosity. expected heterozygosity – observed heterozygosity F = expected heterozygosity • In random mating F=0 • In self-fertilisation F=0. 5, the heterozygote frequency decreases by 50% in each generation • Inbreeding leads to reduced fitness: inbreeding depression Laboratoriumtechniek 51

Forces that change gene frequencies Summary of effects of evolutionary forces on genetic structure Exercises: 22. 15 -22. 34 with a * Laboratoriumtechniek 52

The role of genetics in conservation biology • Reduction of habitat size decreases population size: the gene pool gets smaller which may affect long term survival • Population viability analysis estimates the population size that is necessary for survival • Breeding programmes or action against habitat loss? Laboratoriumtechniek 53

Speciation is division of one species into two new species • A species is a group of interbreeding organisms (or that exchange hereditary material) • Population subdivision may be weak or extreme to the point that two populations never interbreed • If isolation takes place for a long period, fixation of different alleles occurs, leading to • Postzygotic isolation: hybrids are infertile or have low fitness • Individuals that engage interspecific mating suffer disadvantage • Prezygotic isolation: - temporal isolation by changing the mating period - ecological isolation by exploiting different niche • Genetic means to prevent the formation of hybrids: - behavioral incompatibility: both species avoid eachother - mechanical isolation: their genetalia do not fit - gametic isolation: fusion of the gametes fails Laboratoriumtechniek 54

Speciation can be allopatric or sympatric • Allopatric speciation occurs when populations become geographically isolated, are exposed to divergent selection and evolve independantly. After secundary contact postzygotic isolation prevents gene flow, reinforced by (following) prezygotic isolation. • In sympatric speciation gene flow is limited by some form of isolation, but the populations are not totally separated. The most likely cause is selection to use different resources. Example: The fruitfly Rhagoletis polmonella used to lay its eggs in hawthorn. In 1864 Rhagoletis was found in apples. Today we can discriminate between an “apple race” and a “hawthorn race”. Behaviour stimulates divergence: - females prefer to lay eggs in the fruit in which they grew up - males prefer to mate on the fruit from which they emerged Breeding times have diverged, enzym patterns are different. • Fast sympatric speciation is possible by polyploidisation Laboratoriumtechniek 55

Speciation Genetic basis for speciation • Younger species rely mainly on postzygotic barriers • Often male (XY) hybrids are sterile, not the females (XX). Why? - Possibly because many genes are involved in the fertility of males. - By pleiotropic effects of genes that were selected during adaptation to different environments • Sexual selection Example: 800 Drosophila species in Hawaii are the result of allopatric speciation caused by founder effect. Remarkable morphology of males is byproduct of genes controlling courtship, females are often similar. Females choose only their own type. In Lake Victoria in Africa >500 cichlid species evolved in 13. 000 years. Males of related species only differ in colour. Female choice of male colour causes reproductive isolation. Laboratoriumtechniek 56
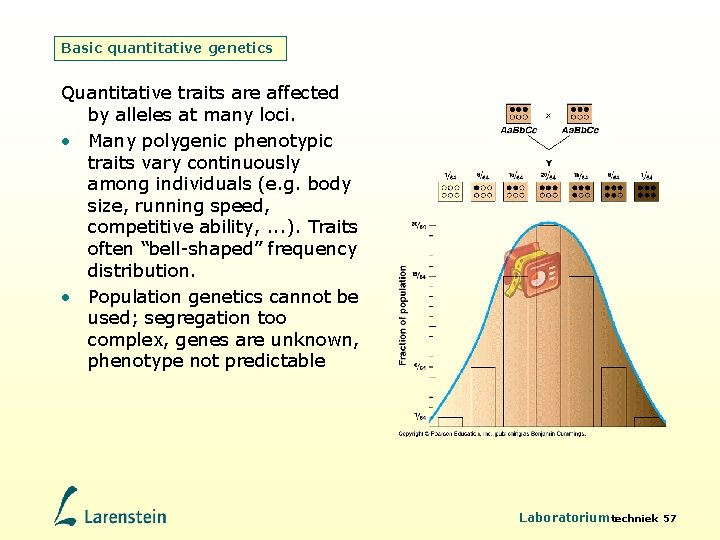
Basic quantitative genetics Quantitative traits are affected by alleles at many loci. • Many polygenic phenotypic traits vary continuously among individuals (e. g. body size, running speed, competitive ability, . . . ). Traits often “bell-shaped” frequency distribution. • Population genetics cannot be used; segregation too complex, genes are unknown, phenotype not predictable Laboratoriumtechniek 57
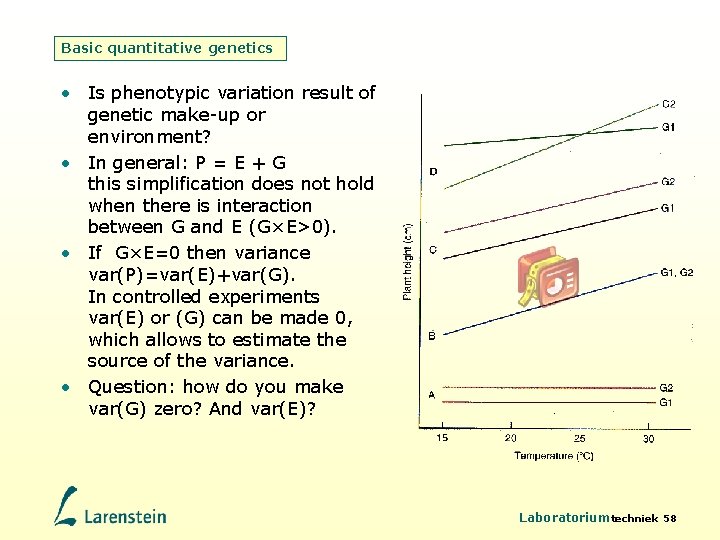
Basic quantitative genetics • Is phenotypic variation result of genetic make-up or environment? • In general: P = E + G this simplification does not hold when there is interaction between G and E (G×E>0). • If G×E=0 then variance var(P)=var(E)+var(G). In controlled experiments var(E) or (G) can be made 0, which allows to estimate the source of the variance. • Question: how do you make var(G) zero? And var(E)? Laboratoriumtechniek 58
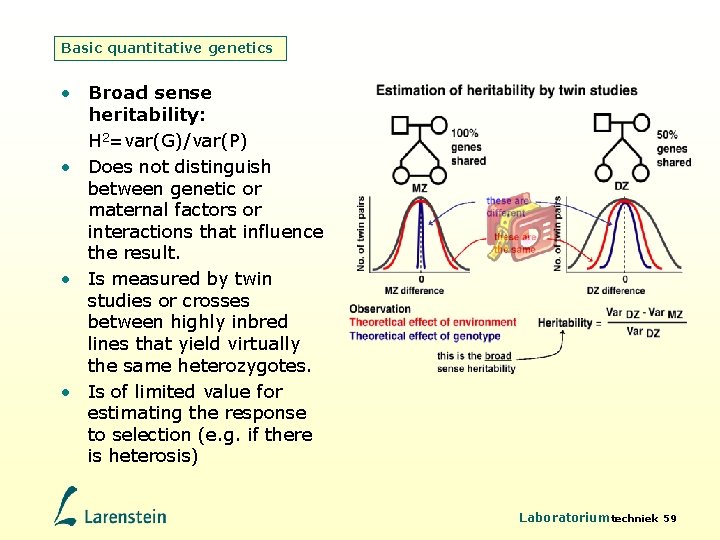
Basic quantitative genetics • Broad sense heritability: H 2=var(G)/var(P) • Does not distinguish between genetic or maternal factors or interactions that influence the result. • Is measured by twin studies or crosses between highly inbred lines that yield virtually the same heterozygotes. • Is of limited value for estimating the response to selection (e. g. if there is heterosis) Laboratoriumtechniek 59

Basic quantitative genetics • Var(G) = var(A)+var(D)+var(I) in which: var(D) = variance due to dominance at one locus var(I) = variance due to epistasis between loci var(A) = additive genetic variance, responds to selection • Narrow sense heritability: h 2=var(A)/var(P) • h 2 is estimated from the response to selection itself, in so-called truncation selection: only individuals with an interesting trait above a certain (truncation) value are selected to breed the next generation. S is the selection differential, R is the selection response, and h 2 = R/S Laboratoriumtechniek 60
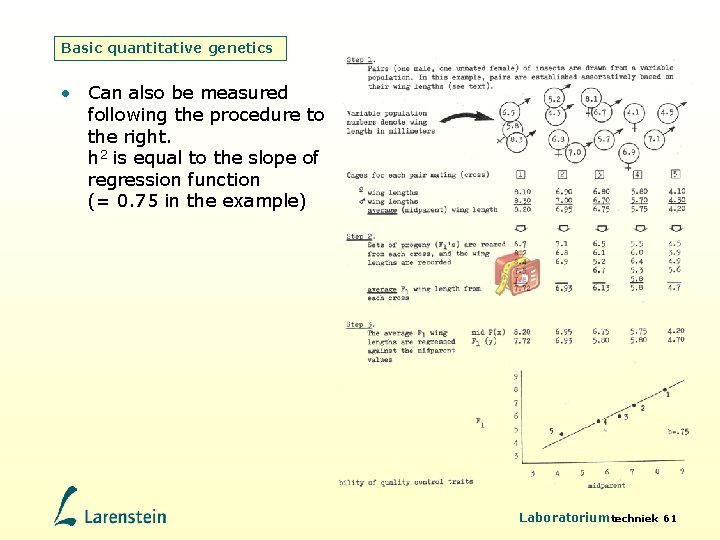
Basic quantitative genetics • Can also be measured following the procedure to the right. h 2 is equal to the slope of regression function (= 0. 75 in the example) Laboratoriumtechniek 61
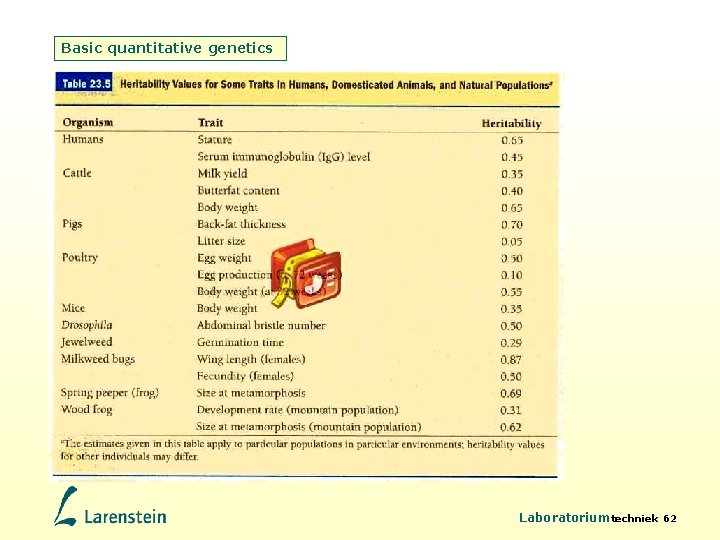
Basic quantitative genetics Laboratoriumtechniek 62
Basic quantitative genetics Question: A botanist measured seed weight and estimated its narrowsense heritability in three widely separated Phlox populations. Mean seed weight Narrow sense heritability Population A 15 mg 0. 60 Population B 12 mg 0. 65 Population C 17 mg 0. 58 Because the heritabilities were high and approximately equal, he concluded that the differences in seed weight between the populations were largely due to genetic differences. Do you agree? Explain. Laboratoriumtechniek 63

Basic quantitative genetics Warnings regarding heritability estimates • Represents contribution of genetic variation in trait: Does not tell how much of trait is genetic or environmental. What does it mean: “intelligence in humans is determined 70% by genes and 30% by environmental conditions” To have a trait you need both. You can only tell to what extent differences between individuals are genetic or environmental. • Depends on population and environment (inbred lab population: lower heritability; Wild type under constant environment: high heritability): h² estimate only valid for populations and conditions tested. • High heritability for a trait does not imply that populations differences in this trait are genetically determined. • An individual does not have heritability; a population does. Laboratoriumtechniek 64

Basic quantitative genetics Evolutionary implications of quantitative genetics • When much additive variation: response to selection rapid and strong • Artificial selection proves that when selection directional, major changes possible • Yet, usually only small changes under natural conditions. Why? – Continuous, directional selection rare due to fluctuations in direction of selection – Pattern of temporal fluctuation may seem to represent average stability – Directional selection may cause negative side-effects, which slow down or halt evolution (trade-offs): After relaxation of selection: return to original state. Laboratoriumtechniek 65

Basic quantitative genetics Integration of population and quantitative genetics • Integration likely because of progress in molecular biology – Discovery of many alleles with quantitative effects at single loci – Link between DNA sequences and phenotypic traits – Progress in computational techniques = possible to analyse multi-gene traits • QTL analysis (Quantitative trait loci analysis) 1. Develop molecular markers (variable DNA sequence). Associate these markers with extreme values for a quantitative, phenotypic trait 2. Localise and characterise gene in vicinity of marker • Conclusions: quantitative traits often controlled by few majoreffect genes and many small-effect genes (modifiers). Laboratoriumtechniek 66

Genetic problems Q 1 In a population of 2000 gaboon vipers, a genetic difference with respect to venom exists at a single locus. The alleles are incompletely dominant. The population shows 100 individuals homozygous for the t allele (genotype tt, non-poisonous), 800 heterozygotes (genotype Tt, mildlypoisonous) and 1100 homozygous for the T-allele (genotype TT, lethally poisonous). a. What is the frequency of the t-allele in the population? b. Are the genotypes in Hardy-Weinberg equilibrium? Q 2 Approximately one normal allele in 30, 000 mutates to the X-linked recessive allele for hemophilia in each human generation. Assume for the purpose of this problem that one h allele in 300, 000 mutates to the normal alternative in each generation. What allelic frequencies would prevail at equilibrium under mutation pressure alone? Laboratoriumtechniek 67

Non-Mendelian inheritance Some traits do not behave in a Mendelian fashion: • Maternal inheritance A type of uniparental inheritance in which all progeny have the genotype and phenotype of the female parent • Maternal effect The effect of the maternal parent's genotype on the phenotype of her offspring • Genetic imprinting The phenomenon in which there is differential expression of a gene depending on whether it was maternally or paternally inherited. Paternal imprinting means that an allele inherited from the father is not expressed in offspring. Maternal imprinting means that an allele inherited from the mother is not expressed in offspring Laboratoriumtechniek 68

Non-Mendelian inheritance • Maternal inheritance Example: four-o-clocks • Maternal effect Example: Lymnea peregra Laboratoriumtechniek 69
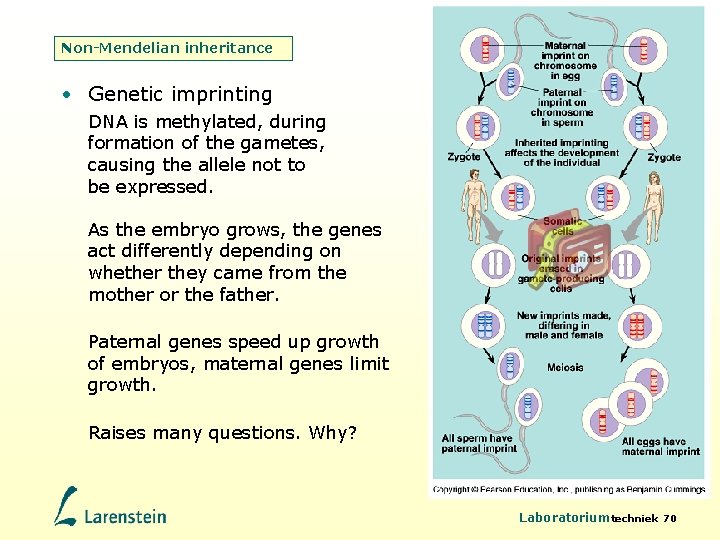
Non-Mendelian inheritance • Genetic imprinting DNA is methylated, during formation of the gametes, causing the allele not to be expressed. As the embryo grows, the genes act differently depending on whether they came from the mother or the father. Paternal genes speed up growth of embryos, maternal genes limit growth. Raises many questions. Why? Laboratoriumtechniek 70

Non-Mendelian inheritance • Genetic imprinting Boys with Prader-Willi (left) and Angelman (right) syndromes Both of these conditions are caused by deletions or other mutations in the same region of chromosome 15. However, part of this region is imprinted (or inactivated) on the maternally inherited chromosome (the PWS region), and part is imprinted on the paternal chromosome (the AS gene, which is called UBE 3 A). If, for example, there is a deletion in this region on the paternal chromosome 15, neither chromosome 15 of the child will express the genes in the PWS region (the copy inherited from the mother is inactivated). As a result, the child will have PWS. However, if the deletion is on the maternal chromosome 15, then neither chromosome 15 of the child will express the genes in the AS region, and hence, this child will have AS. Laboratoriumtechniek 71
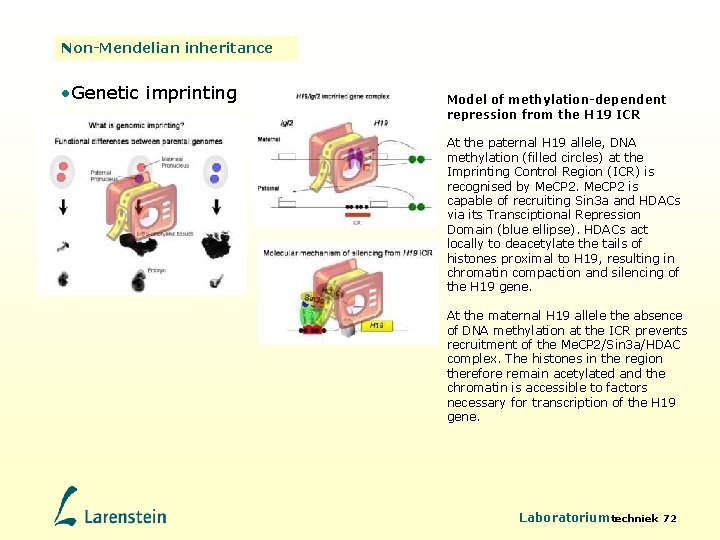
Non-Mendelian inheritance • Genetic imprinting Model of methylation-dependent repression from the H 19 ICR At the paternal H 19 allele, DNA methylation (filled circles) at the Imprinting Control Region (ICR) is recognised by Me. CP 2 is capable of recruiting Sin 3 a and HDACs via its Transciptional Repression Domain (blue ellipse). HDACs act locally to deacetylate the tails of histones proximal to H 19, resulting in chromatin compaction and silencing of the H 19 gene. At the maternal H 19 allele the absence of DNA methylation at the ICR prevents recruitment of the Me. CP 2/Sin 3 a/HDAC complex. The histones in the region therefore remain acetylated and the chromatin is accessible to factors necessary for transcription of the H 19 gene. Laboratoriumtechniek 72
- Slides: 72