Intro Bio Lecture 5 1 Proteins amino acids

Intro Bio Lecture 5 1
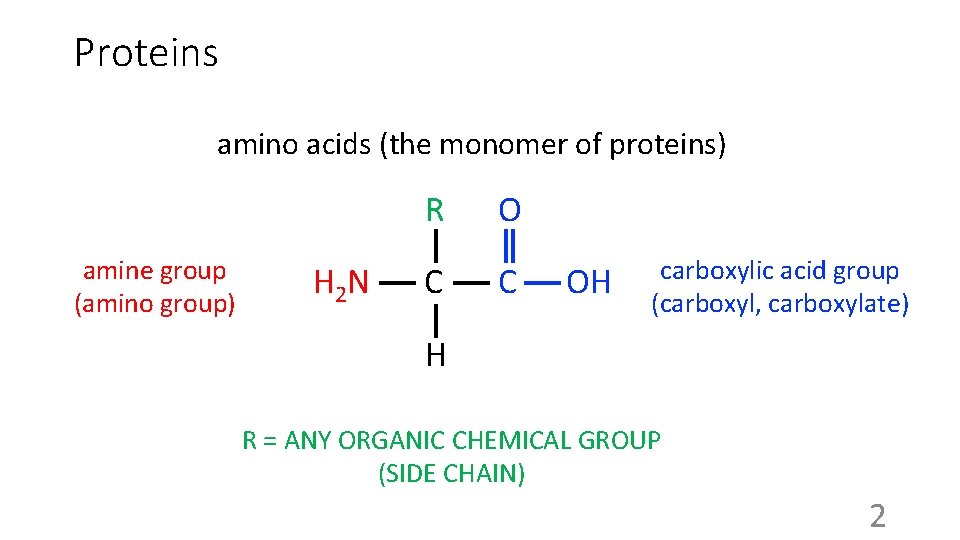
Proteins amino acids (the monomer of proteins) R amine group (amino group) H 2 N C OH carboxylic acid group (carboxyl, carboxylate) H R = ANY ORGANIC CHEMICAL GROUP (SIDE CHAIN) 2
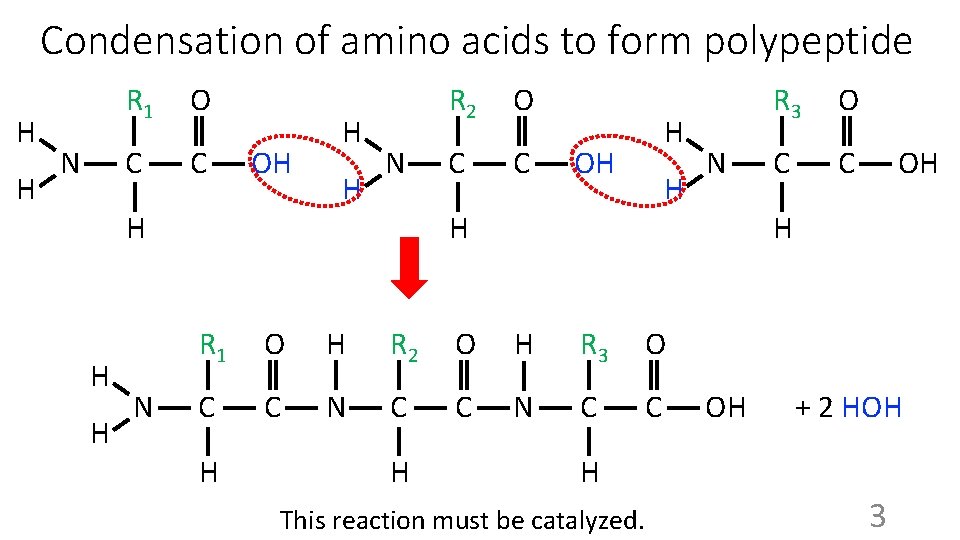
Condensation of amino acids to form polypeptide H H N R 1 O C C OH H N R 2 O C C H OH H H R 1 O H R 2 O H R 3 O C C N C C H H N R 3 O C C OH H OH + 2 HOH H This reaction must be catalyzed. 3
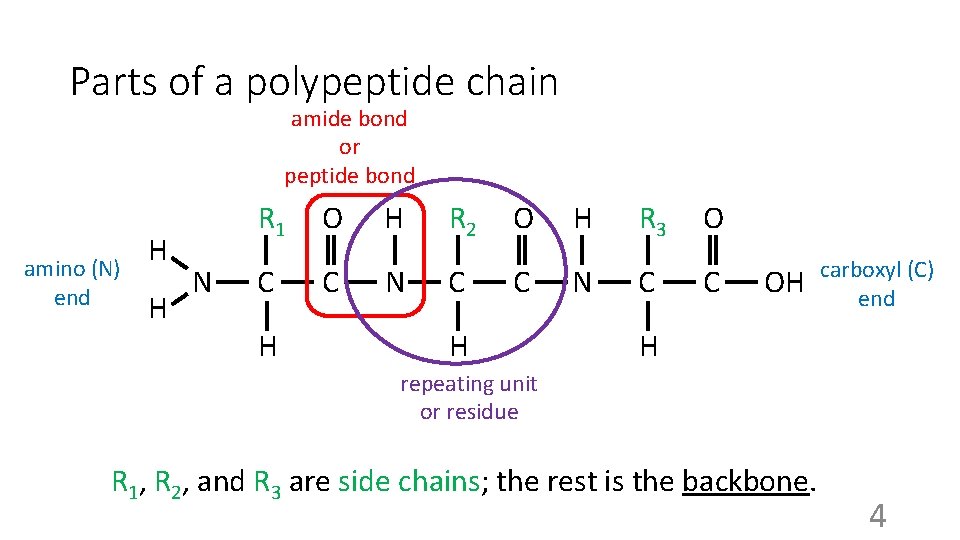
Parts of a polypeptide chain amide bond or peptide bond amino (N) end H H N R 1 O H R 2 O H R 3 O C C N C C H H OH carboxyl (C) end H repeating unit or residue R 1, R 2, and R 3 are side chains; the rest is the backbone. 4

Parts of a polypeptide chain amide bonds amino (N) end H H N carboxyl (C) end R 1 O H R 2 O H R 3 O H R 4 O H R 5 O C N C N C H C H C C OH H 5

Parts of a polypeptide chain amino (N) end H H N carboxyl (C) end R 1 O H R 2 O H R 3 O H R 4 O H R 5 O C N C N C H C C H repeating unit or residue H C C OH H 6
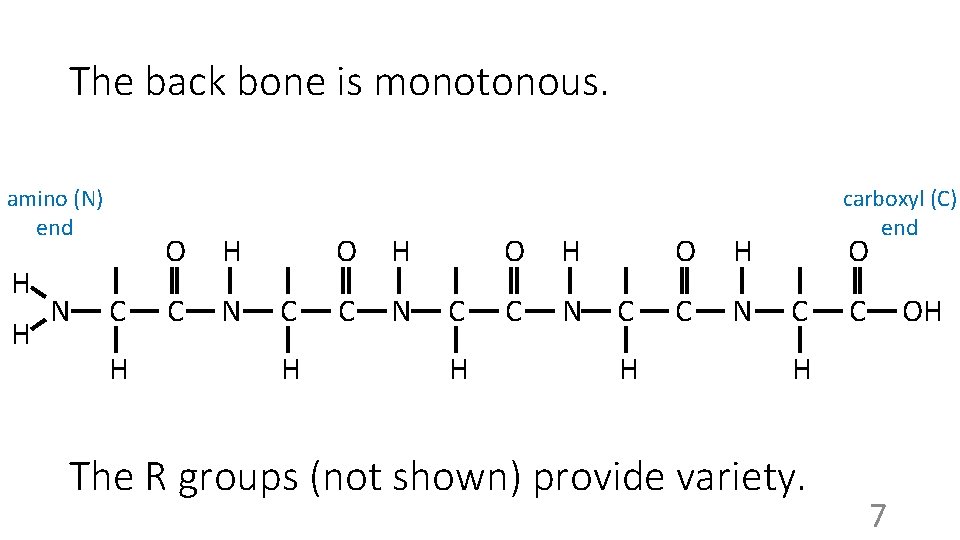
The back bone is monotonous. amino (N) end H H N C H O H C N carboxyl (C) end O C C OH H The R groups (not shown) provide variety. 7
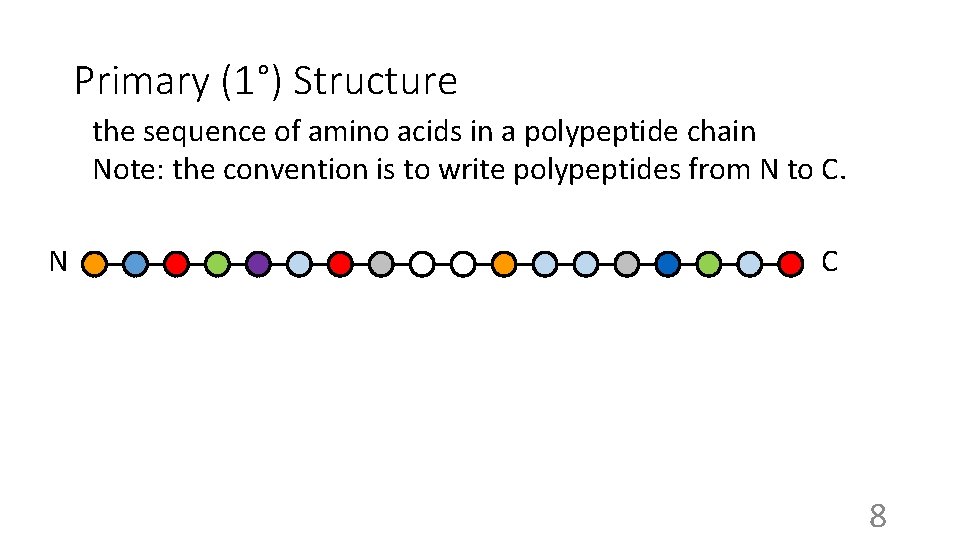
Primary (1°) Structure the sequence of amino acids in a polypeptide chain Note: the convention is to write polypeptides from N to C. N C 8

Sequence Determination Method Phase I Carboxypeptidase: hydrolyzes this peptide bond One way: use an enzyme: (an older method, but useful for teaching) N C 9
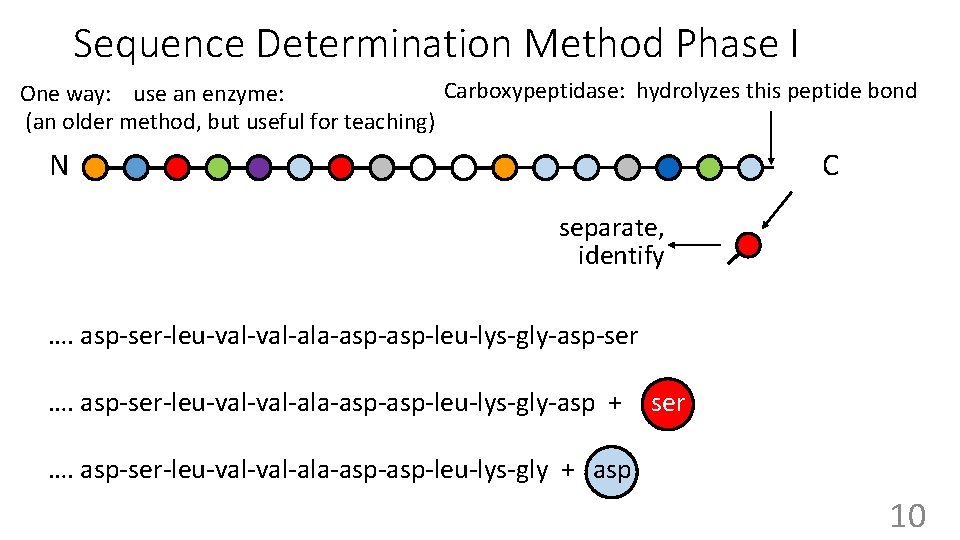
Sequence Determination Method Phase I Carboxypeptidase: hydrolyzes this peptide bond One way: use an enzyme: (an older method, but useful for teaching) N C separate, identify …. asp-ser-leu-val-ala-asp-leu-lys-gly-asp-ser …. asp-ser-leu-val-ala-asp-leu-lys-gly-asp + ser …. asp-ser-leu-val-ala-asp-leu-lys-gly + asp 10
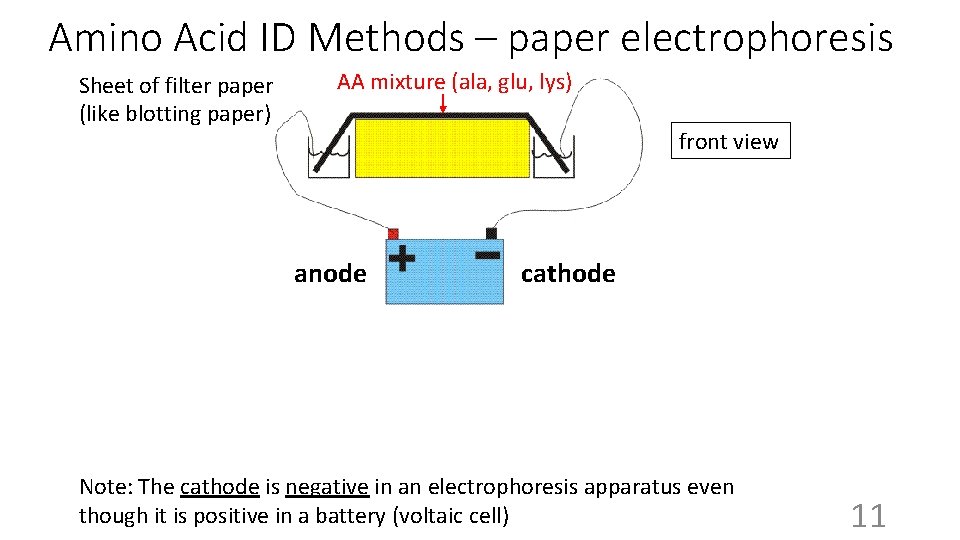
Amino Acid ID Methods – paper electrophoresis AA mixture (ala, glu, lys) Sheet of filter paper (like blotting paper) front view anode Anode cathode (-) (+) Cathode Note: The cathode is negative in an electrophoresis apparatus even though it is positive in a battery (voltaic cell) 11
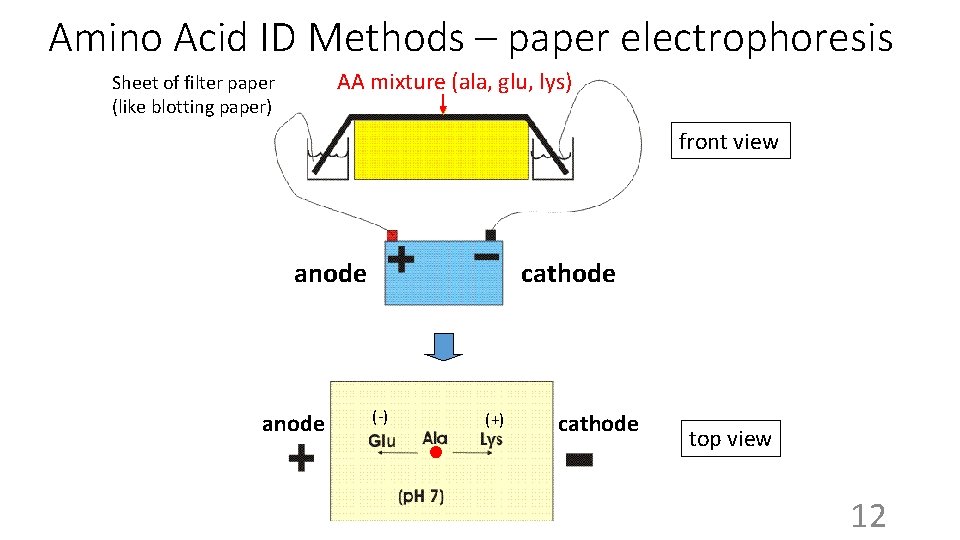
Amino Acid ID Methods – paper electrophoresis AA mixture (ala, glu, lys) Sheet of filter paper (like blotting paper) front view anode cathode (-) (+) Cathode cathode top view 12

A paper electrophoresis apparatus ~3000 volts DC! buffer reservoirs 13
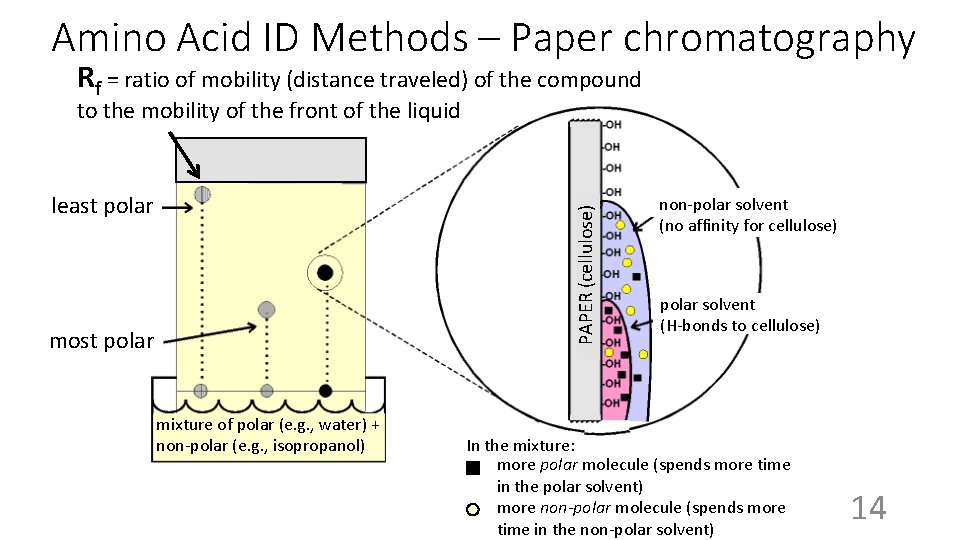
Amino Acid ID Methods – Paper chromatography Rf = ratio of mobility (distance traveled) of the compound to the mobility of the front of the liquid PAPER (cellulose) least polar most polar mixture of polar (e. g. , water) + non-polar (e. g. , isopropanol) non-polar solvent (no affinity for cellulose) polar solvent (H-bonds to cellulose) In the mixture: more polar molecule (spends more time in the polar solvent) more non-polar molecule (spends more time in the non-polar solvent) 14

Amino Acid ID Methods – Paper chromatography “Rf” “front” = 1. 00 After stopping the paper chromatography and staining for the amino acids: Rf = distance relative to the front ile 0. 82 trp 0. 69 gly 0. 45 ser 0. 27 asp 0. 11 15

Paper chromatography apparatus Dyes here (black Magic Marker). Amino acids are colorless, and must be stained to visualize. 16
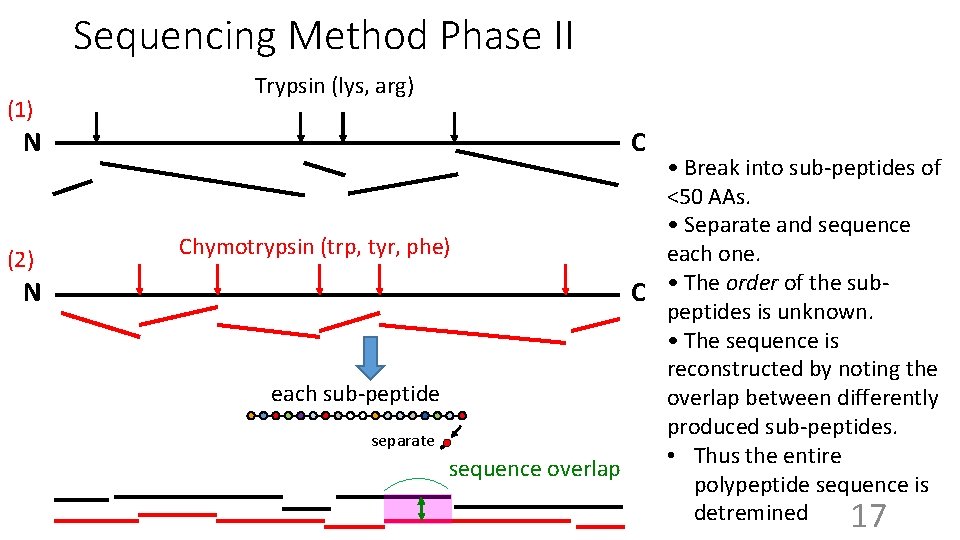
Sequencing Method Phase II (1) N (2) N Trypsin (lys, arg) C • Break into sub-peptides of <50 AAs. • Separate and sequence Chymotrypsin (trp, tyr, phe) each one. C • The order of the subpeptides is unknown. • The sequence is reconstructed by noting the each sub-peptide overlap between differently produced sub-peptides. separate • Thus the entire sequence overlap polypeptide sequence is detremined 17

00000000000000000000000000 After protease A (cleaves after n): ation ghaswin gsbutn in whohasimagin man ofeet withoutlearn 52 letters (AAs, say) After protease B (cleaves after i): magi manwhohasi nghaswi ngsbutnofeet onwi thoutlearni nati onwi ation nati onwi manwhohasi magi nati onwi thoutlearni nghaswi ngsbutnofeet ||| \\\|\ |||||| |\\\\\\ man whohasimagin ation withoutlearn in ghaswin gsbutn ofeet Man who has imagination without learning has wings but no feet 18
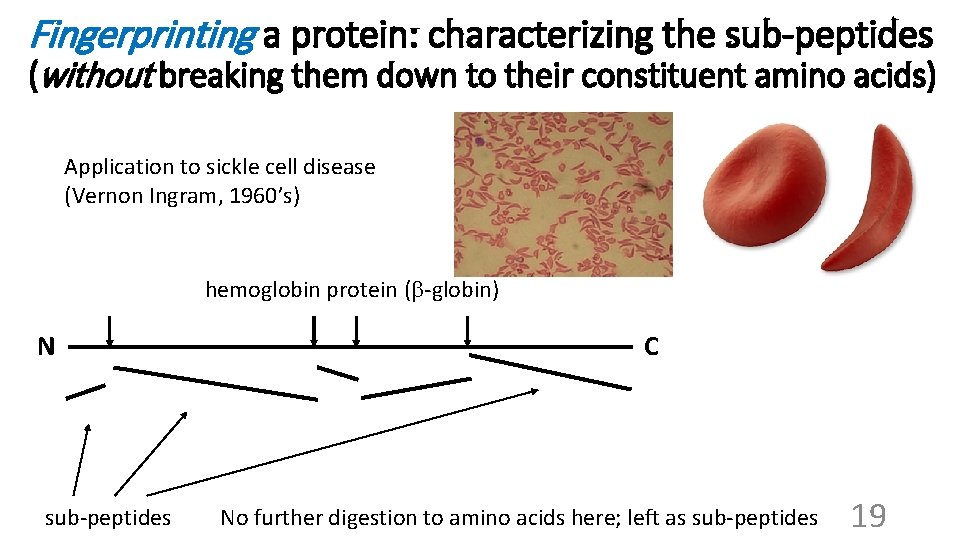
Fingerprinting a protein: characterizing the sub-peptides (without breaking them down to their constituent amino acids) Application to sickle cell disease (Vernon Ingram, 1960’s) hemoglobin protein (b-globin) N sub-peptides C No further digestion to amino acids here; left as sub-peptides 19
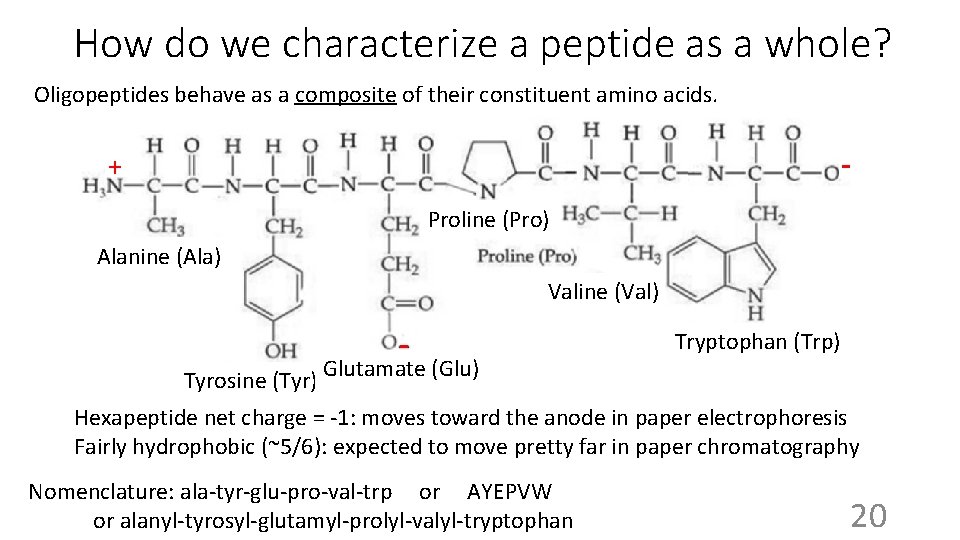
How do we characterize a peptide as a whole? Oligopeptides behave as a composite of their constituent amino acids. - + Proline (Pro) Alanine (Ala) Valine (Val) - Tyrosine (Tyr) Glutamate (Glu) Tryptophan (Trp) Hexapeptide net charge = -1: moves toward the anode in paper electrophoresis Fairly hydrophobic (~5/6): expected to move pretty far in paper chromatography Nomenclature: ala-tyr-glu-pro-val-trp or AYEPVW or alanyl-tyrosyl-glutamyl-prolyl-valyl-tryptophan 20
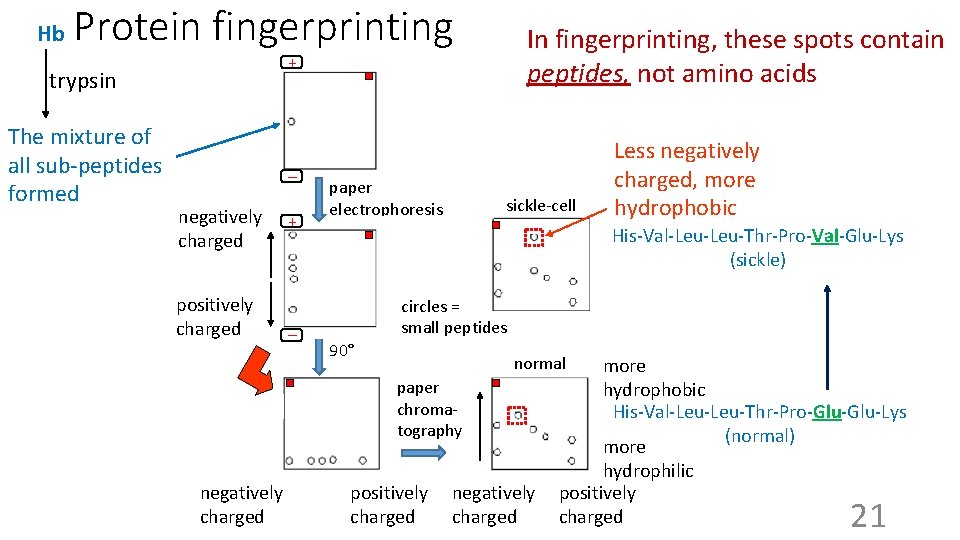
Hb Protein fingerprinting + trypsin The mixture of all sub-peptides formed In fingerprinting, these spots contain peptides, not amino acids _ negatively charged positively charged + paper electrophoresis sickle-cell His-Val-Leu-Thr-Pro-Val-Glu-Lys (sickle) circles = small peptides _ 90° normal paper chromatography negatively charged Less negatively charged, more hydrophobic positively charged negatively charged more hydrophobic His-Val-Leu-Thr-Pro-Glu-Lys (normal) more hydrophilic positively charged 21

Fingerprints for different polypeptides more hydrophobic charge on _ peptide: + _ + _ + more hydrophilic Every different polypeptide has a different primary structure (sequence). So ~every polypeptide will have a different arrangement of spots after fingerprinting. 22

3 -dimensional structure of proteins One given purified polypeptide • Molecule #1: N-met-leu-ala-asp-val-lys-. . • Molecule #2: N-met-leu-ala-asp-val-lys-. . . • Molecule #3: N-met-leu-ala-asp-val-lys-. . . • Molecule #4: N-met-leu-ala-asp-val-lys-. . . etc. clothesline model. . 23

3 D structure of proteins – How does a polypeptide fold correctly? Determining protein 3 -dimensional structure X-ray diffraction of protein crystals Nuclear magnetic resonance Electron microscopy • A very general principle: hydrophobics inside, hydrophilics outside. • Still difficult to predict completely correctly, except for smaller proteins. • Better results using analogies to known structures (not exactly first principles) • Where is the information for choosing the correct folded structure? • Is it being provided by another source (e. g, a scaffold, or cast using some kind of mold)? • Or does it reside in the primary structure itself? 24
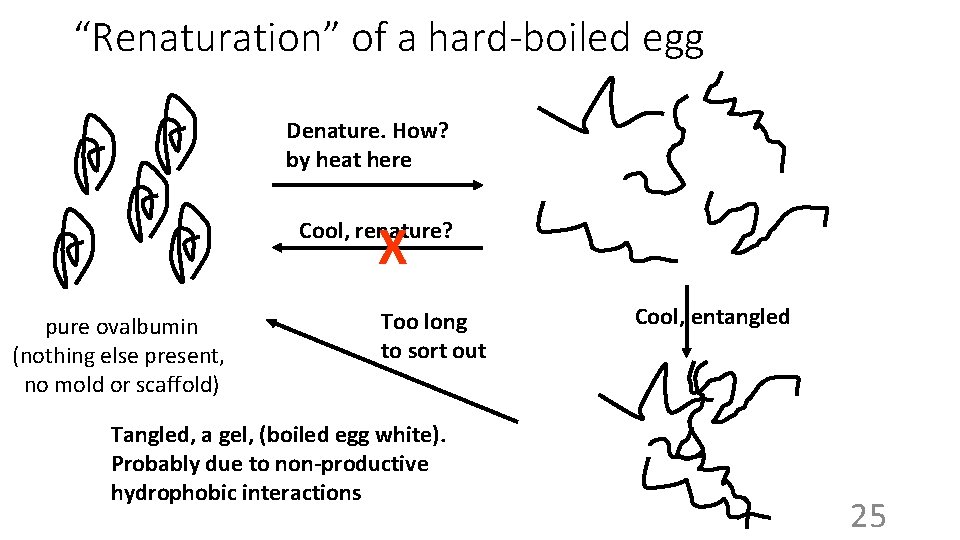
“Renaturation” of a hard-boiled egg Denature. How? by heat here X Cool, renature? pure ovalbumin (nothing else present, no mold or scaffold) Too long to sort out Tangled, a gel, (boiled egg white). Probably due to non-productive hydrophobic interactions Cool, entangled 25
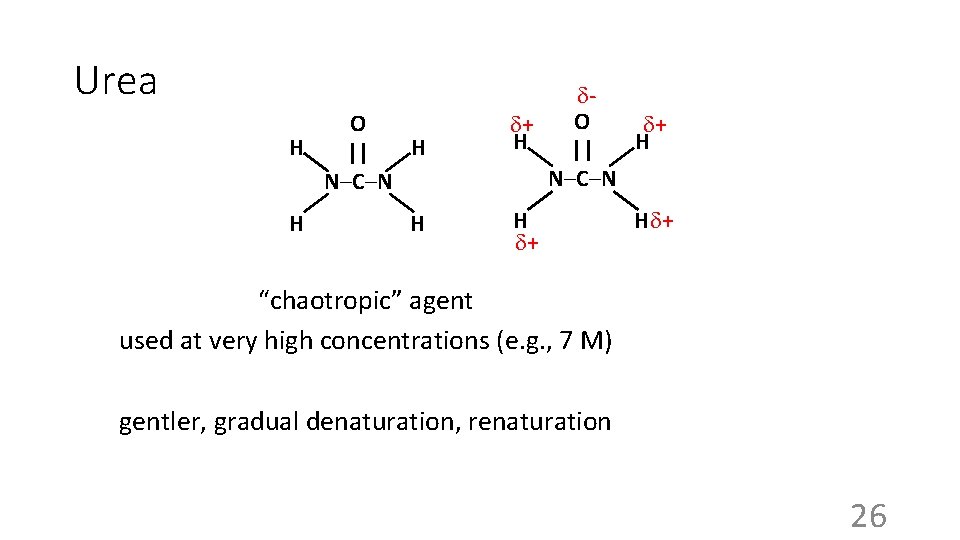
Urea O H H || N–C–N H H O + + H H || -N–C–N + H H + + “chaotropic” agent used at very high concentrations (e. g. , 7 M) gentler, gradual denaturation, renaturation 26
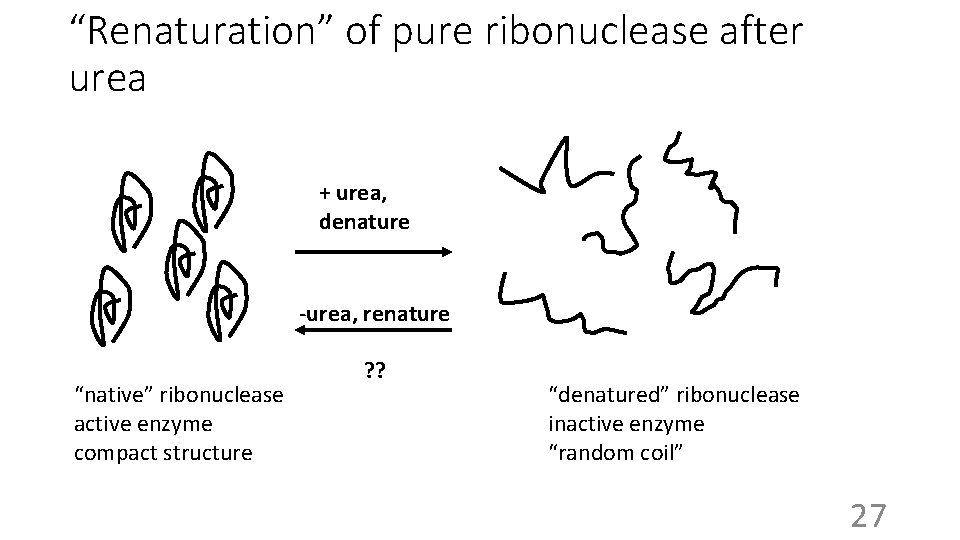
“Renaturation” of pure ribonuclease after urea + urea, denature -urea, renature “native” ribonuclease active enzyme compact structure ? ? “denatured” ribonuclease inactive enzyme “random coil” 27

Slow denaturation and renaturation of pure ribonuclease by urea O || Urea = H 2 N—C—NH 2 urea protein Dialysis Bag is made of a semipermeable membrane Macromolecules (protein here) cannot permeate the bag, but small molecules (H 20, urea) can. Urea will move from areas of high concentration to areas of low concentration. 28
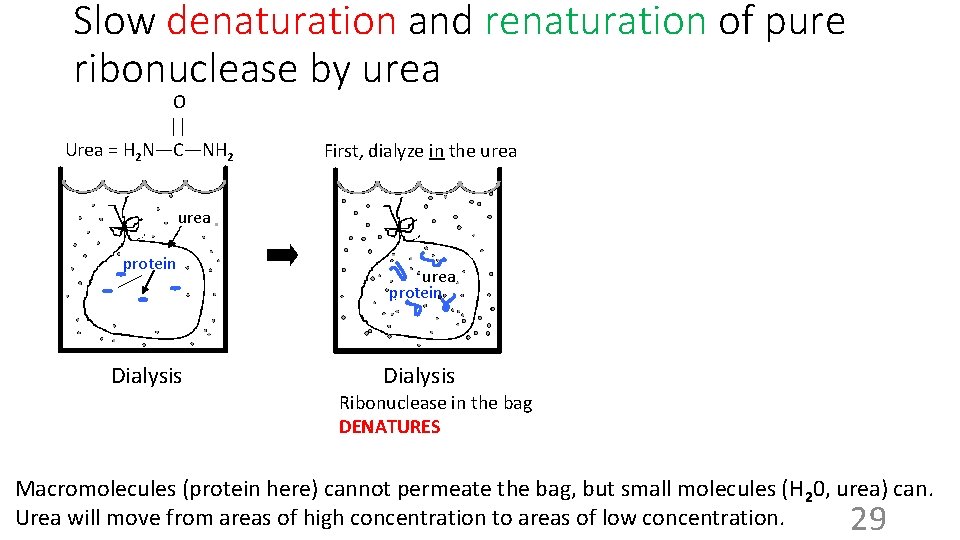
Slow denaturation and renaturation of pure ribonuclease by urea O || Urea = H 2 N—C—NH 2 First, dialyze in the urea protein Dialysis Ribonuclease in the bag DENATURES Macromolecules (protein here) cannot permeate the bag, but small molecules (H 20, urea) can. Urea will move from areas of high concentration to areas of low concentration. 29
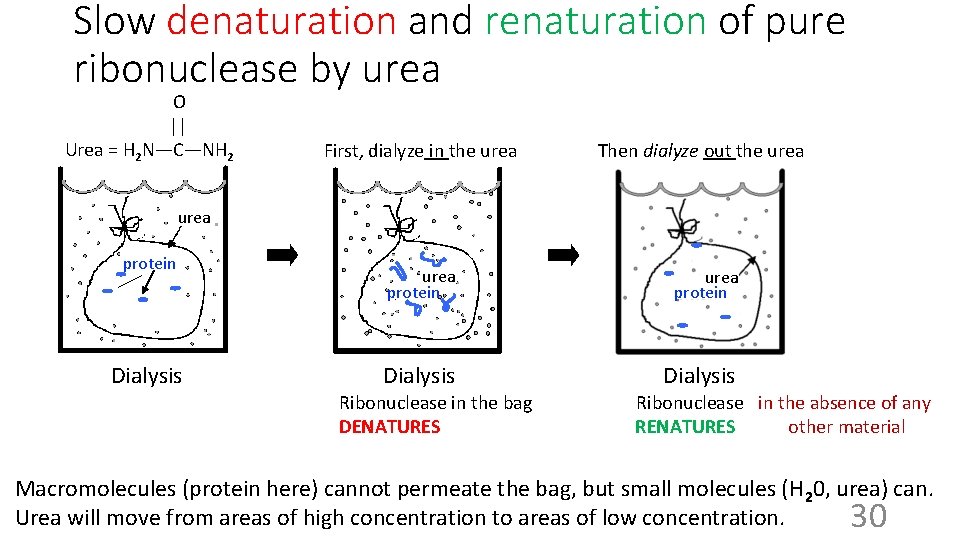
Slow denaturation and renaturation of pure ribonuclease by urea O || Urea = H 2 N—C—NH 2 First, dialyze in the urea Then dialyze out the urea protein Dialysis urea protein Dialysis Ribonuclease in the bag DENATURES Ribonuclease in the absence of any RENATURES other material Macromolecules (protein here) cannot permeate the bag, but small molecules (H 20, urea) can. Urea will move from areas of high concentration to areas of low concentration. 30

PRIMARY structure determines TERTIARY structure Christian Anfinsen: + urea, denatures - urea, renatures “The Anfinsen Experiment” 31
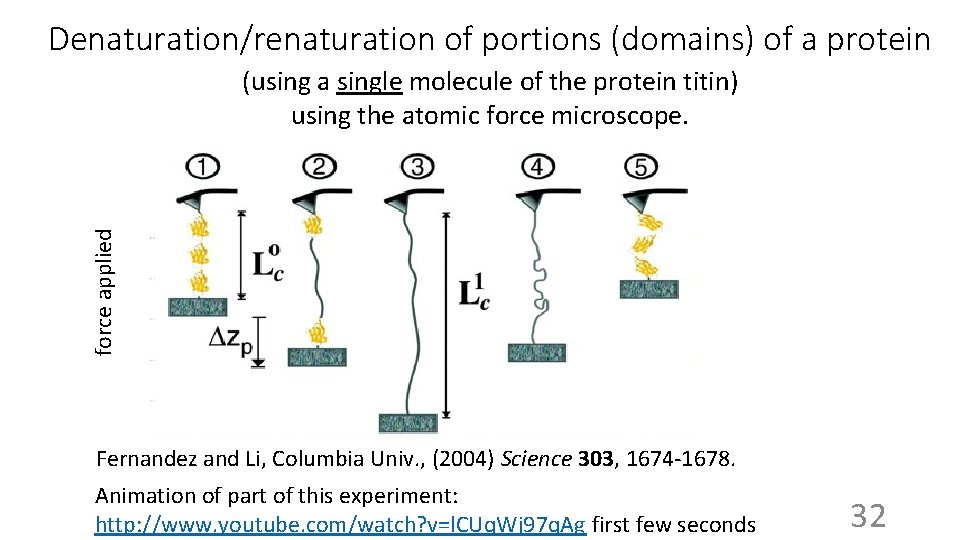
Denaturation/renaturation of portions (domains) of a protein force applied (using a single molecule of the protein titin) using the atomic force microscope. Fernandez and Li, Columbia Univ. , (2004) Science 303, 1674 -1678. Animation of part of this experiment: http: //www. youtube. com/watch? v=l. CUq. Wj 97 q. Ag first few seconds 32

BUT: Chaperonins (proteins themselves) • Help fold proteins during synthesis • Perhaps by preventing illegitimate interactions, like intermolecular contacts via exposed hydrophobic groups of partially folded proteins • Also help re-fold proteins that have denatured after passing through a membrane’s phospholipid bilayer, e. g. , during transport into a mitochondrion (organelle). 33
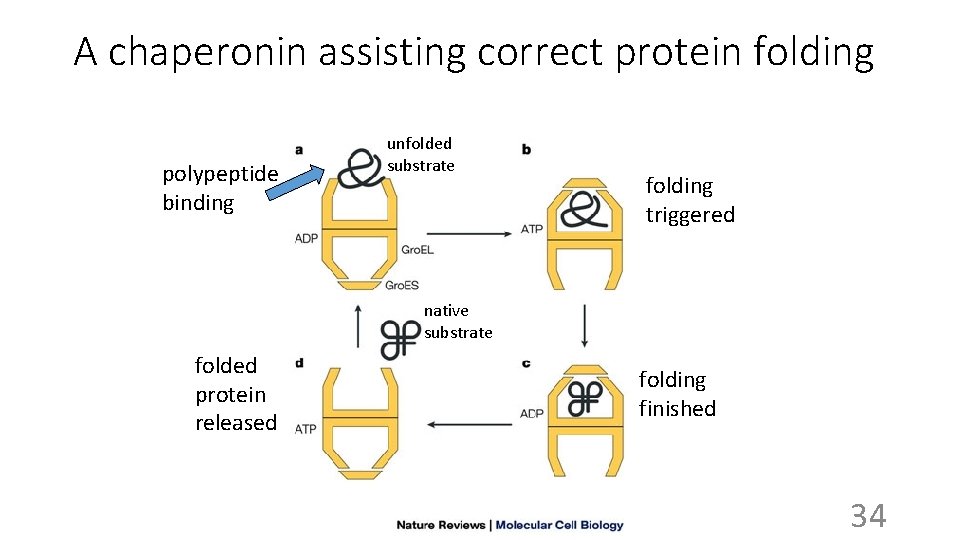
A chaperonin assisting correct protein folding polypeptide binding unfolded substrate folding triggered native substrate folded protein released folding finished 34
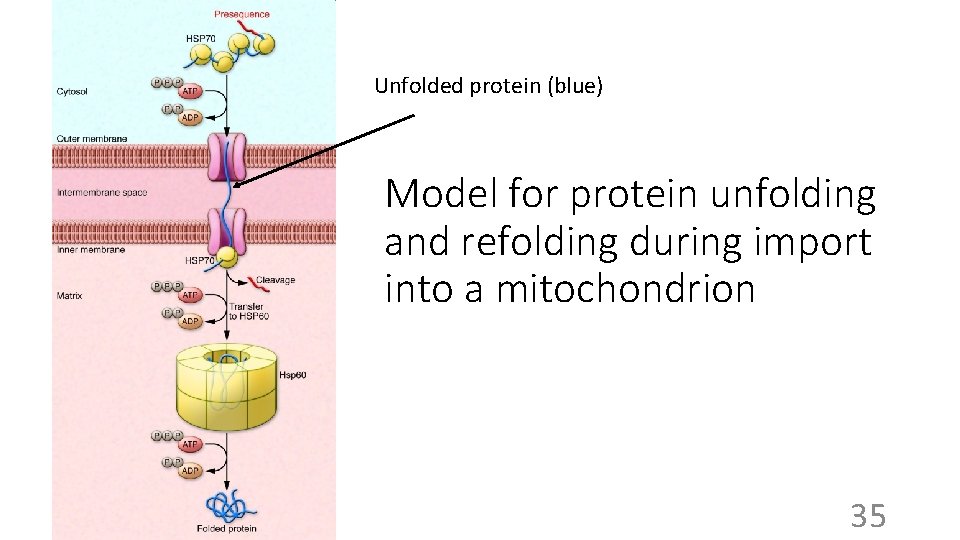
Unfolded protein (blue) Model for protein unfolding and refolding during import into a mitochondrion 35
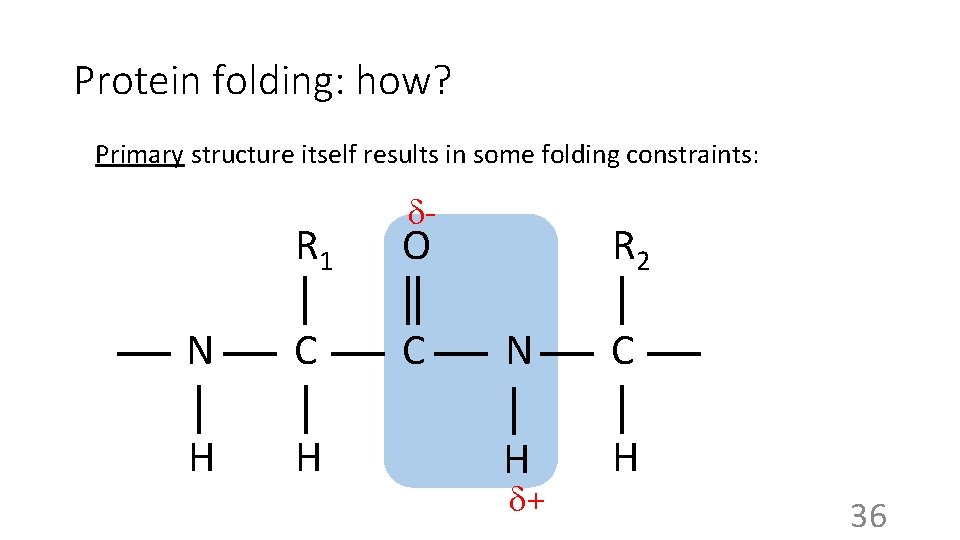
Protein folding: how? Primary structure itself results in some folding constraints: - R 1 O N C C H H R 2 N C H H + 36
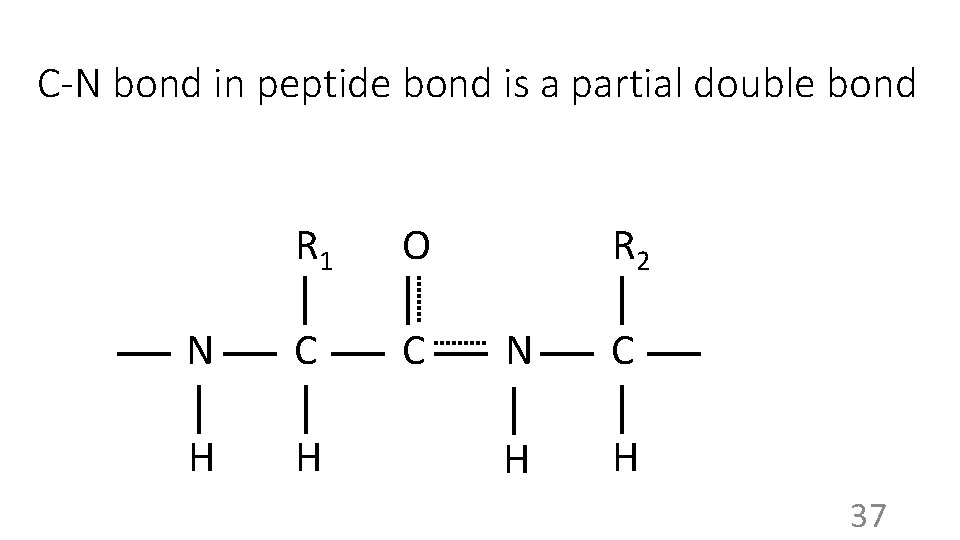
C-N bond in peptide bond is a partial double bond R 1 O N C C H H R 2 N C H H 37
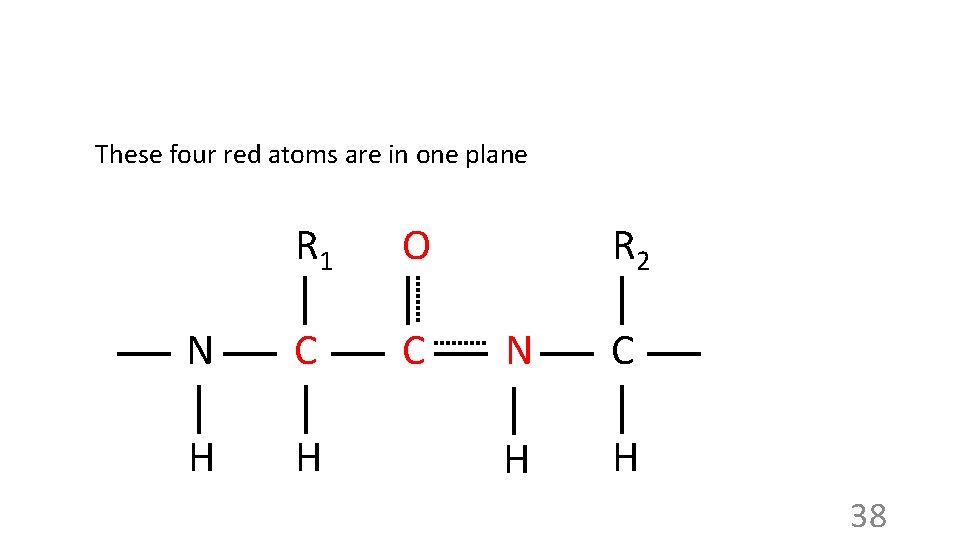
These four red atoms are in one plane R 1 O N C C H H R 2 N C H H 38
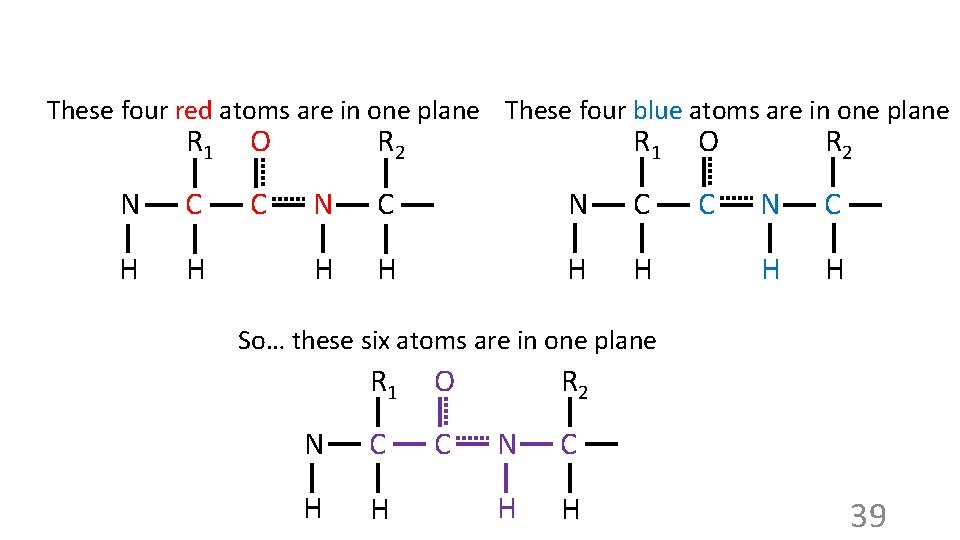
These four red atoms are in one plane These four blue atoms are in one plane R 1 O N C C H H R 2 R 1 O C N C H H R 2 N C H H So… these six atoms are in one plane R 1 O N C C H H R 2 N C H H 39
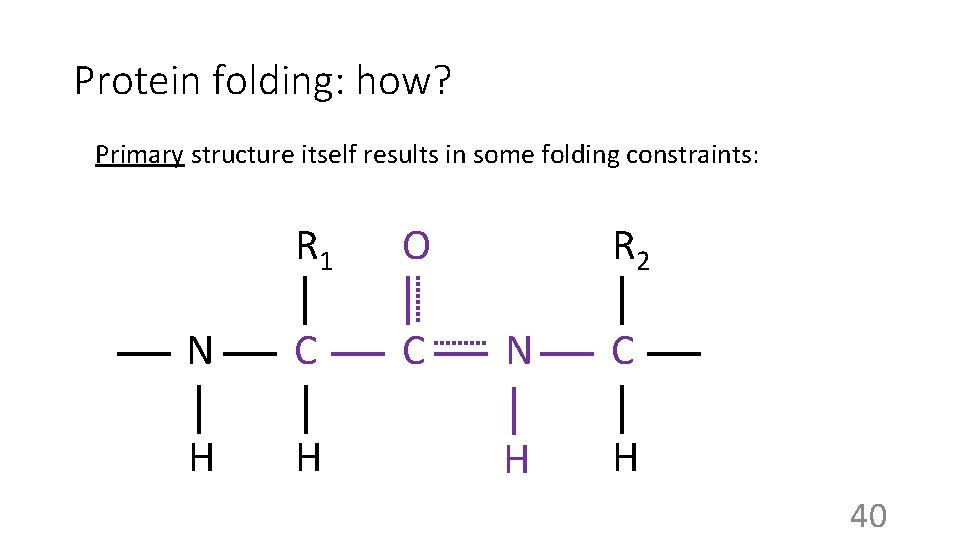
Protein folding: how? Primary structure itself results in some folding constraints: R 1 O N C C H H R 2 N C H H 40
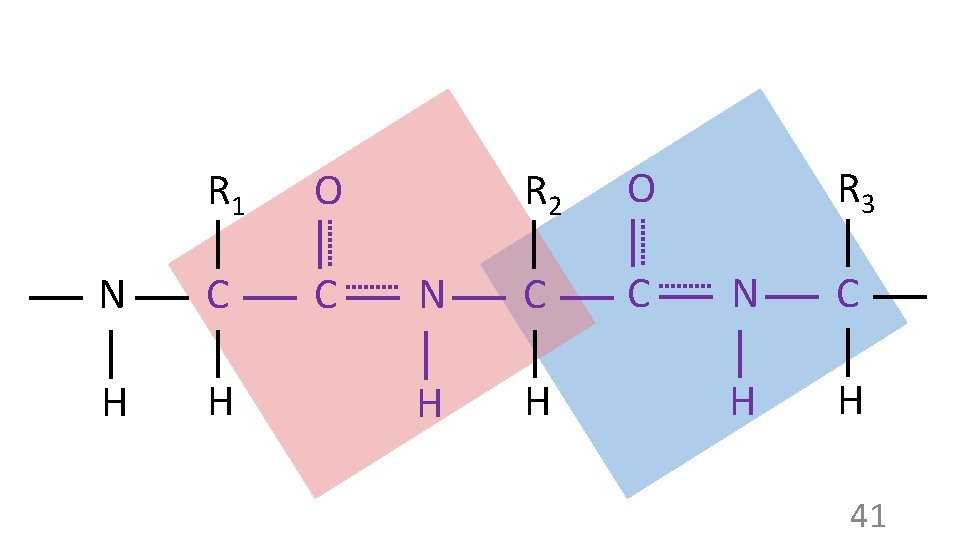
R 1 O N C C H H R 2 O N C C H H R 3 N C H H 41
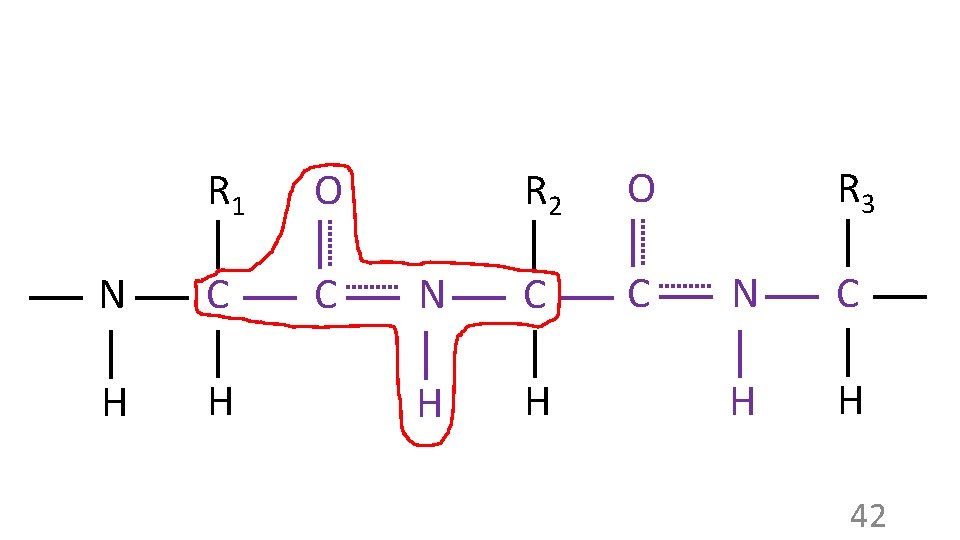
R 1 O N C C H H R 2 O N C C H H R 3 N C H H 42
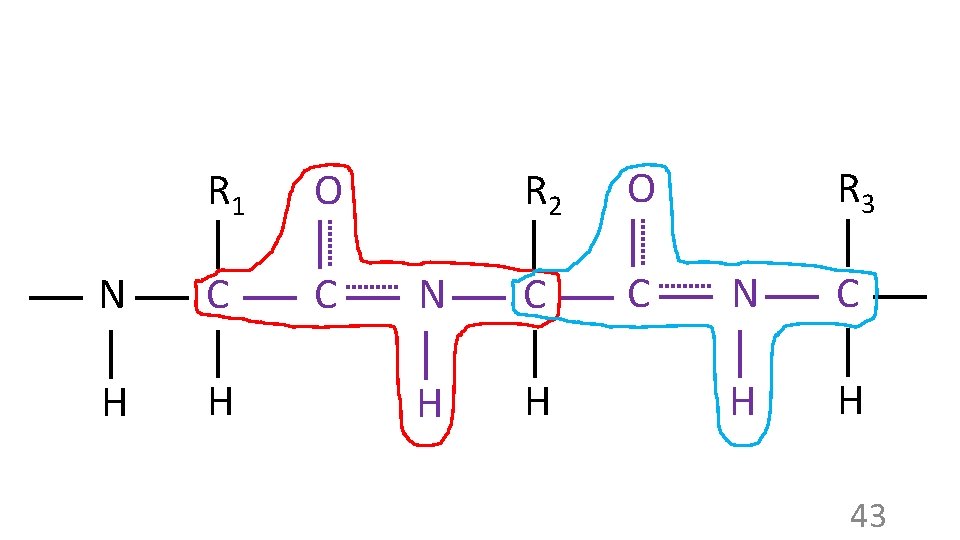
R 1 O N C C H H R 2 O N C C H H R 3 N C H H 43
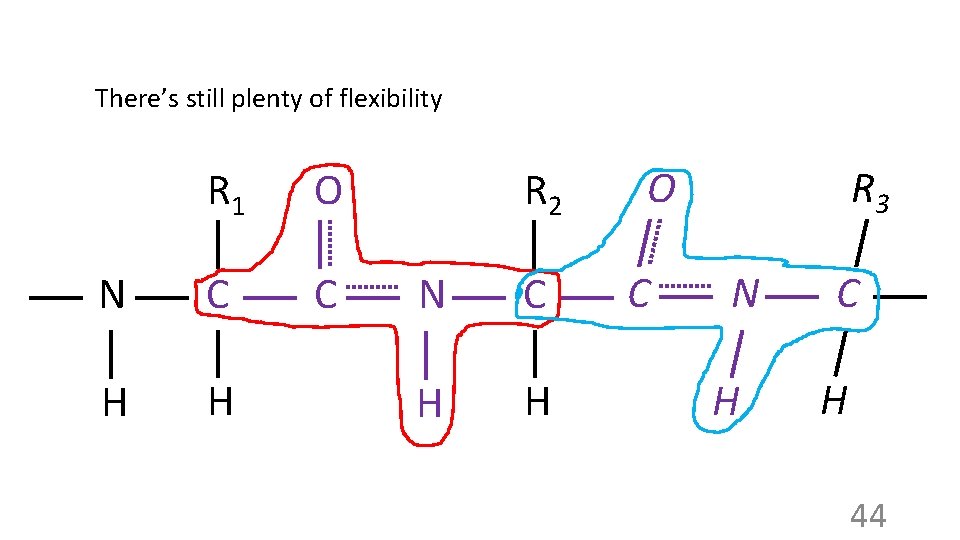
There’s still plenty of flexibility R 1 O N C C H H R 2 N C H H O C R 3 N H C H 44
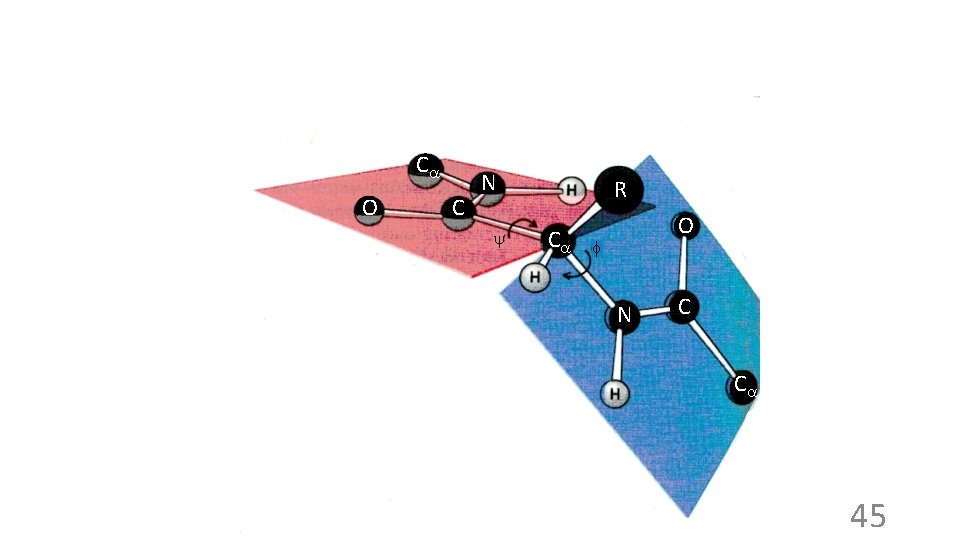
Ca O C N y R Ca O f N C Ca 45
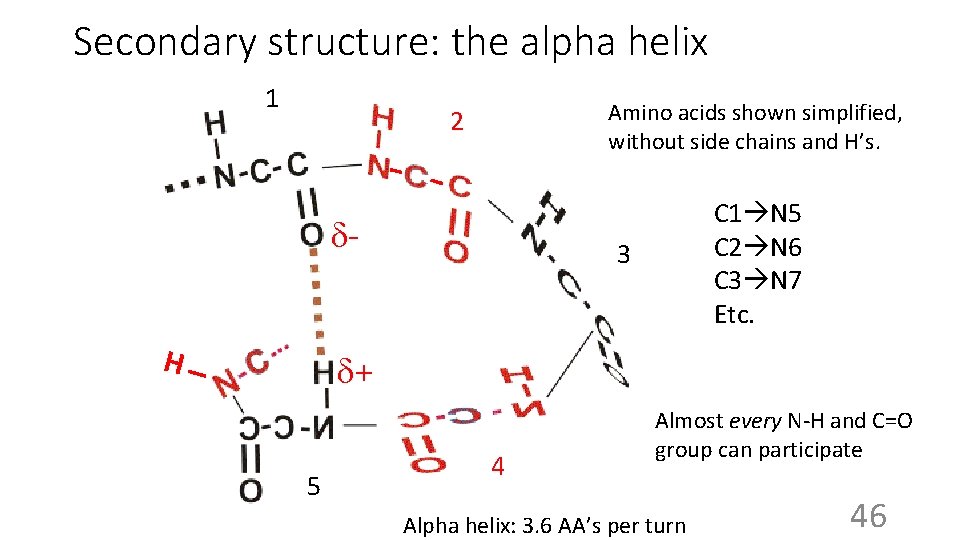
Secondary structure: the alpha helix 1 Amino acids shown simplified, without side chains and H’s. 2 H C 1 N 5 C 2 N 6 C 3 N 7 Etc. 3 + 5 4 Almost every N-H and C=O group can participate Alpha helix: 3. 6 AA’s per turn 46
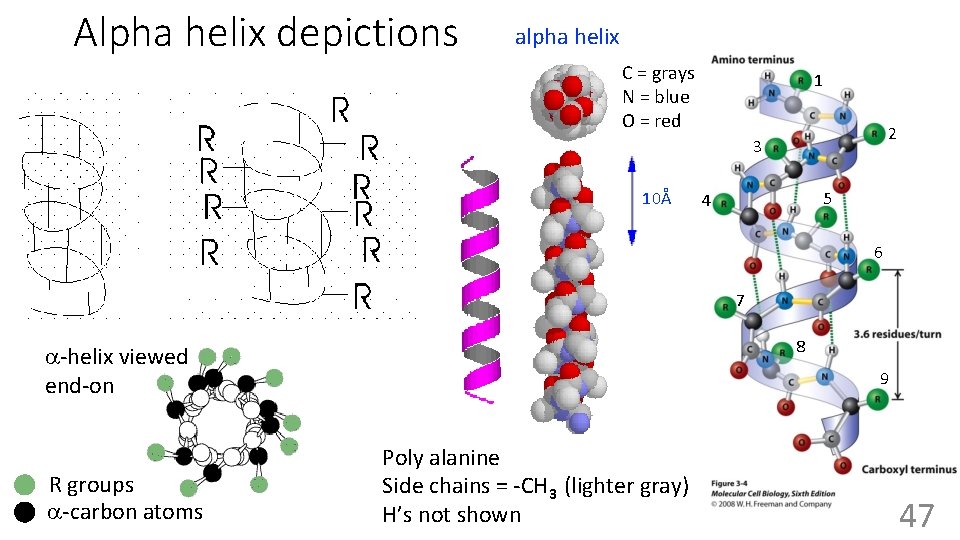
Alpha helix depictions alpha helix C = grays N = blue O = red 1 2 3 10Å 5 4 6 7 8 a-helix viewed end-on R groups a-carbon atoms 9 Poly alanine Side chains = -CH 3 (lighter gray) H’s not shown 47

Linus Pauling and a model of the a helix Linus Pauling and the alpha helix 1963 48
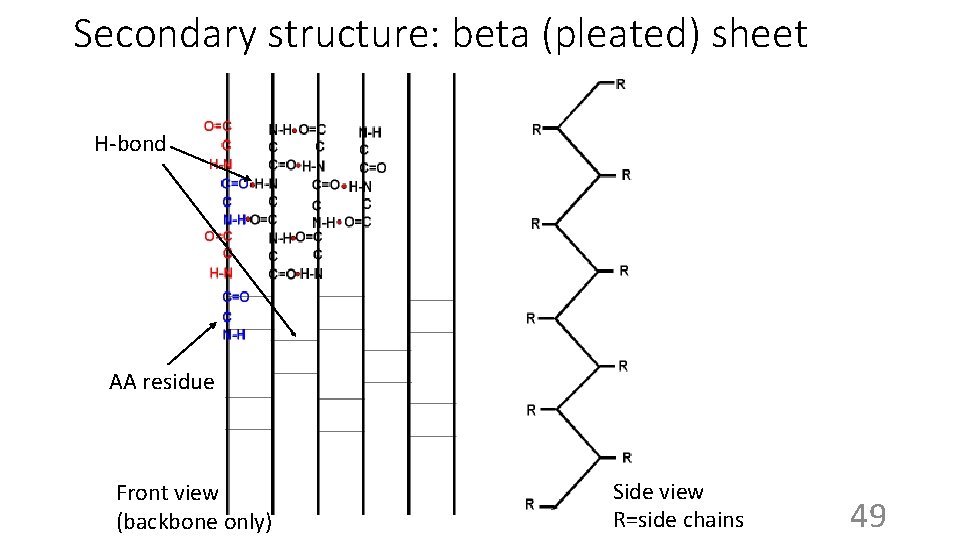
Secondary structure: beta (pleated) sheet H-bond AA residue Front view (backbone only) Side view R=side chains 49
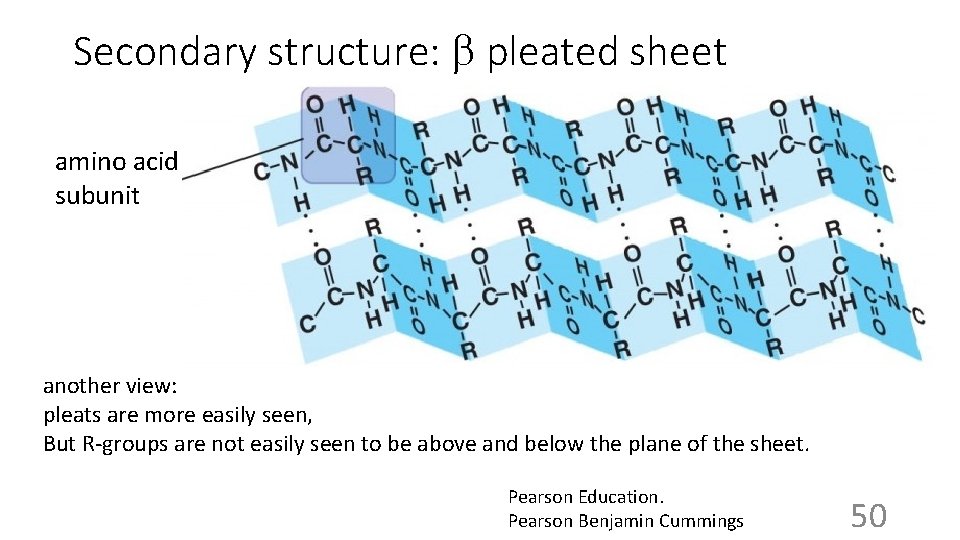
Secondary structure: b pleated sheet amino acid subunit another view: pleats are more easily seen, But R-groups are not easily seen to be above and below the plane of the sheet. Pearson Education. Pearson Benjamin Cummings 50
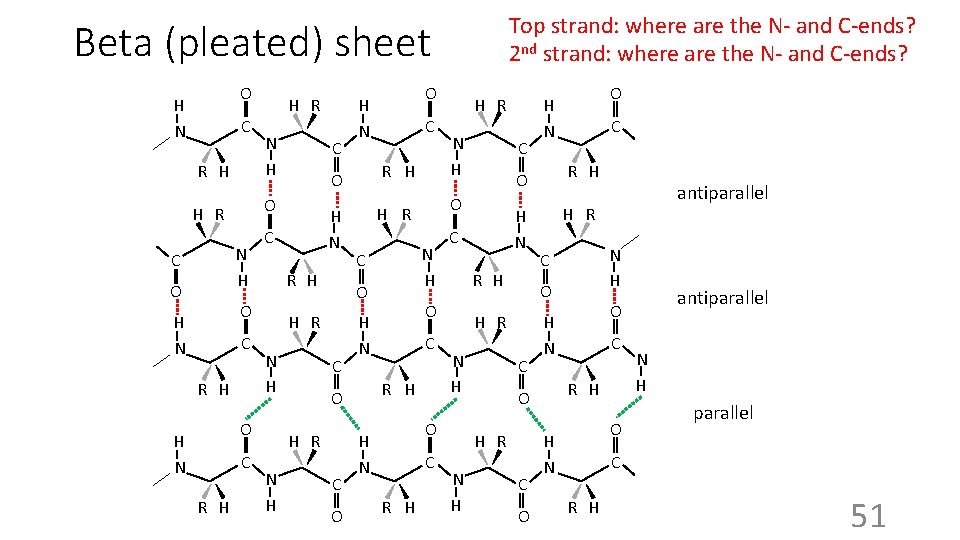
Top strand: where are the N- and C-ends? 2 nd strand: where are the N- and C-ends? Beta (pleated) sheet H N O C R H N H C R H C H R C C O C R H H N H R N H C C R H C O antiparallel H R N H C antiparallel O H N C R H O H R N H O H N O R H O H N C C O H N O H R N H R H N H O H R O H N C O H N O O H R C C C H R O R H N H O H N C H R O O H N N H parallel C R H 51
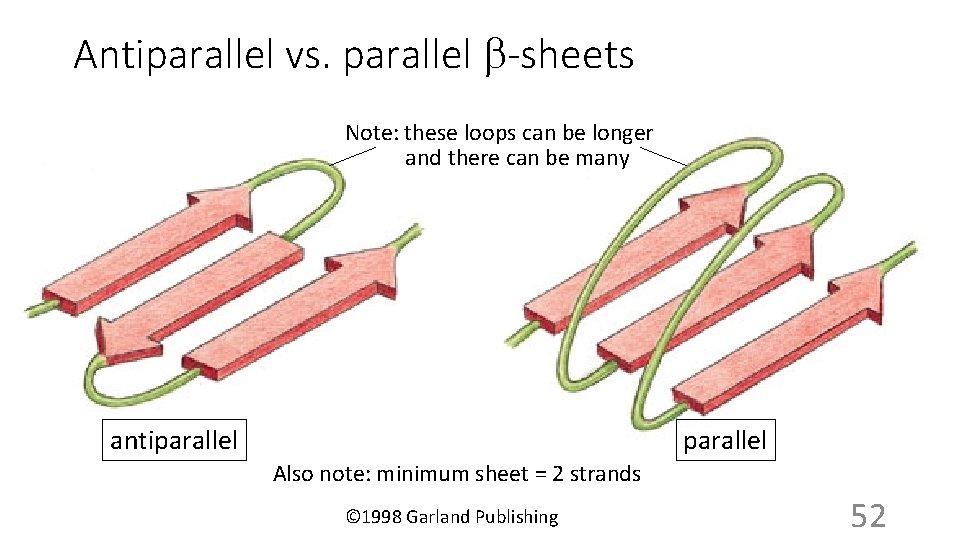
Antiparallel vs. parallel b-sheets Note: these loops can be longer and there can be many antiparallel Also note: minimum sheet = 2 strands © 1998 Garland Publishing 52

Secondary structure (my definition) • Structure produced by regular repeated interactions between atoms of the backbone • If these are backbone atoms • What type of interaction/bond are we talking about? • Which atoms are forming them? 53
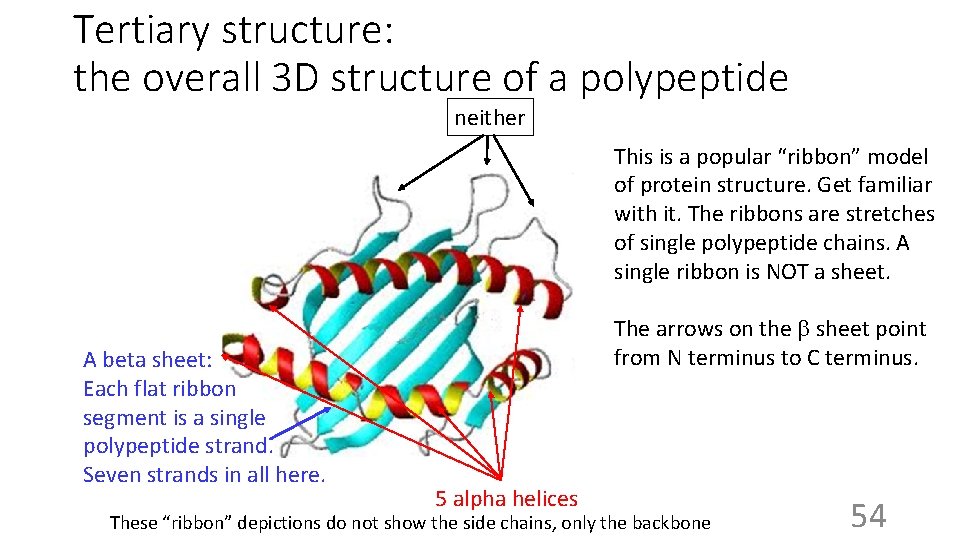
Tertiary structure: the overall 3 D structure of a polypeptide neither This is a popular “ribbon” model of protein structure. Get familiar with it. The ribbons are stretches of single polypeptide chains. A single ribbon is NOT a sheet. A beta sheet: Each flat ribbon segment is a single polypeptide strand. Seven strands in all here. The arrows on the b sheet point from N terminus to C terminus. 5 alpha helices These “ribbon” depictions do not show the side chains, only the backbone 54
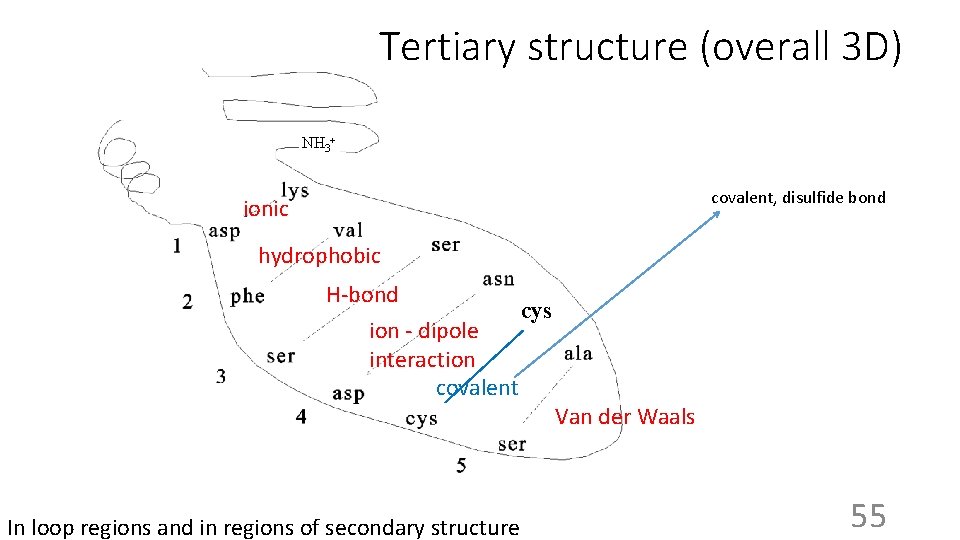
Tertiary structure (overall 3 D) NH 3+ covalent, disulfide bond ionic hydrophobic H-bond cys ion - dipole interaction covalent In loop regions and in regions of secondary structure Van der Waals 55
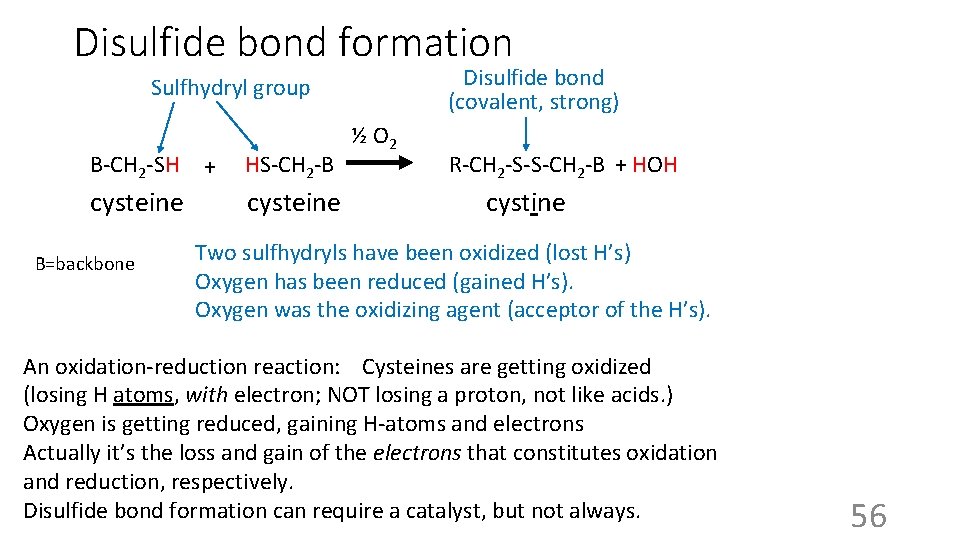
Disulfide bond formation Disulfide bond (covalent, strong) Sulfhydryl group B-CH 2 -SH cysteine B=backbone + HS-CH 2 -B cysteine ½ O 2 R-CH 2 -S-S-CH 2 -B + HOH cystine Two sulfhydryls have been oxidized (lost H’s) Oxygen has been reduced (gained H’s). Oxygen was the oxidizing agent (acceptor of the H’s). An oxidation-reduction reaction: Cysteines are getting oxidized (losing H atoms, with electron; NOT losing a proton, not like acids. ) Oxygen is getting reduced, gaining H-atoms and electrons Actually it’s the loss and gain of the electrons that constitutes oxidation and reduction, respectively. Disulfide bond formation can require a catalyst, but not always. 56
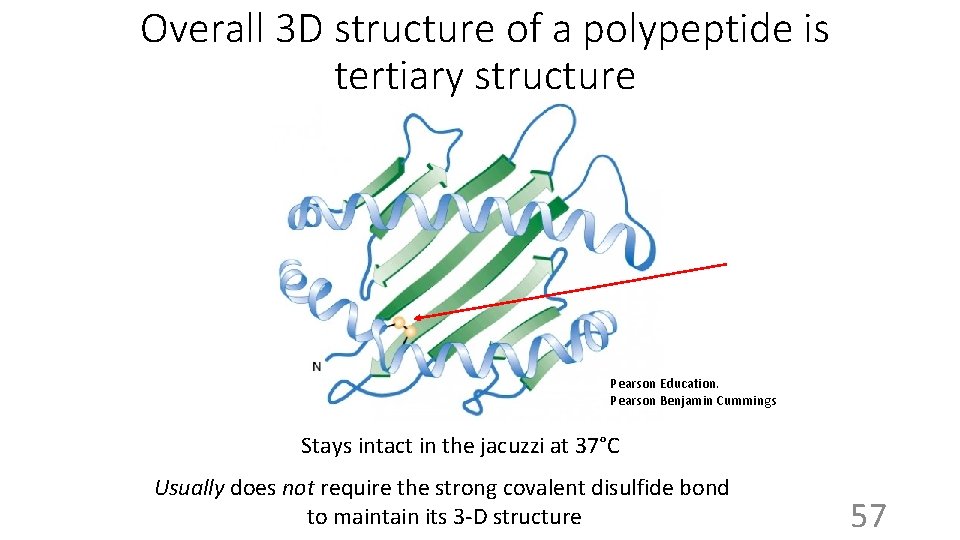
Overall 3 D structure of a polypeptide is tertiary structure Pearson Education. Pearson Benjamin Cummings Stays intact in the jacuzzi at 37°C Usually does not require the strong covalent disulfide bond to maintain its 3 -D structure 57
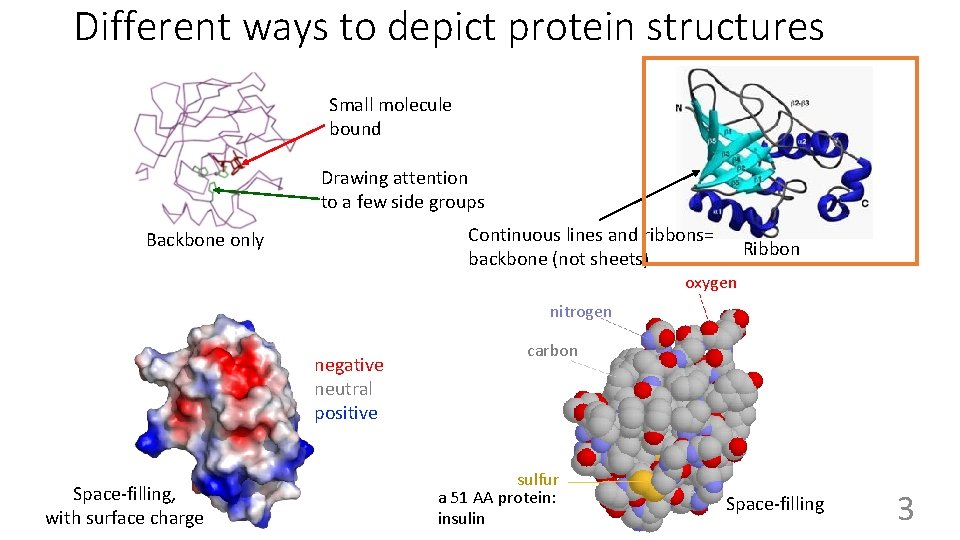
Different ways to depict protein structures Small molecule bound Drawing attention to a few side groups Continuous lines and ribbons= backbone (not sheets) Backbone only Ribbon oxygen nitrogen negative neutral positive Space-filling, with surface 58 charge carbon sulfur a 51 AA protein: insulin Space-filling 3
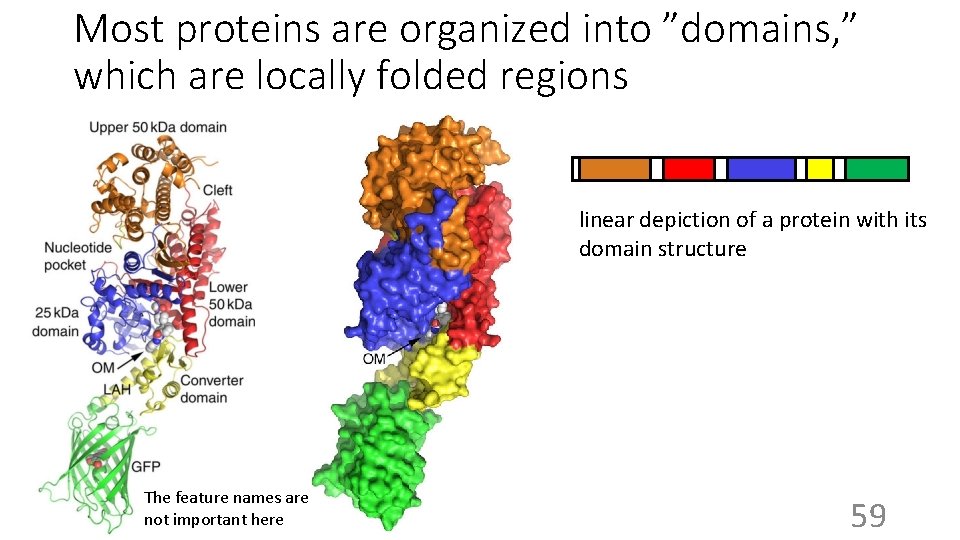
Most proteins are organized into ”domains, ” which are locally folded regions linear depiction of a protein with its domain structure The feature names are not important here 59
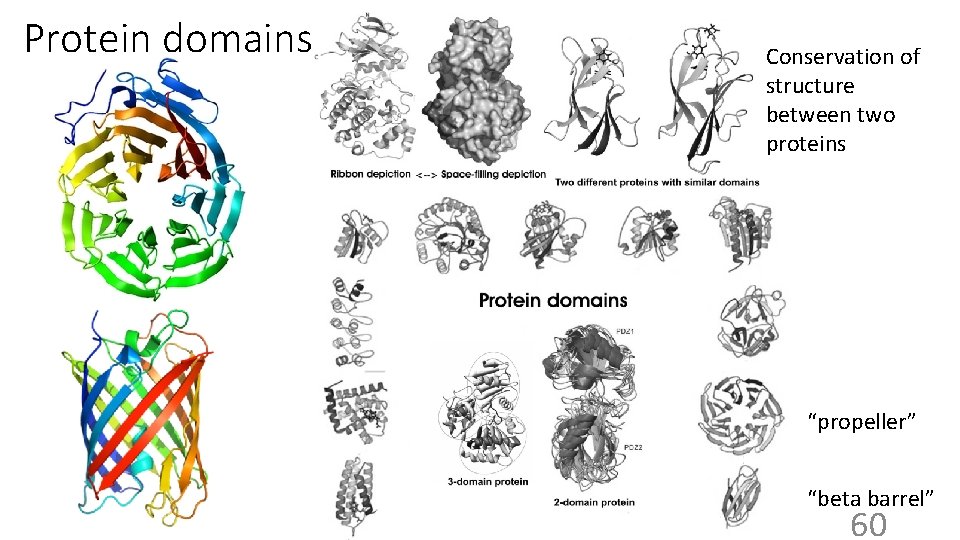
Protein domains Conservation of structure between two proteins “propeller” “beta barrel” 60

Quaternary Structure (4°): The association of more than one polypeptide chain in a protein Monomeric protein (no quaternary structure) Dimeric protein (a homodimer) Dimeric protein (a heterodimer) A heterotetramer A heteropolymeric protein (large one) Also called: multimeric proteins The individual polypeptides in a multimeric protein are called subunits. 61
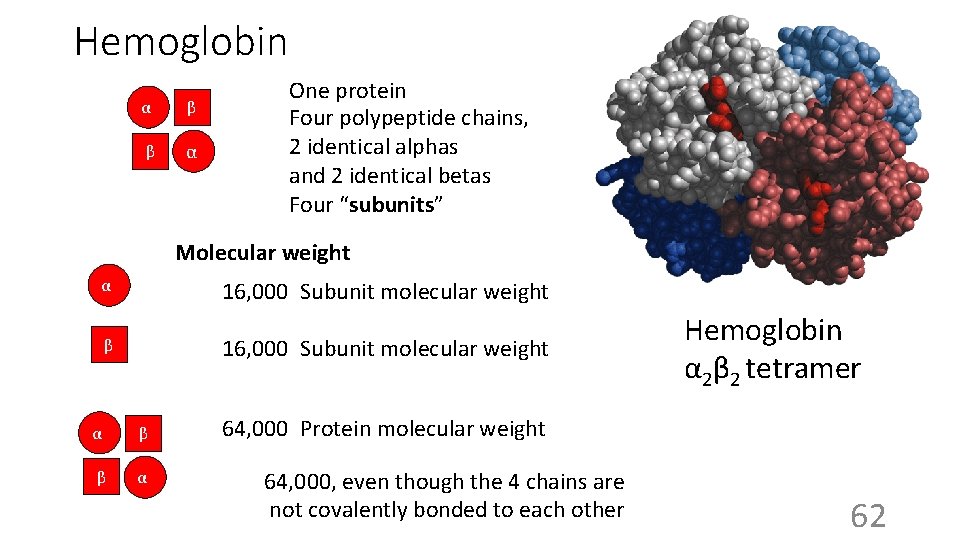
Hemoglobin α β β α One protein Four polypeptide chains, 2 identical alphas and 2 identical betas Four “subunits” Molecular weight α 16, 000 Subunit molecular weight β α β β α Hemoglobin α 2β 2 tetramer 64, 000 Protein molecular weight 64, 000, even though the 4 chains are not covalently bonded to each other 62
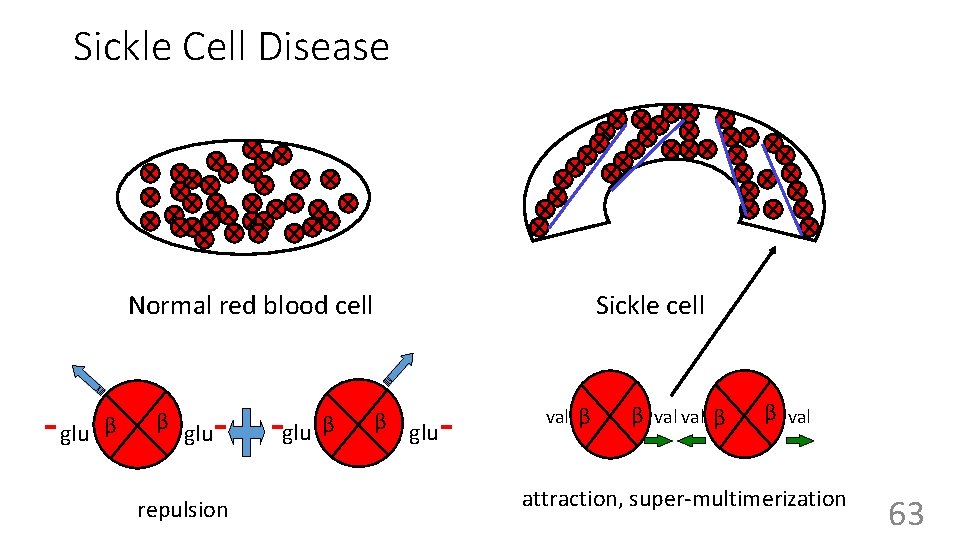
Sickle Cell Disease Normal red blood cell glu β β glu repulsion glu β Sickle cell β glu val β β val attraction, super-multimerization 63
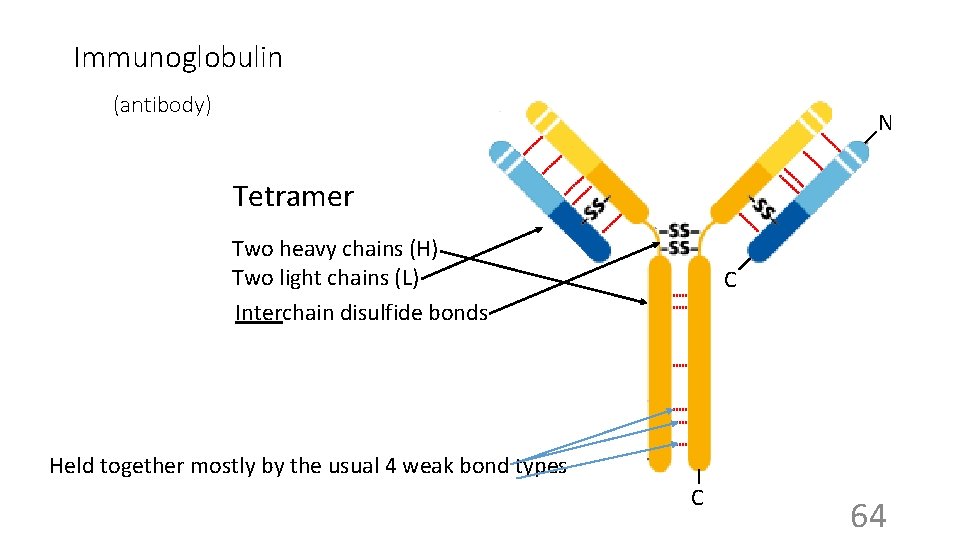
Immunoglobulin (antibody) N Tetramer Two heavy chains (H) Two light chains (L) Interchain disulfide bonds C Held together mostly by the usual 4 weak bond types C 64
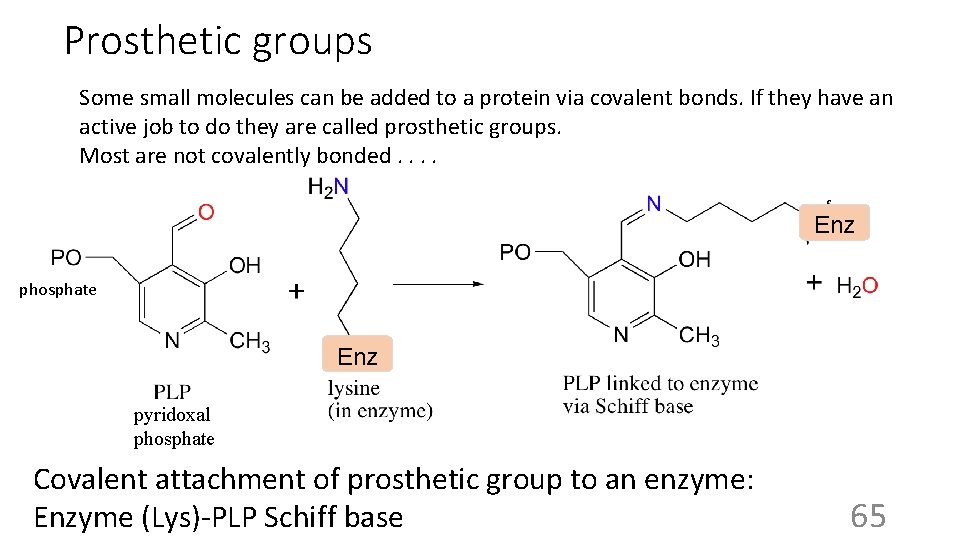
Prosthetic groups Some small molecules can be added to a protein via covalent bonds. If they have an active job to do they are called prosthetic groups. Most are not covalently bonded. . Enz phosphate Enz pyridoxal phosphate Covalent attachment of prosthetic group to an enzyme: Enzyme (Lys)-PLP Schiff base 65
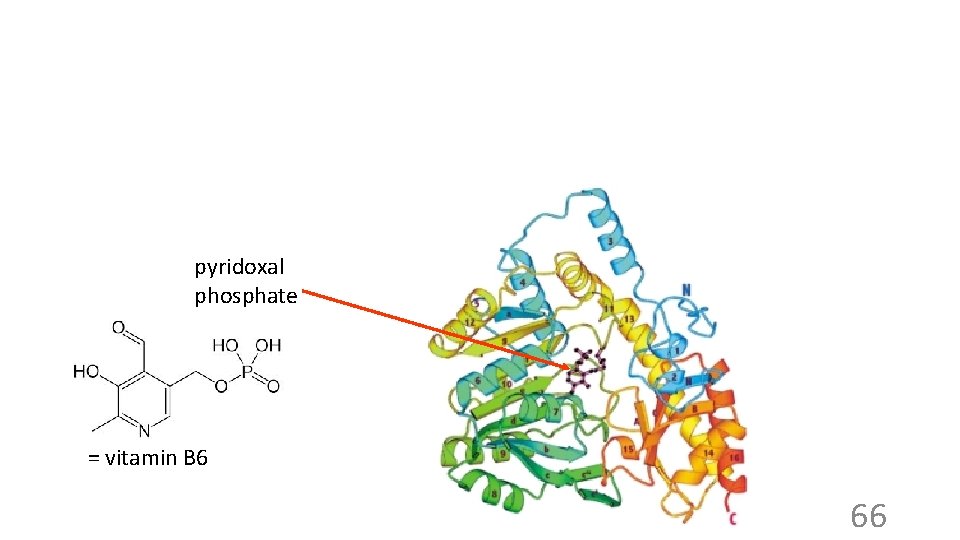
pyridoxal phosphate = vitamin B 6 66
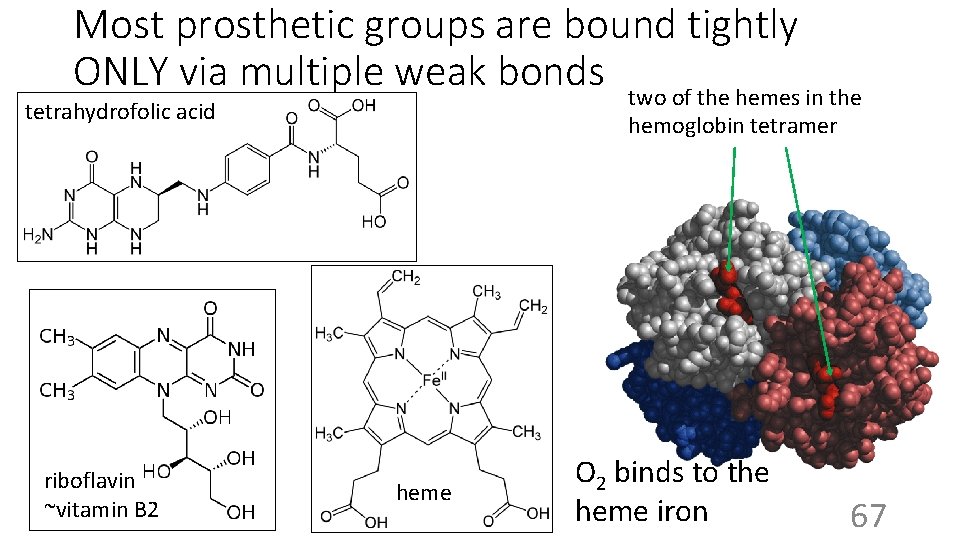
Most prosthetic groups are bound tightly ONLY via multiple weak bonds two of the hemes in the tetrahydrofolic acid riboflavin ~vitamin B 2 hemoglobin tetramer heme O 2 binds to the heme iron 67
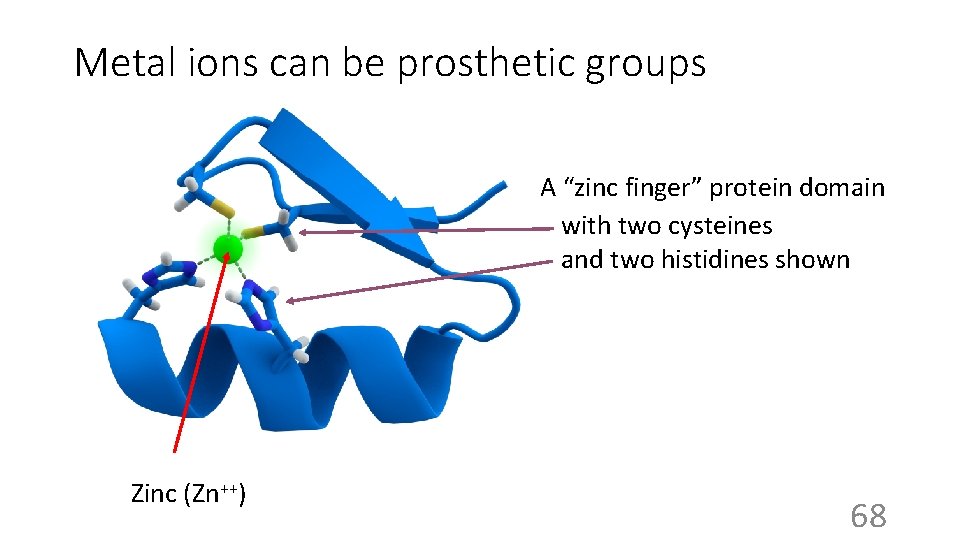
Metal ions can be prosthetic groups A “zinc finger” protein domain with two cysteines and two histidines shown Zinc (Zn++) 68

Covalent protein modifications Acetylation: CH 3 -C- joined to free amino groups (e. g. , lys and N-terminus) to form an amide. O Methylation: Methyl group added to lys amino or arg guanidino. Glycosylation: sugars can be added to hydroxyls (ser, thr) or and amide (asn) And especially: Phosphorylation: phosphate added to hydroxyls (ser, thr, tyr) to form an ester. Modifies the properties of the protein. Turns a protein’s function on or off. Covered next semester. 69
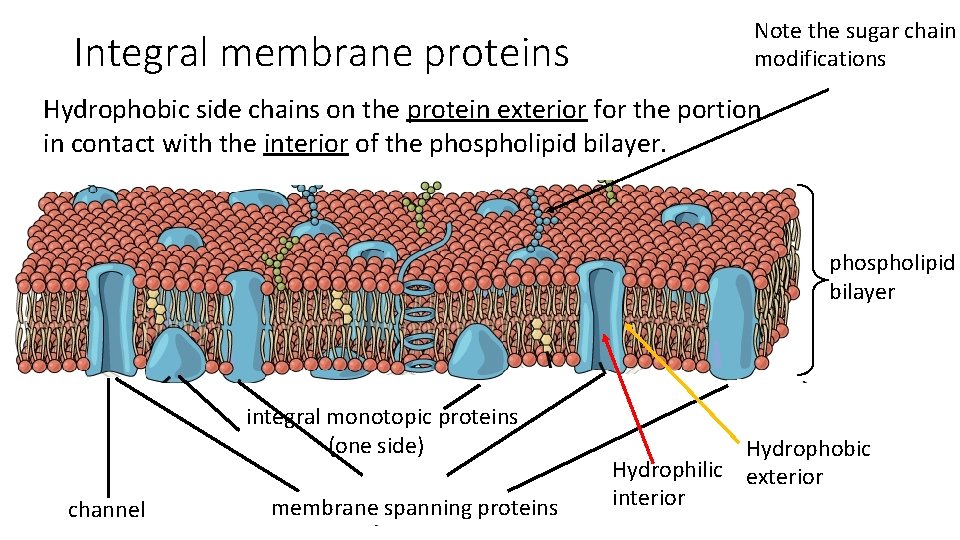
Integral membrane proteins Note the sugar chain modifications Hydrophobic side chains on the protein exterior for the portion in contact with the interior of the phospholipid bilayer integral monotopic proteins (one side) channel membrane spanning proteins Hydrophobic Hydrophilic exterior interior 70
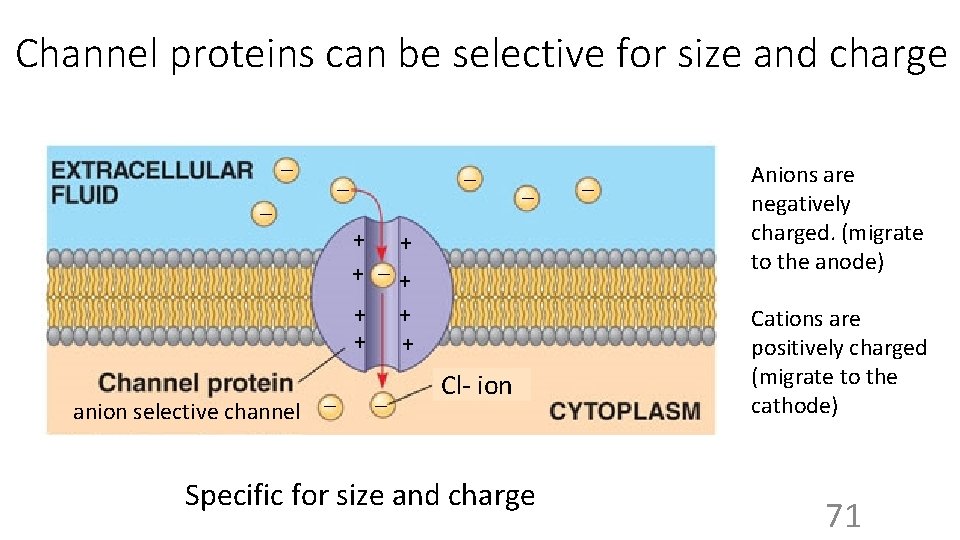
Channel proteins can be selective for size and charge _ _ _ + + + _+ + + anion selective channel _ _ Cl- ion Specific for size and charge _ Anions are negatively charged. (migrate to the anode) Cations are positively charged (migrate to the cathode) 71
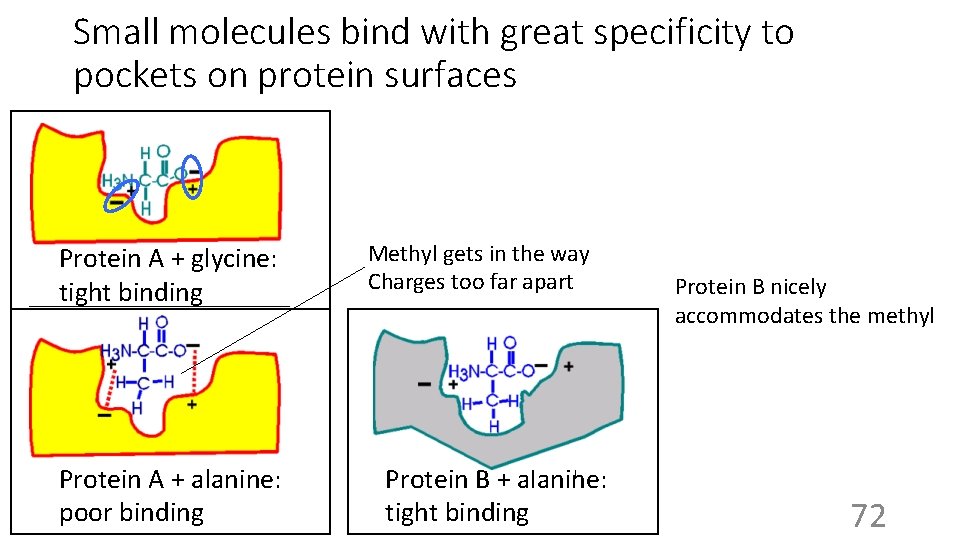
Small molecules bind with great specificity to pockets on protein surfaces Protein A + glycine: tight binding Protein A + alanine: poor binding Methyl gets in the way Charges too far apart B + alanine: BProtein nicely accommodates the methyl tight binding Protein B nicely accommodates the methyl 72
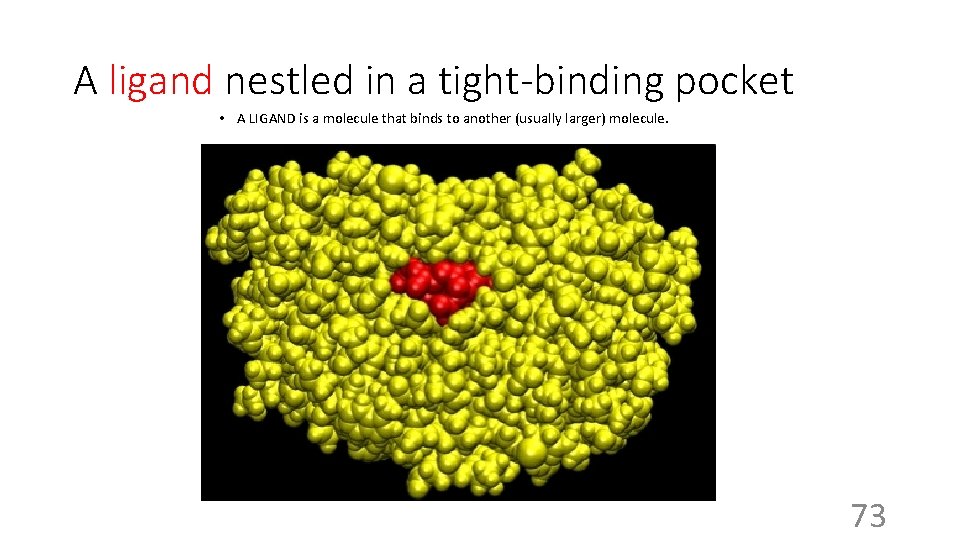
A ligand nestled in a tight-binding pocket • A LIGAND is a molecule that binds to another (usually larger) molecule. 73
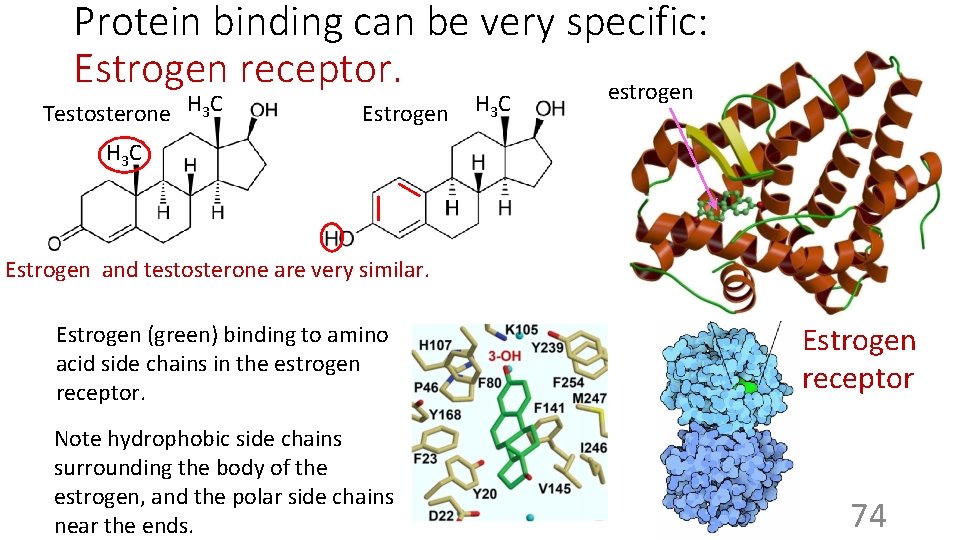
Protein binding can be very specific: Estrogen receptor. estrogen Testosterone H 3 C Estrogen H 3 C Estrogen and testosterone are very similar. Estrogen (green) binding to amino acid side chains in the estrogen receptor. Note hydrophobic side chains surrounding the body of the estrogen, and the polar side chains near the ends. Estrogen receptor 74
- Slides: 74