Interpretation of Agilent 2100 Bioanalyzer Data Intact Total

Interpretation of Agilent 2100 Bioanalyzer Data
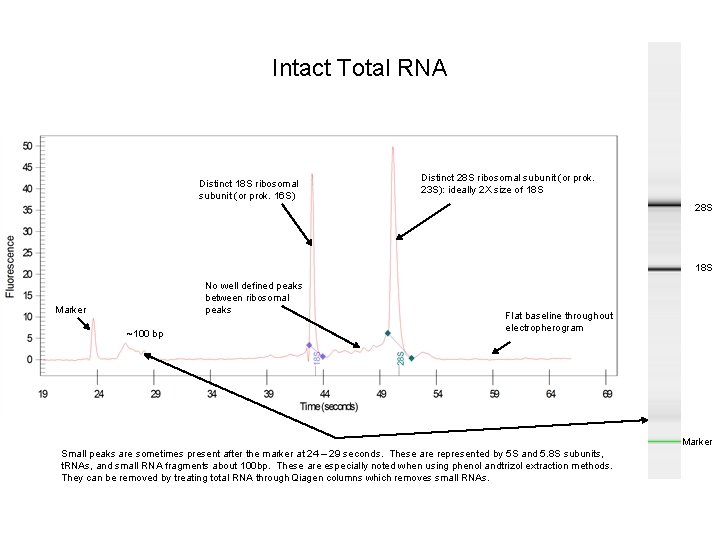
Intact Total RNA Distinct 18 S ribosomal subunit (or prok. 16 S) Distinct 28 S ribosomal subunit (or prok. 23 S): ideally 2 X size of 18 S 28 S 18 S No well defined peaks between ribosomal peaks Marker ~100 bp Flat baseline throughout electropherogram Marker Small peaks are sometimes present after the marker at 24 – 29 seconds. These are represented by 5 S and 5. 8 S subunits, t. RNAs, and small RNA fragments about 100 bp. These are especially noted when using phenol andtrizol extraction methods. They can be removed by treating total RNA through Qiagen columns which removes small RNAs.
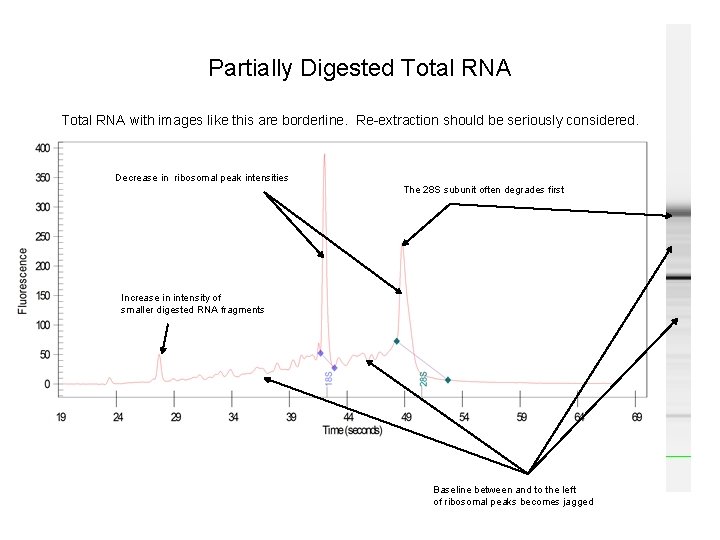
Partially Digested Total RNA with images like this are borderline. Re-extraction should be seriously considered. Decrease in ribosomal peak intensities The 28 S subunit often degrades first Increase in intensity of smaller digested RNA fragments Baseline between and to the left of ribosomal peaks becomes jagged
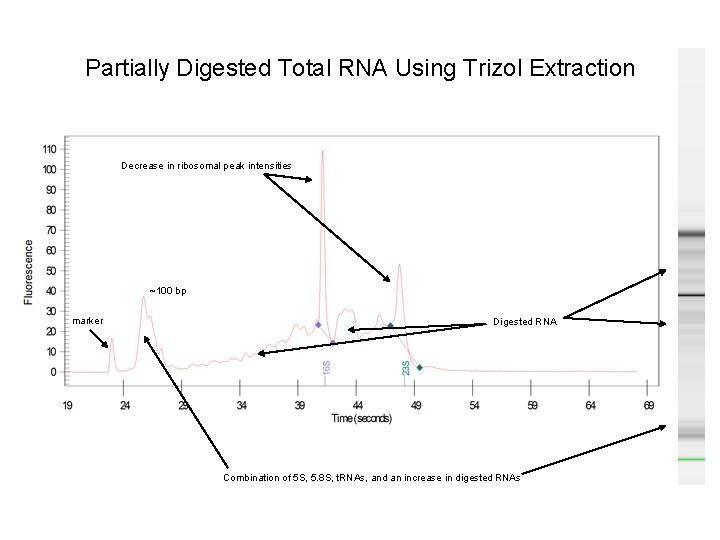
Partially Digested Total RNA Using Trizol Extraction Decrease in ribosomal peak intensities ~100 bp marker Digested RNA Combination of 5 S, 5. 8 S, t. RNAs, and an increase in digested RNAs
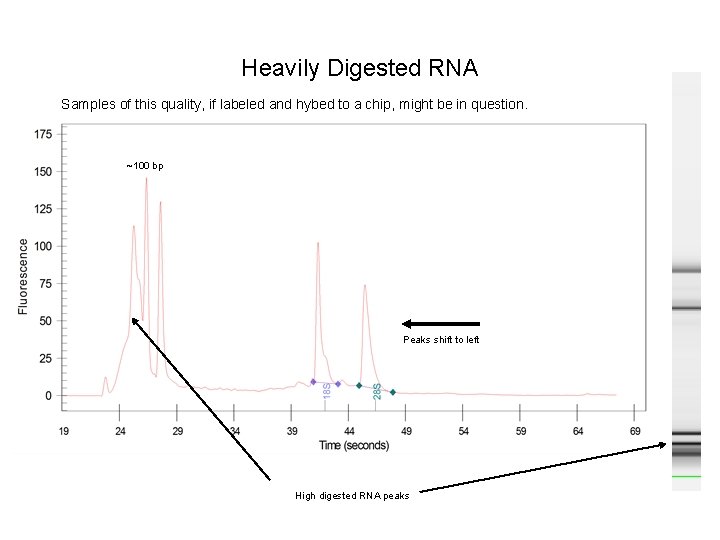
Heavily Digested RNA Samples of this quality, if labeled and hybed to a chip, might be in question. ~100 bp Peaks shift to left High digested RNA peaks
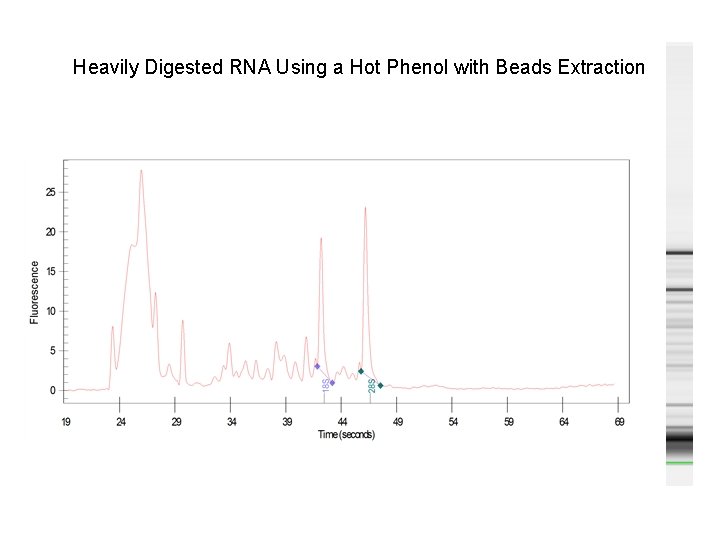
Heavily Digested RNA Using a Hot Phenol with Beads Extraction
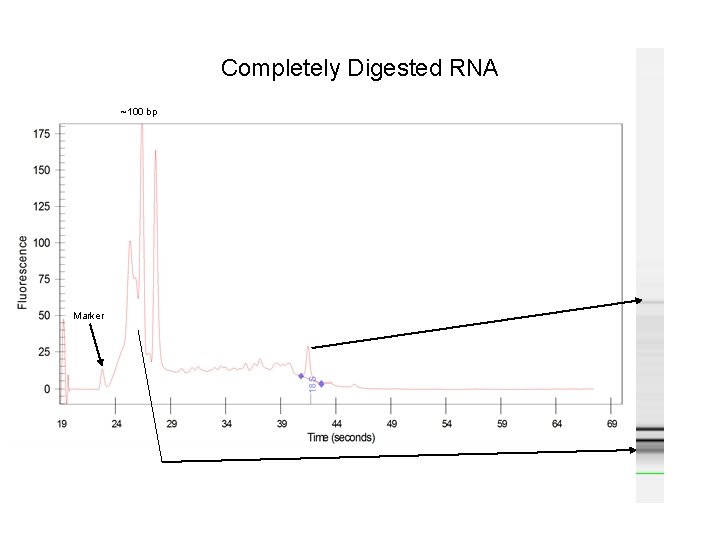
Completely Digested RNA ~100 bp Marker
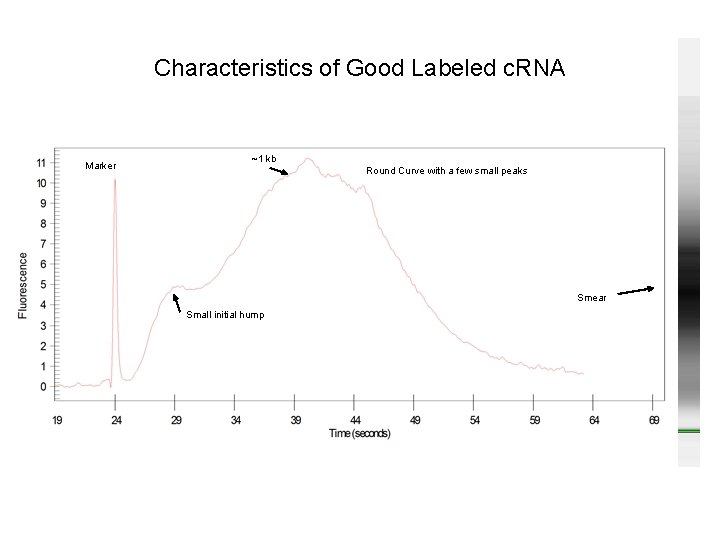
Characteristics of Good Labeled c. RNA Marker ~1 kb Round Curve with a few small peaks Smear Small initial hump
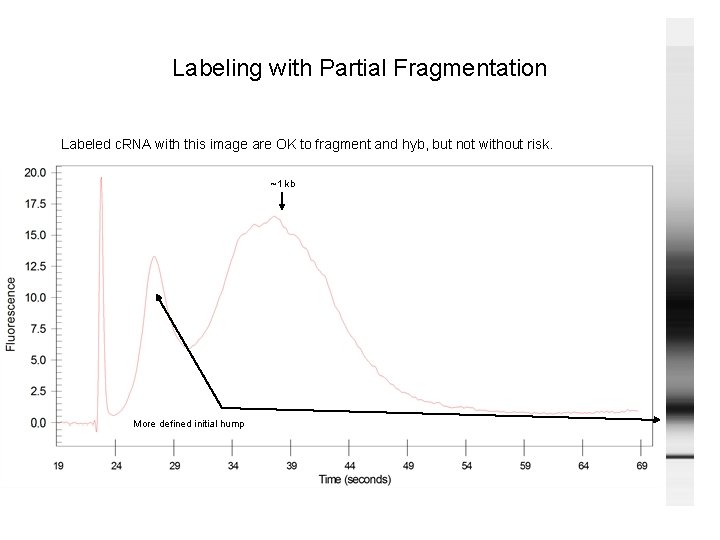
Labeling with Partial Fragmentation Labeled c. RNA with this image are OK to fragment and hyb, but not without risk. ~1 kb More defined initial hump
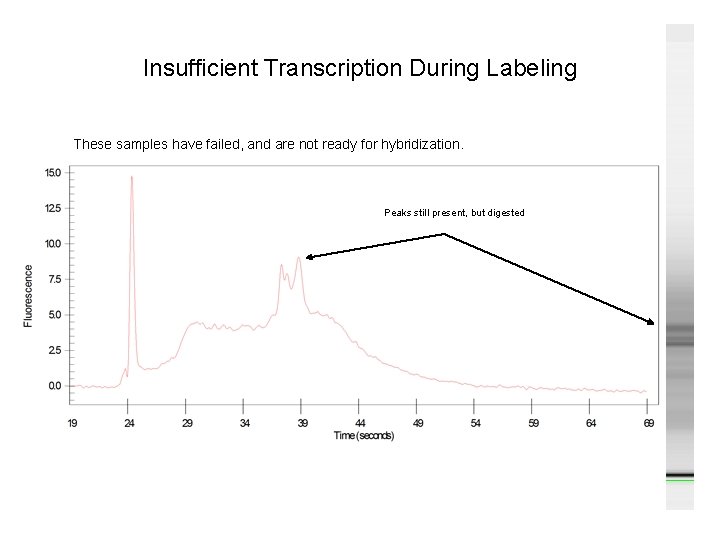
Insufficient Transcription During Labeling These samples have failed, and are not ready for hybridization. Peaks still present, but digested
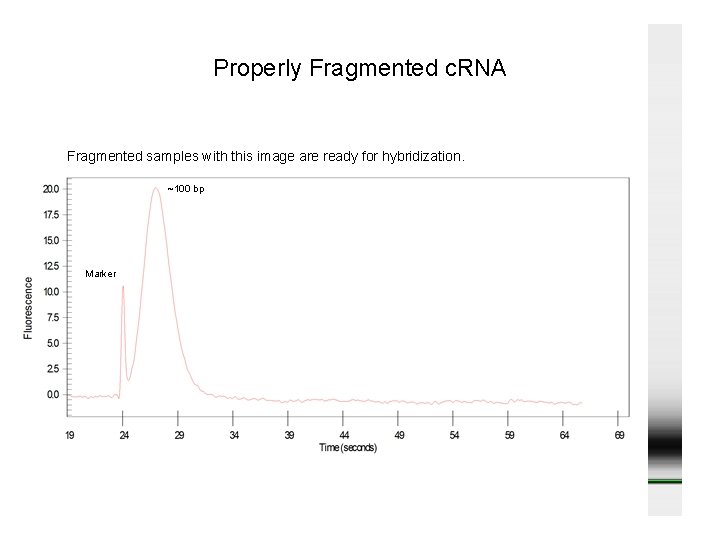
Properly Fragmented c. RNA Fragmented samples with this image are ready for hybridization. ~100 bp Marker
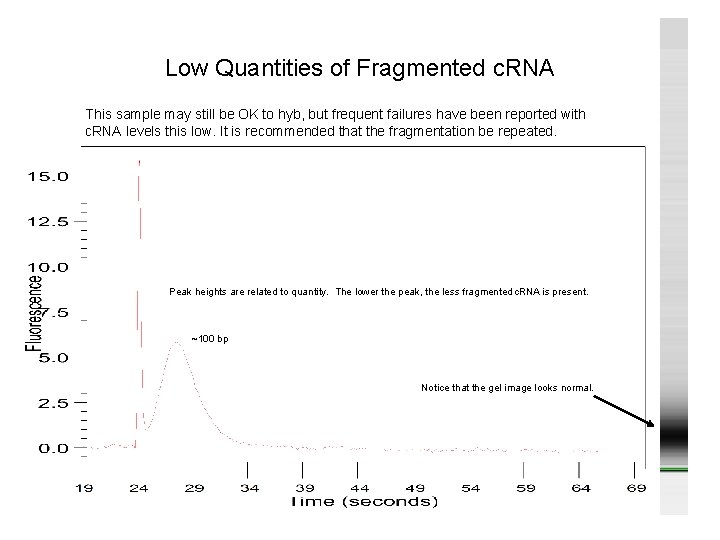
Low Quantities of Fragmented c. RNA This sample may still be OK to hyb, but frequent failures have been reported with c. RNA levels this low. It is recommended that the fragmentation be repeated. Peak heights are related to quantity. The lower the peak, the less fragmented c. RNA is present. ~100 bp Notice that the gel image looks normal.
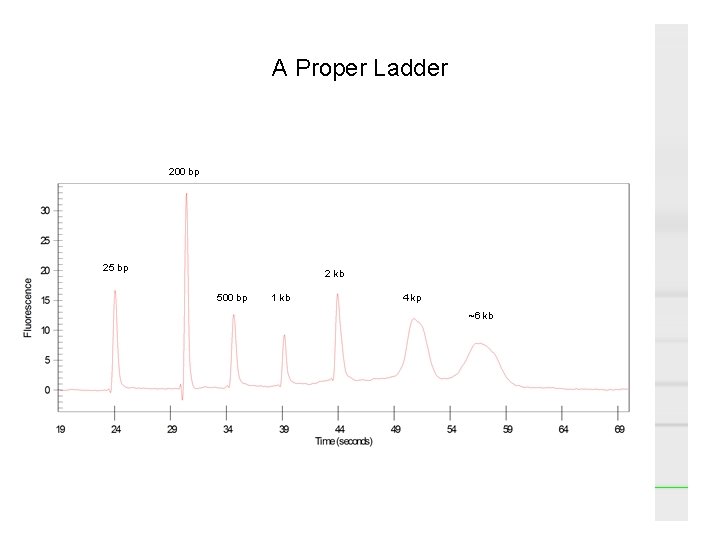
A Proper Ladder 200 bp 25 bp 2 kb 500 bp 1 kb 4 kp ~6 kb
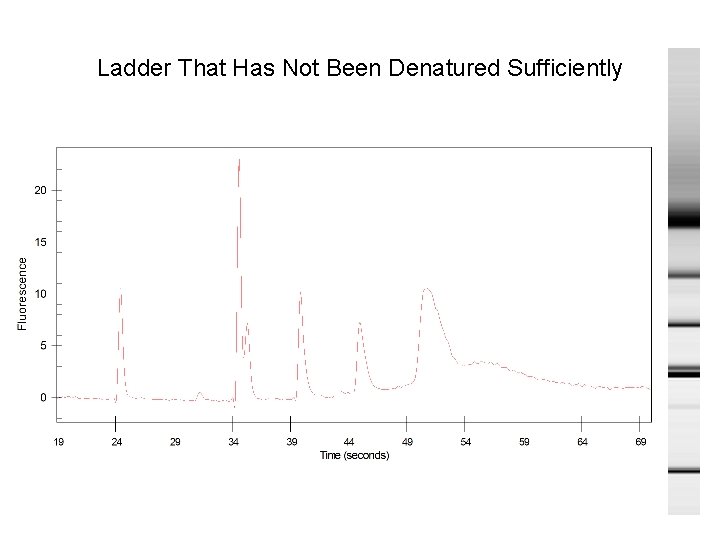
Ladder That Has Not Been Denatured Sufficiently
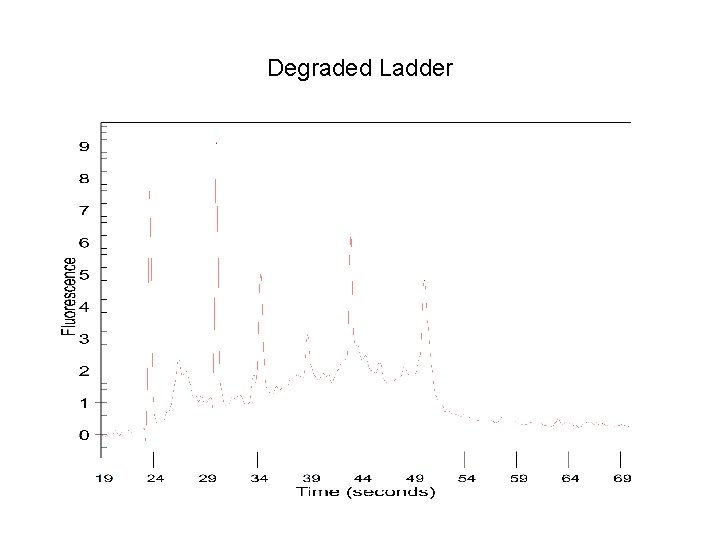
Degraded Ladder
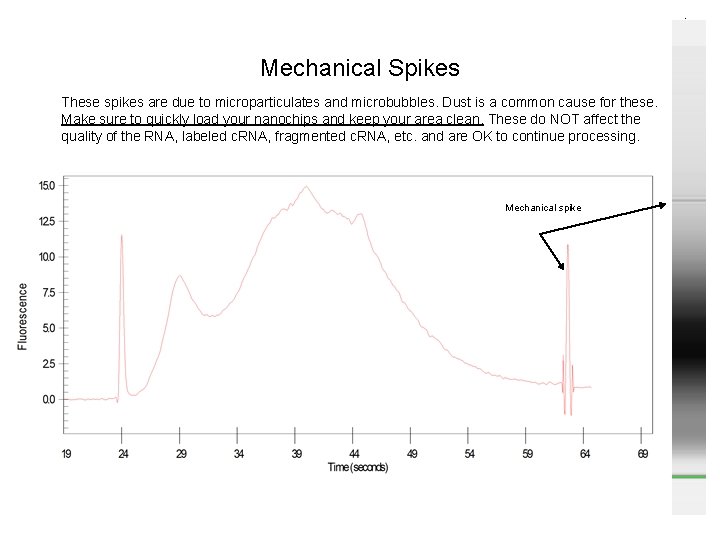
Mechanical Spikes These spikes are due to microparticulates and microbubbles. Dust is a common cause for these. Make sure to quickly load your nanochips and keep your area clean. These do NOT affect the quality of the RNA, labeled c. RNA, fragmented c. RNA, etc. and are OK to continue processing. Mechanical spike
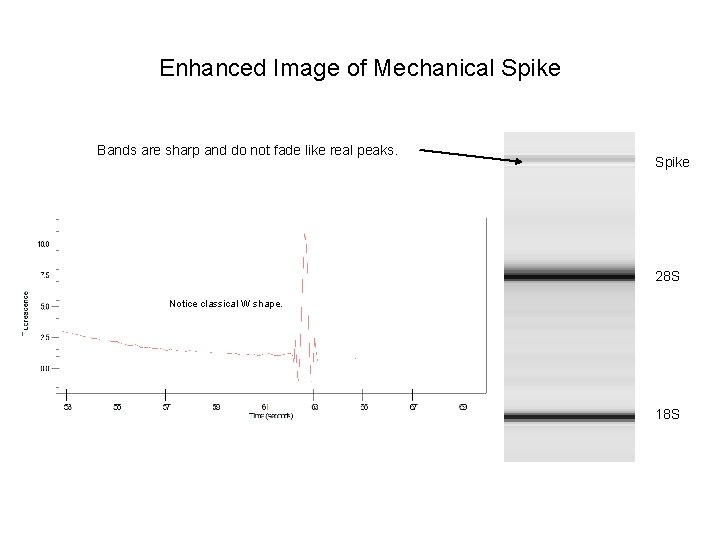
Enhanced Image of Mechanical Spike Bands are sharp and do not fade like real peaks. Spike 28 S Notice classical W shape. 18 S
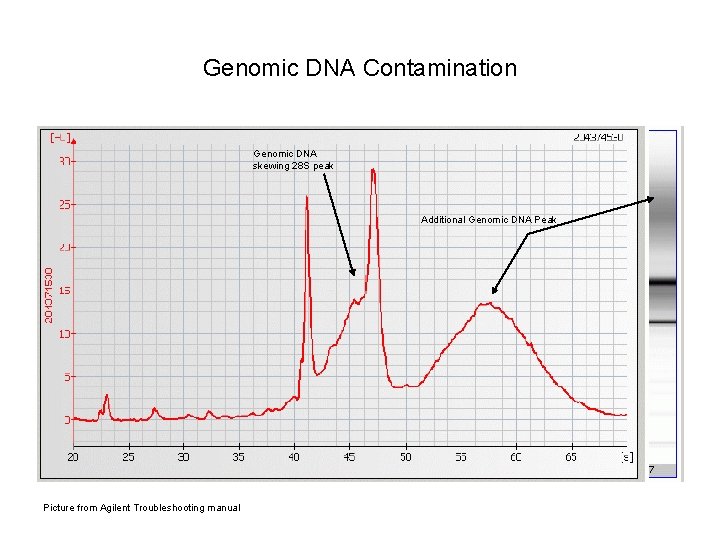
Genomic DNA Contamination Genomic DNA skewing 28 S peak Additional Genomic DNA Peak Picture from Agilent Troubleshooting manual
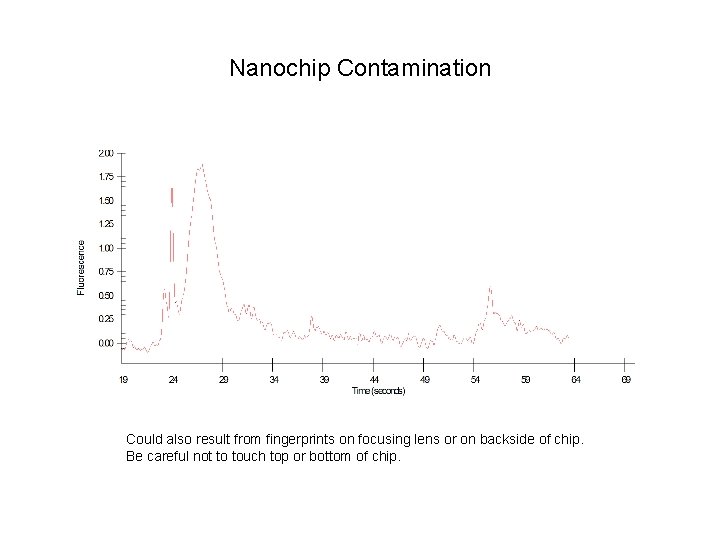
Nanochip Contamination Could also result from fingerprints on focusing lens or on backside of chip. Be careful not to touch top or bottom of chip.
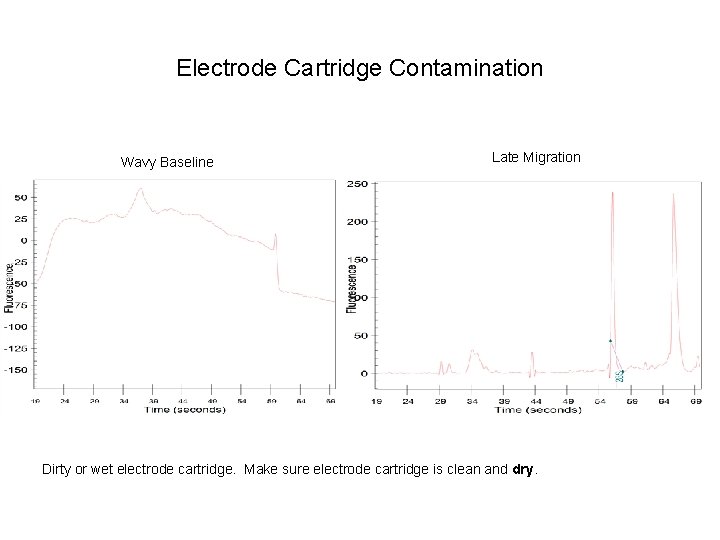
Electrode Cartridge Contamination Wavy Baseline Late Migration Dirty or wet electrode cartridge. Make sure electrode cartridge is clean and dry.
- Slides: 20