Internal Use Only DIA on HRAM Orbitrap Based

Internal Use Only! DIA on HR/AM Orbitrap Based Instruments Demonstrating complete workflows on the Q Exactive Plus and the Orbitrap Fusion Presenter: Reiko Kiyonami, Ph. D. Sales webinar, San Jose, January 21 st, 2014 Contributors: Shannon Eliuk, Vlad Zabrouskov, Amol Prakash, Scott Peterman, Xuan Yue, Maciej Bromirsky, Markus Kellmann Andreas Huhmer
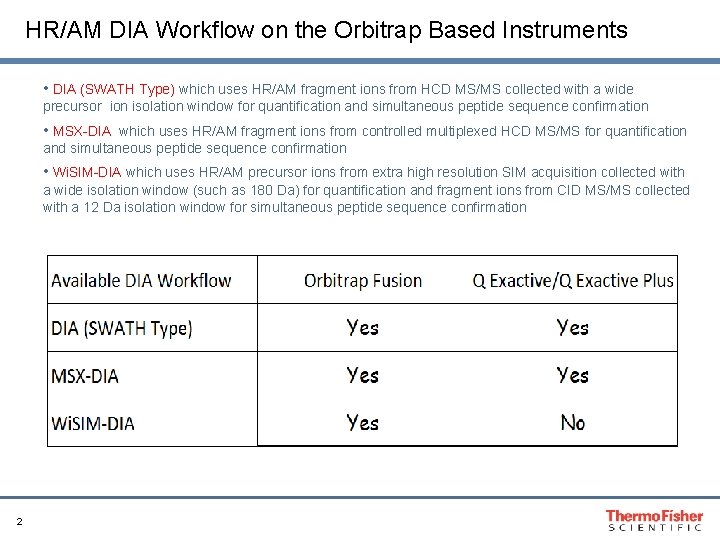
HR/AM DIA Workflow on the Orbitrap Based Instruments • DIA (SWATH Type) which uses HR/AM fragment ions from HCD MS/MS collected with a wide precursor ion isolation window for quantification and simultaneous peptide sequence confirmation • MSX-DIA which uses HR/AM fragment ions from controlled multiplexed HCD MS/MS for quantification and simultaneous peptide sequence confirmation • Wi. SIM-DIA which uses HR/AM precursor ions from extra high resolution SIM acquisition collected with a wide isolation window (such as 180 Da) for quantification and fragment ions from CID MS/MS collected with a 12 Da isolation window for simultaneous peptide sequence confirmation 2

Huge Interests about DIA on Orbitrap Based Instruments 72 ASMS 2013 contents added to Planet Orbitrap. Wi. SIM-DIA on OT Fusion is number 1 in downloads DIA/MSX-DIA on QE is number 3 in downloads 3
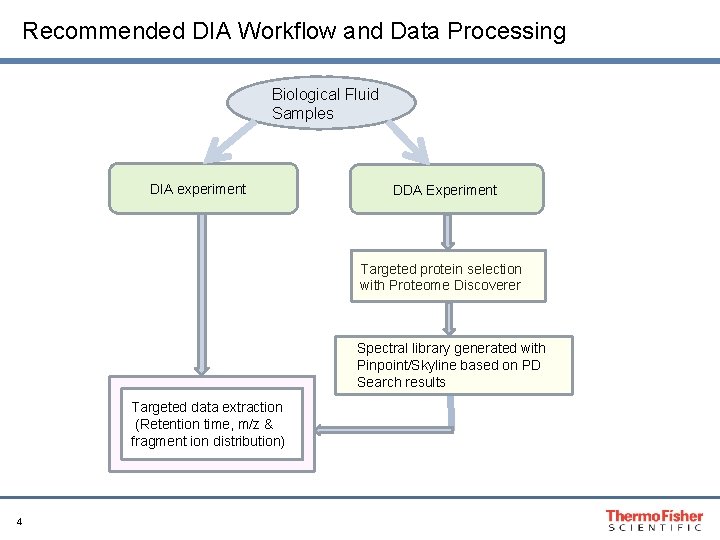
Recommended DIA Workflow and Data Processing Biological Fluid Samples DIA experiment DDA Experiment Targeted protein selection with Proteome Discoverer Spectral library generated with Pinpoint/Skyline based on PD Search results Targeted data extraction (Retention time, m/z & fragment ion distribution) 4
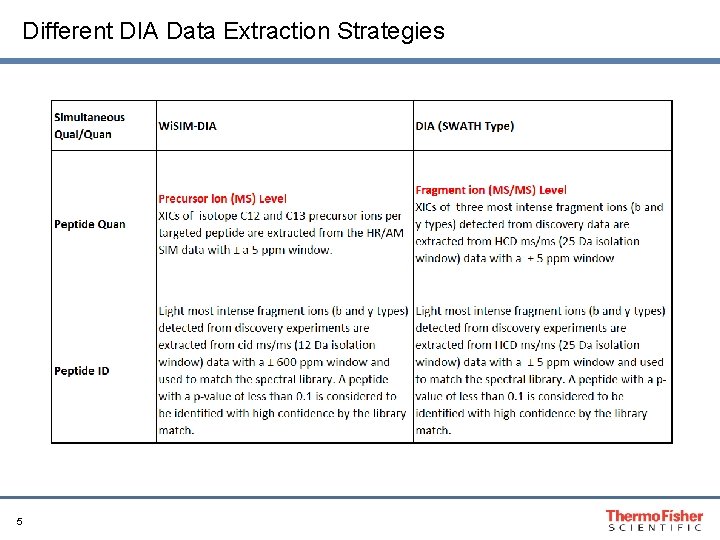
Different DIA Data Extraction Strategies 5
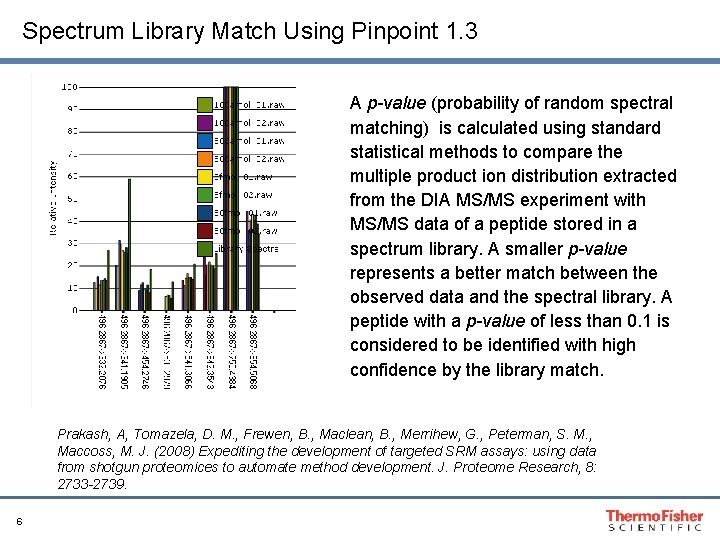
Spectrum Library Match Using Pinpoint 1. 3 A p-value (probability of random spectral matching) is calculated using standard statistical methods to compare the multiple product ion distribution extracted from the DIA MS/MS experiment with MS/MS data of a peptide stored in a spectrum library. A smaller p-value represents a better match between the observed data and the spectral library. A peptide with a p-value of less than 0. 1 is considered to be identified with high confidence by the library match. Prakash, A, Tomazela, D. M. , Frewen, B. , Maclean, B. , Merrihew, G. , Peterman, S. M. , Maccoss, M. J. (2008) Expediting the development of targeted SRM assays: using data from shotgun proteomices to automate method development. J. Proteome Research, 8: 2733 -2739. 6
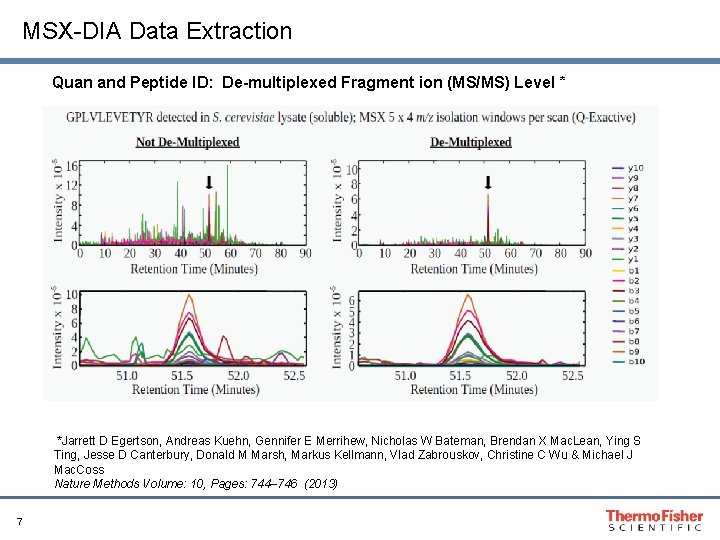
MSX-DIA Data Extraction Quan and Peptide ID: De-multiplexed Fragment ion (MS/MS) Level * *Jarrett D Egertson, Andreas Kuehn, Gennifer E Merrihew, Nicholas W Bateman, Brendan X Mac. Lean, Ying S Ting, Jesse D Canterbury, Donald M Marsh, Markus Kellmann, Vlad Zabrouskov, Christine C Wu & Michael J Mac. Coss Nature Methods Volume: 10, Pages: 744– 746 (2013) 7
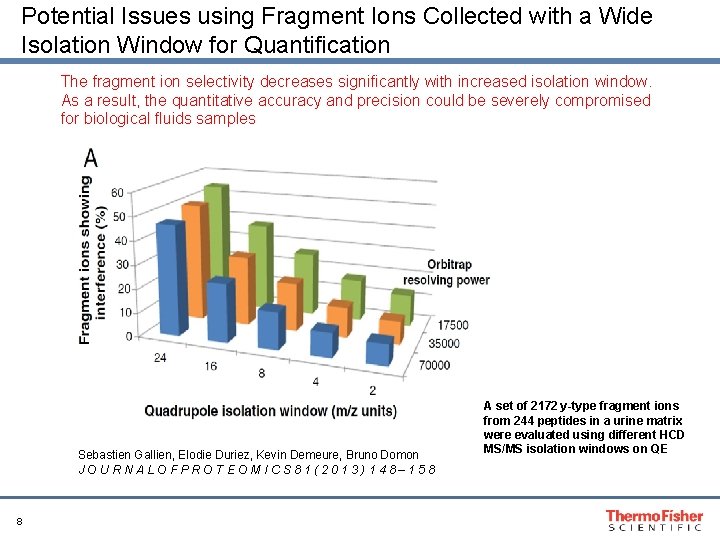
Potential Issues using Fragment Ions Collected with a Wide Isolation Window for Quantification The fragment ion selectivity decreases significantly with increased isolation window. As a result, the quantitative accuracy and precision could be severely compromised for biological fluids samples Sebastien Gallien, Elodie Duriez, Kevin Demeure, Bruno Domon JOURNALOFPROTEOMICS 81(2013)148– 158 8 A set of 2172 y-type fragment ions from 244 peptides in a urine matrix were evaluated using different HCD MS/MS isolation windows on QE
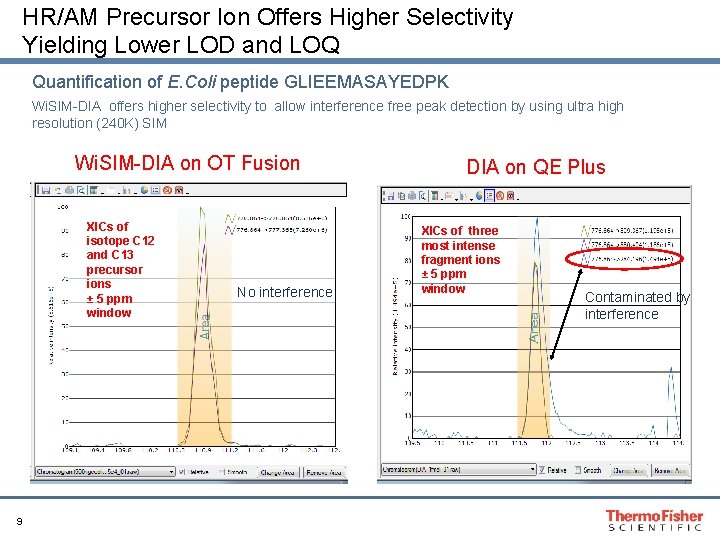
HR/AM Precursor Ion Offers Higher Selectivity Yielding Lower LOD and LOQ Quantification of E. Coli peptide GLIEEMASAYEDPK Wi. SIM-DIA offers higher selectivity to allow interference free peak detection by using ultra high resolution (240 K) SIM Wi. SIM-DIA on OT Fusion XICs of isotope C 12 and C 13 precursor ions ± 5 ppm window 9 No interference DIA on QE Plus XICs of three most intense fragment ions ± 5 ppm window Contaminated by interference
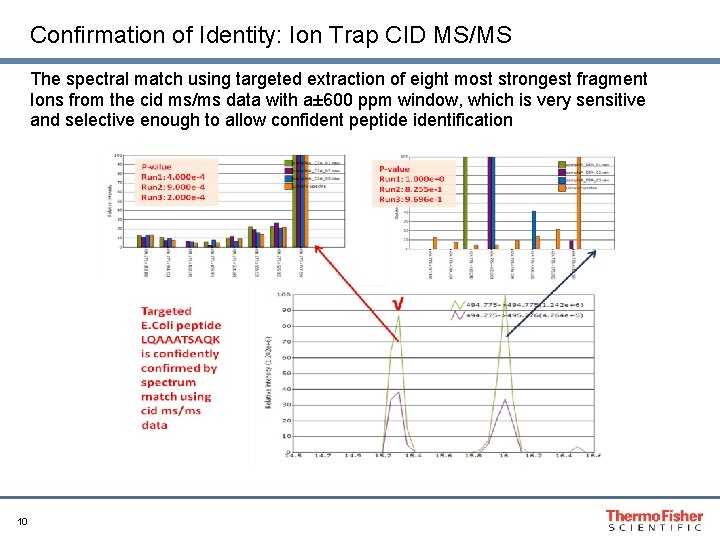
Confirmation of Identity: Ion Trap CID MS/MS The spectral match using targeted extraction of eight most strongest fragment Ions from the cid ms/ms data with a± 600 ppm window, which is very sensitive and selective enough to allow confident peptide identification 10

Quantifying Large Number of Proteins of Interest Using DIA Workflow on the Q Exactive Plus and Wi. SIM-DIA Workflow on the Orbitrap Fusion Instruments The world leader in serving science 11

Objectives • Evaluate the quantitative performances of DIA approach on the QE Plus instrument and Wi. SIM-DIA approach on the OT Fusion instrument to quantify large number of proteins in E. Coli digest using targeted data extraction and spectral library search. • DIA on QE Plus and Wi. SIM-DIA on OT Fusion are used to determine the relative expression ratios of 806 E. Coli proteins between 250 ng E. Coli digest and 500 ng E. Coli digest. 12
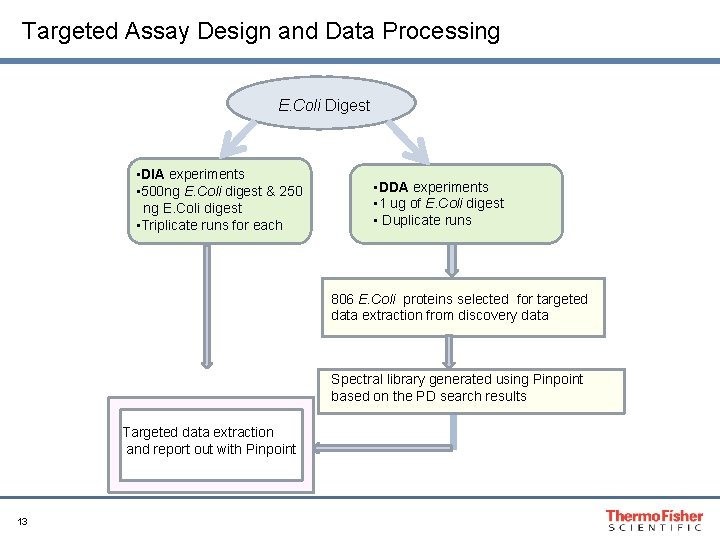
Targeted Assay Design and Data Processing E. Coli Digest • DIA experiments • 500 ng E. Coli digest & 250 ng E. Coli digest • Triplicate runs for each • DDA experiments • 1 ug of E. Coli digest • Duplicate runs 806 E. Coli proteins selected for targeted data extraction from discovery data Spectral library generated using Pinpoint based on the PD search results Targeted data extraction and report out with Pinpoint 13
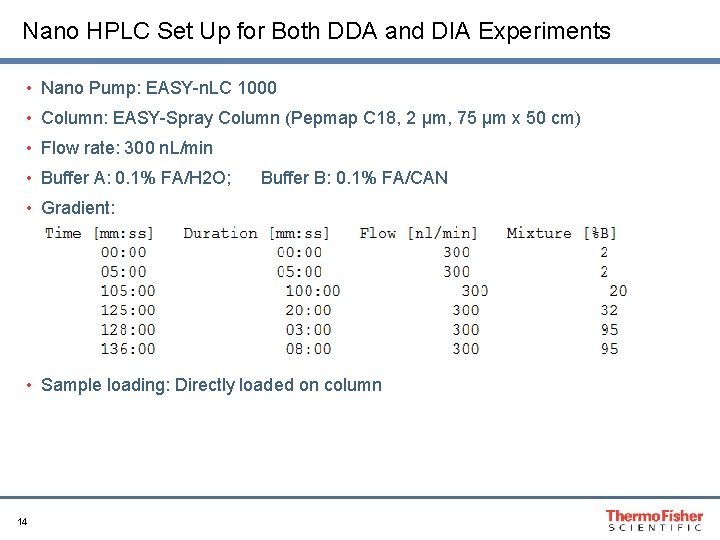
Nano HPLC Set Up for Both DDA and DIA Experiments • Nano Pump: EASY-n. LC 1000 • Column: EASY-Spray Column (Pepmap C 18, 2 μm, 75 μm x 50 cm) • Flow rate: 300 n. L/min • Buffer A: 0. 1% FA/H 2 O; Buffer B: 0. 1% FA/CAN • Gradient: • Sample loading: Directly loaded on column 14

Large Scale Quantitative Results on the Q Exactive Plus Instrument using DIA (SWATH Type) Workflow 15
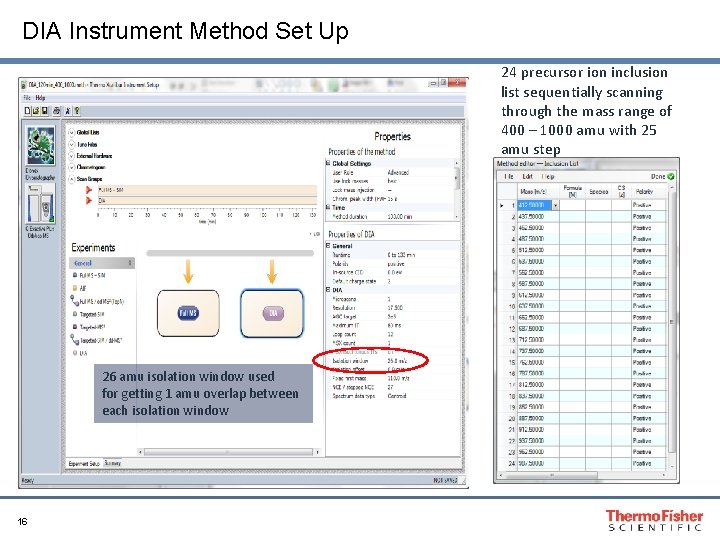
DIA Instrument Method Set Up 24 precursor ion inclusion list sequentially scanning through the mass range of 400 – 1000 amu with 25 amu step 26 amu isolation window used for getting 1 amu overlap between each isolation window 16

DIA Data Processing • Pinpoint 1. 3 • Spectral library established using DDA discovery data • Simultaneous peptide sequence confirmation and quantification + Three most intense fragment ions (b and y types) for quantification + Eight to ten most intense fragment ions (b and y types) for confirmation • Automatic quality control of transitions to eliminate transitions with significant interferences • ± 5 ppm XIC window 17
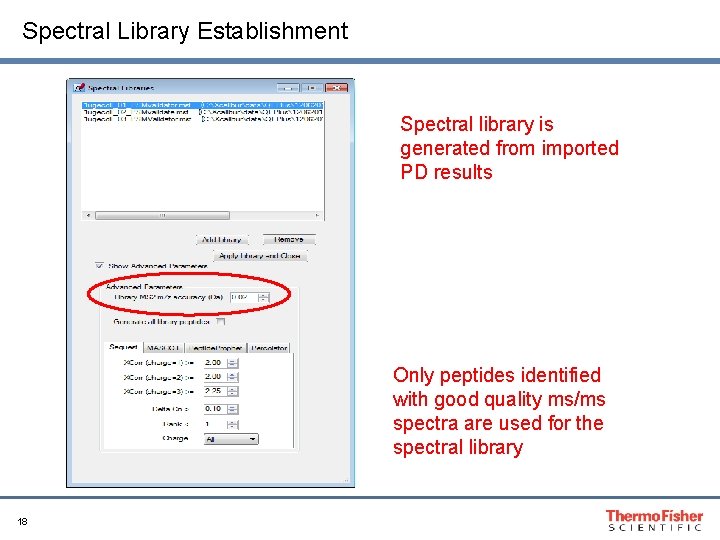
Spectral Library Establishment Spectral library is generated from imported PD results Only peptides identified with good quality ms/ms spectra are used for the spectral library 18
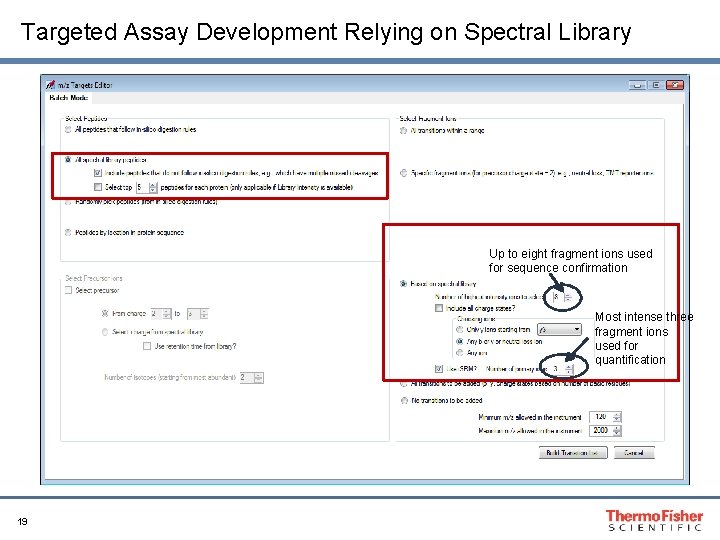
Targeted Assay Development Relying on Spectral Library Up to eight fragment ions used for sequence confirmation Most intense three fragment ions used for quantification 19
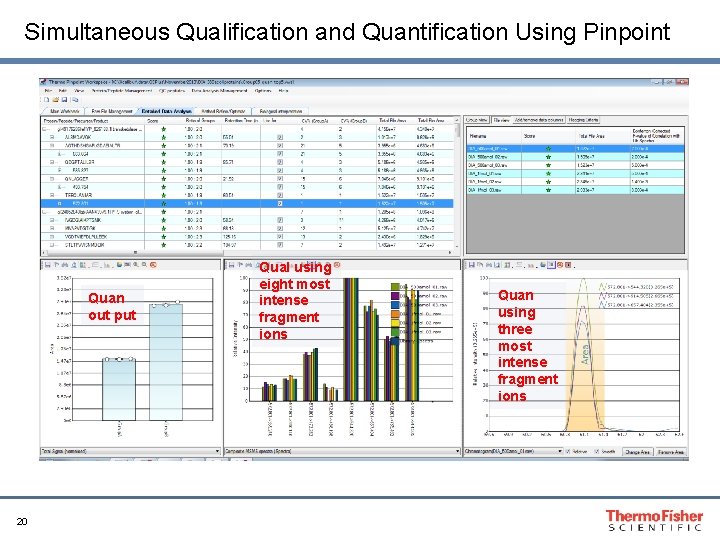
Simultaneous Qualification and Quantification Using Pinpoint Quan out put 20 Qual using eight most intense fragment ions Quan using three most intense fragment ions
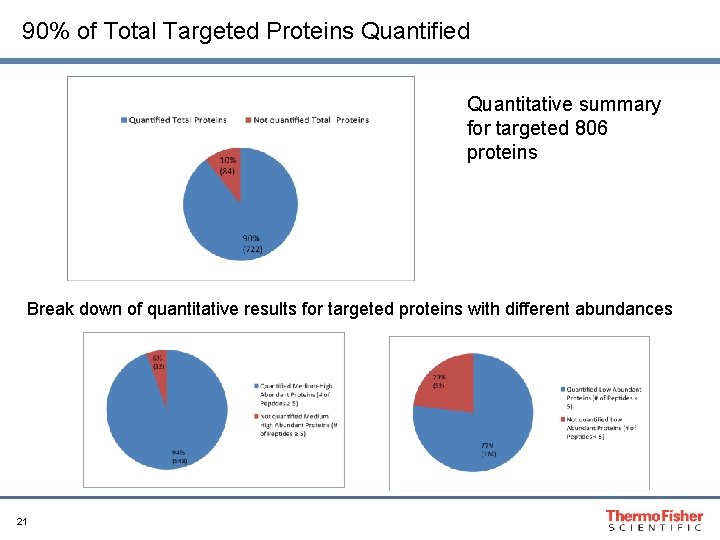
90% of Total Targeted Proteins Quantified Quantitative summary for targeted 806 proteins Break down of quantitative results for targeted proteins with different abundances 21
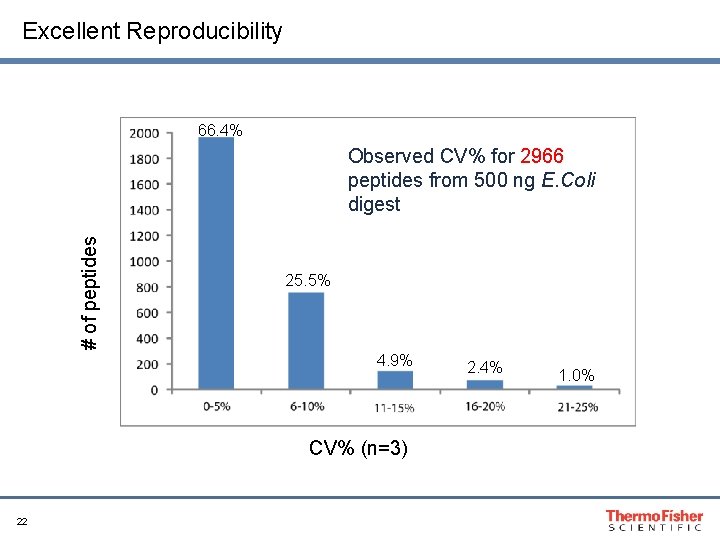
Excellent Reproducibility 66. 4% # of peptides Observed CV% for 2966 peptides from 500 ng E. Coli digest 25. 5% 4. 9% CV% (n=3) 22 2. 4% 1. 0%
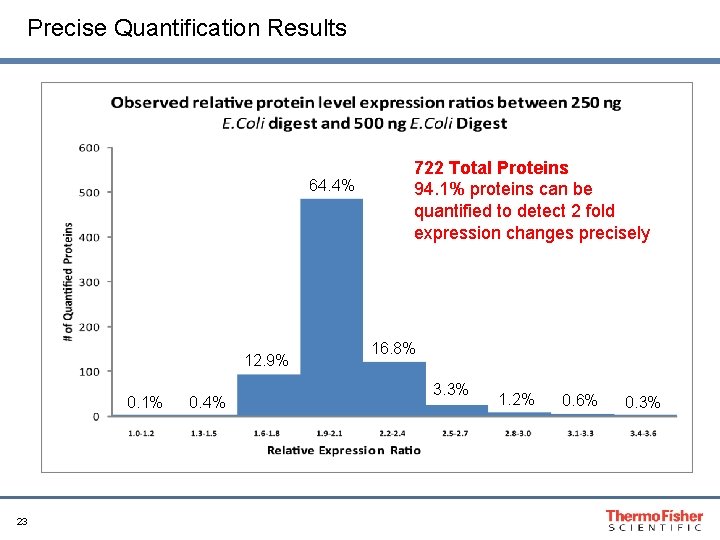
Precise Quantification Results 64. 4% 12. 9% 0. 1% 23 0. 4% 722 Total Proteins 94. 1% proteins can be quantified to detect 2 fold expression changes precisely 16. 8% 3. 3% 1. 2% 0. 6% 0. 3%

Large Scale Quantitative Results on the OT Fusion Instrument using Wi. SIM-DIA Workflow 24
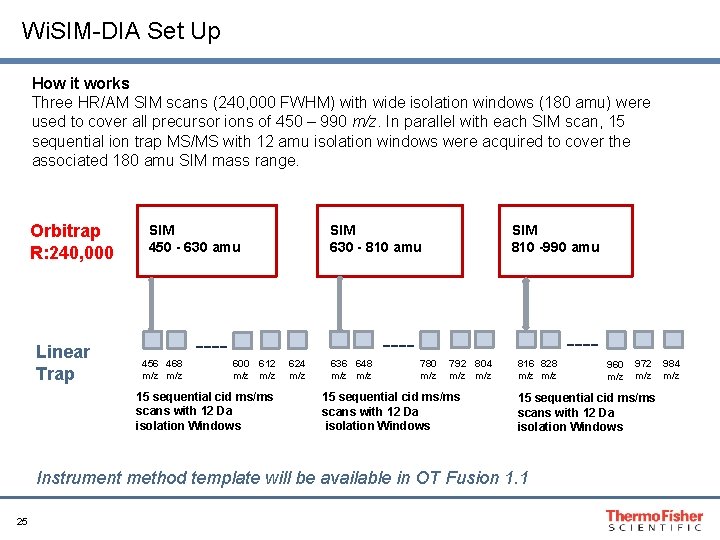
Wi. SIM-DIA Set Up How it works Three HR/AM SIM scans (240, 000 FWHM) with wide isolation windows (180 amu) were used to cover all precursor ions of 450 – 990 m/z. In parallel with each SIM scan, 15 sequential ion trap MS/MS with 12 amu isolation windows were acquired to cover the associated 180 amu SIM mass range. Orbitrap R: 240, 000 Linear Trap SIM 630 - 810 amu SIM 450 - 630 amu 456 468 m/z ---- SIM 810 -990 amu ---- ---600 612 m/z 15 sequential cid ms/ms scans with 12 Da isolation Windows 624 m/z 636 648 m/z 780 m/z 792 804 m/z 15 sequential cid ms/ms scans with 12 Da isolation Windows 816 828 m/z 972 m/z 15 sequential cid ms/ms scans with 12 Da isolation Windows Instrument method template will be available in OT Fusion 1. 1 25 960 m/z 984 m/z

Wi. SIM-DIA Data Processing • Pinpoint 1. 3 software • A spectral library is established using PD search results from the short gun data dependent discovery experiments. • For quantification, the XICs of isotope C 12 and C 13 precursor ions per targeted peptide are extracted from the HR/AM SIM data with ± a 5 ppm window. • For peptide sequence confirmation, eight most intense fragment ions (b and y types) detected from discovery data are extracted from cid ms/ms with ± 600 ppm window and used to match the spectral library. • A peptide with a P-value of correlation with library spectra that is less than 0. 1 is considered to be confirmed with high confidence by the spectral library match. 26
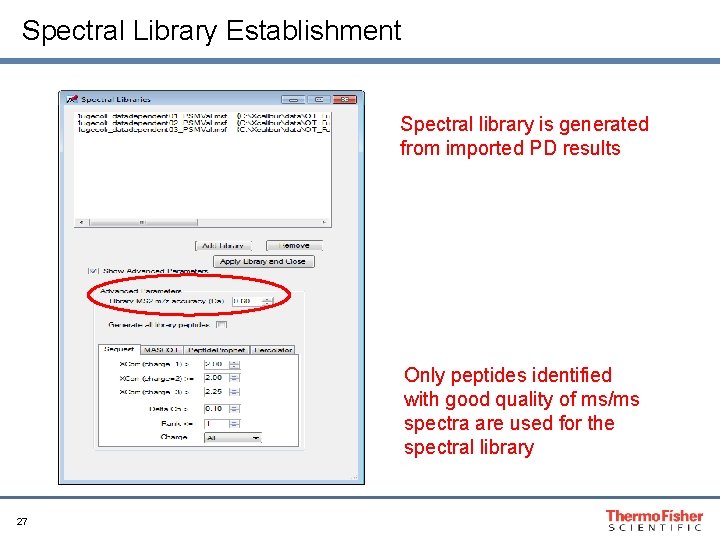
Spectral Library Establishment Spectral library is generated from imported PD results Only peptides identified with good quality of ms/ms spectra are used for the spectral library 27
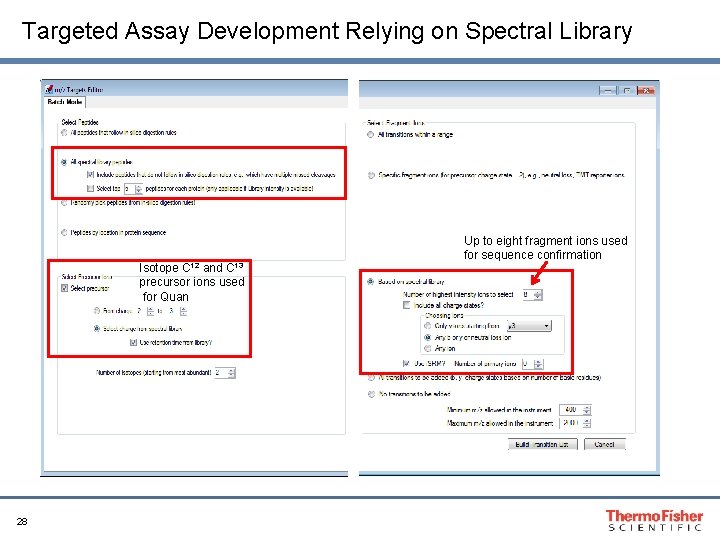
Targeted Assay Development Relying on Spectral Library Isotope C 12 and C 13 precursor ions used for Quan 28 Up to eight fragment ions used for sequence confirmation
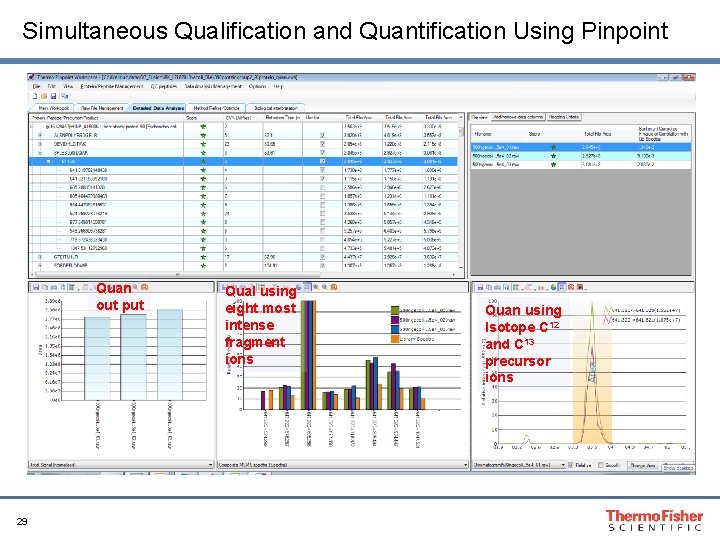
Simultaneous Qualification and Quantification Using Pinpoint Quan out put 29 Qual using eight most intense fragment ions Quan using Isotope C 12 and C 13 precursor ions
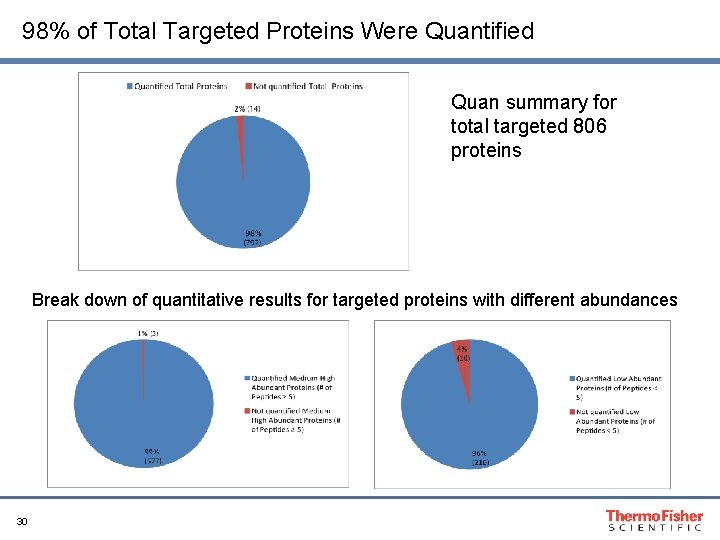
98% of Total Targeted Proteins Were Quantified Quan summary for total targeted 806 proteins Break down of quantitative results for targeted proteins with different abundances 30
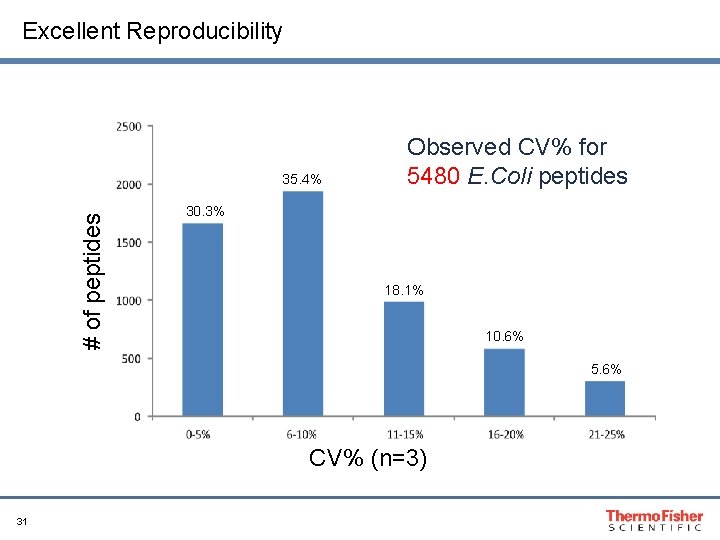
Excellent Reproducibility # of peptides 35. 4% Observed CV% for 5480 E. Coli peptides 30. 3% 18. 1% 10. 6% 5. 6% CV% (n=3) 31
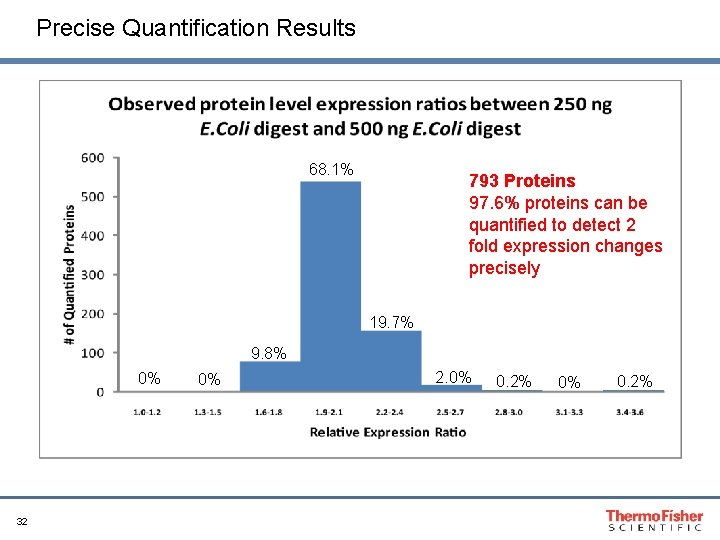
Precise Quantification Results 68. 1% 793 Proteins 97. 6% proteins can be quantified to detect 2 fold expression changes precisely 19. 7% 9. 8% 0% 32 0% 2. 0% 0. 2%

Summary 33
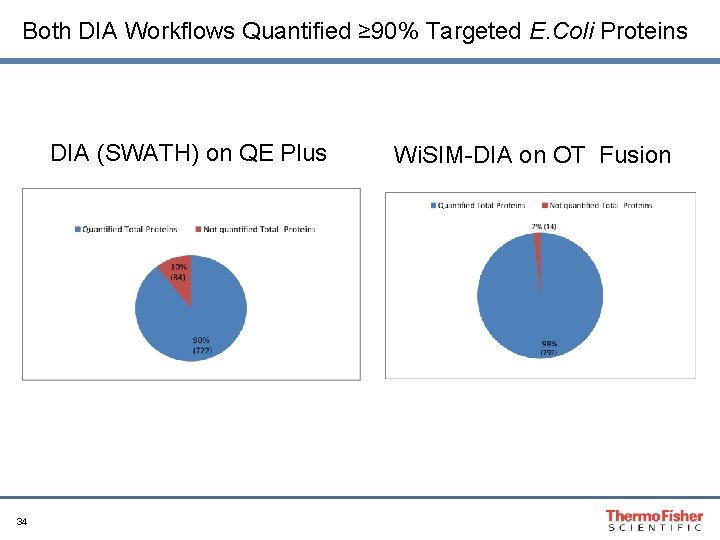
Both DIA Workflows Quantified ≥ 90% Targeted E. Coli Proteins DIA (SWATH) on QE Plus 34 Wi. SIM-DIA on OT Fusion
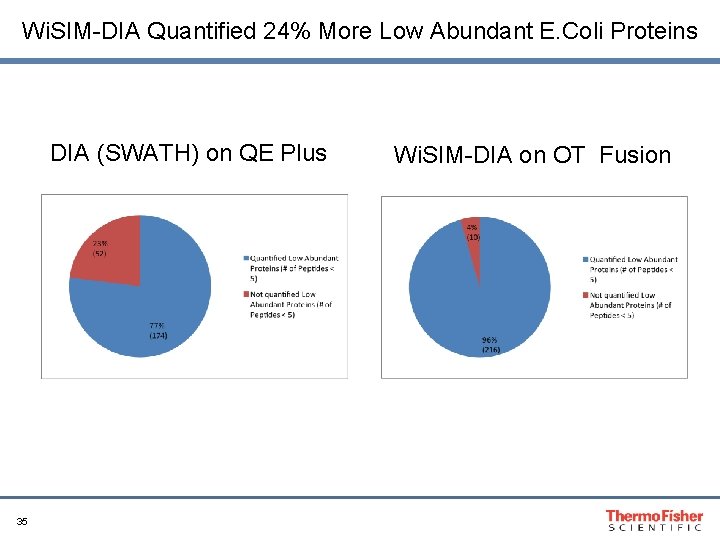
Wi. SIM-DIA Quantified 24% More Low Abundant E. Coli Proteins DIA (SWATH) on QE Plus 35 Wi. SIM-DIA on OT Fusion

Comparison of Available DIA Workflows on Orbitrap Based Instruments Wi. SIM-DIA (SWATH Type) MSX DIA* LOD 10 attomole 50 attomole 100 attomole LOQ (at 15% RSD) 100 attomole 500 attomole 1 fmol Selectivity Very High Medium High *Jarrett D Egertson, Andreas Kuehn, Gennifer E Merrihew, Nicholas W Bateman, Brendan X Mac. Lean, Ying S Ting, Jesse D Canterbury, Donald M Marsh, Markus Kellmann, Vlad Zabrouskov, Christine C Wu & Michael J Mac. Coss Nature Methods Volume: 10, Pages: 744– 746 (2013) 36

Comparison of DIA Processing Software 37
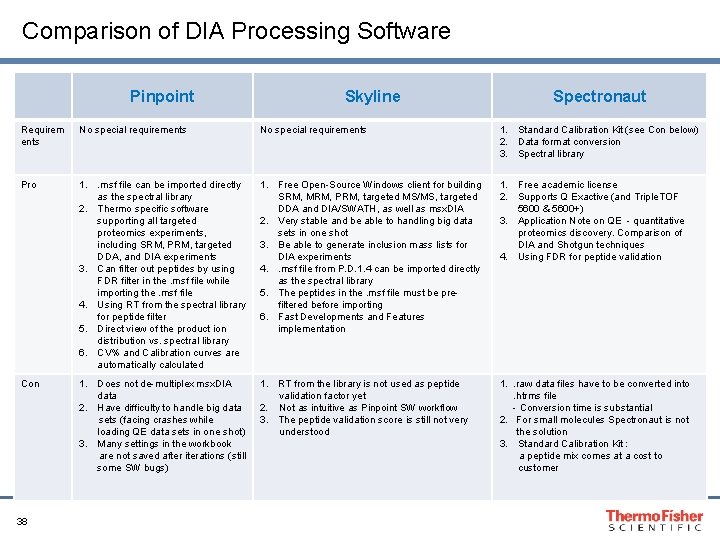
Comparison of DIA Processing Software Pinpoint Skyline Spectronaut Requirem ents No special requirements 1. Standard Calibration Kit (see Con below) 2. Data format conversion 3. Spectral library Pro 1. . msf file can be imported directly as the spectral library 2. Thermo specific software supporting all targeted proteomics experiments, including SRM, PRM, targeted DDA, and DIA experiments 3. Can filter out peptides by using FDR filter in the. msf file while importing the. msf file 4. Using RT from the spectral library for peptide filter 5. Direct view of the product ion distribution vs. spectral library 6. CV% and Calibration curves are automatically calculated 1. Free Open-Source Windows client for building SRM, MRM, PRM, targeted MS/MS, targeted DDA and DIA/SWATH, as well as msx. DIA 2. Very stable and be able to handling big data sets in one shot 3. Be able to generate inclusion mass lists for DIA experiments 4. . msf file from P. D. 1. 4 can be imported directly as the spectral library 5. The peptides in the. msf file must be prefiltered before importing 6. Fast Developments and Features implementation 1. Free academic license 2. Supports Q Exactive (and Triple. TOF 5600 & 5600+) 3. Application Note on QE - quantitative proteomics discovery. Comparison of DIA and Shotgun techniques 4. Using FDR for peptide validation Con 1. Does not de-multiplex msx. DIA data 2. Have difficulty to handle big data sets (facing crashes while loading QE data sets in one shot) 3. Many settings in the workbook are not saved after iterations (still some SW bugs) 1. RT from the library is not used as peptide validation factor yet 2. Not as intuitive as Pinpoint SW workflow 3. The peptide validation score is still not very understood 1. . raw data files have to be converted into. htrms file - Conversion time is substantial 2. For small molecules Spectronaut is not the solution 3. Standard Calibration Kit : a peptide mix comes at a cost to customer 38

Conclusion • The Wi. SIM-DIA is unique on the OT Fusion Instrument. • The Wi. SIM-DIA provides lower detection limits and higher selectivity by using precursor ions collected on SIM with extremely high resolving power of 240, 000 for quantification, which enables separation of most background interferences from analyte signal at MS level. • The MSX-DIA is unique on Orbitrap based instruments. • The MSX-DIA provides higher selectivity than DIA and could provide better quantitative results than DIA for complex samples. • The high resolving power of Orbitrap based instruments enable precise quantification by using XICs with ± 5 ppm windows. • Innovative Thermo Scientific™ EASY-Spray™ source and EASY-Spray columns enable stable peptide signal intensities and retention times across multiple sample runs which assure reproducible quantitative results. 39

Next Steps • Publish the DIA and Wi. SIM-DIA data discussed today • Multiple application notes including data processing using Pinpoint • Additional data with collaborators 40
- Slides: 40