Integrating Free Surfer and FSL FEAT 1 Outline

















- Slides: 36

Integrating Free. Surfer and FSL FEAT 1

Outline • Registering FEAT Free. Surfer Anatomical • Automatic (FLIRT) • Manual (tkregister 2) • Surface-based Group f. MRI Analysis • Individual Analysis • Viewing FEAT output on Anatomical • Sampling FEAT output on the surface • Mapping Free. Surfer Segmentations to FEAT 2

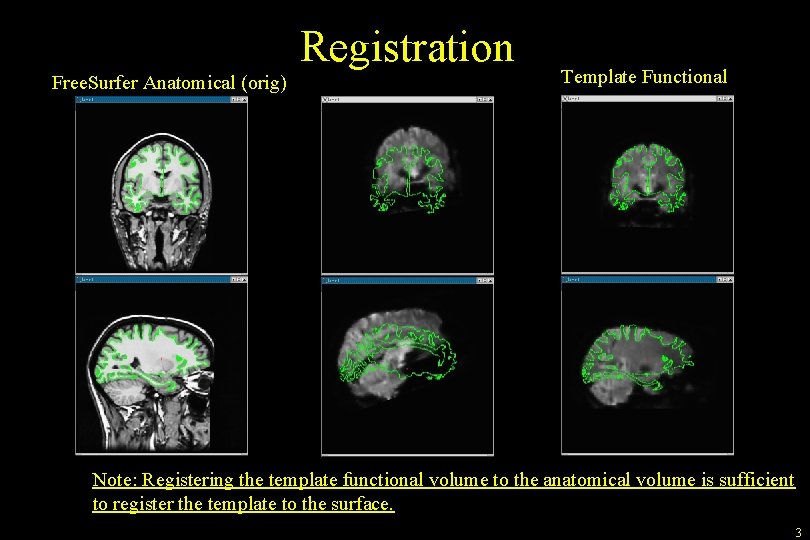
Registration Free. Surfer Anatomical (orig) Template Functional Note: Registering the template functional volume to the anatomical volume is sufficient to register the template to the surface. 3

Free. Surfer Registration Free. Surfer Subject-Specific • Volumes • Surfaces • Thickness • ROIs Registration Your Data/Software • f. MRI (FSL, etc) • EEG/MEG • DTI • … Registration Matrix • Affine 4 x 4 • As many as 12 DOF (usually 6) • Text file 4

Automatic Registration First: analyze your data with FEAT reg-feat 2 anat –featdir fbert 1. feat –subject bert Uses BBR to perform 6 DOF registration: fbert 1. feat/example_func. nii. gz $SUBJECTS_DIR/bert/mri/brainmask. mgz Creates Free. Surfer registration matrix: fbert 1. feat/reg/freesurfer/anat 2 exf. register. dat Also creates: fbert 1. feat/reg/freesurfer/anat 2 std. register. dat This matrix maps standard space to anatomical space and can be used when combining data within subject across data sets reg-feat 2 anat --help 5

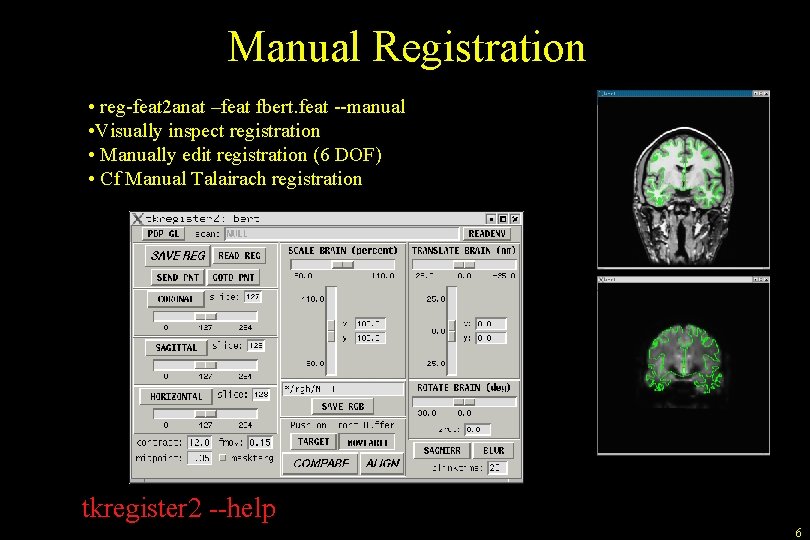
Manual Registration • reg-feat 2 anat –feat fbert. feat --manual • Visually inspect registration • Manually edit registration (6 DOF) • Cf Manual Talairach registration tkregister 2 --help 6

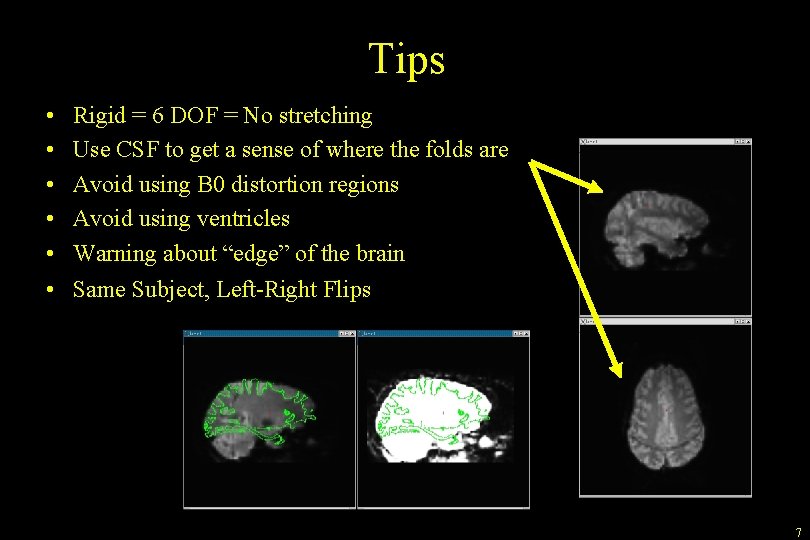
Tips • • • Rigid = 6 DOF = No stretching Use CSF to get a sense of where the folds are Avoid using B 0 distortion regions Avoid using ventricles Warning about “edge” of the brain Same Subject, Left-Right Flips 7

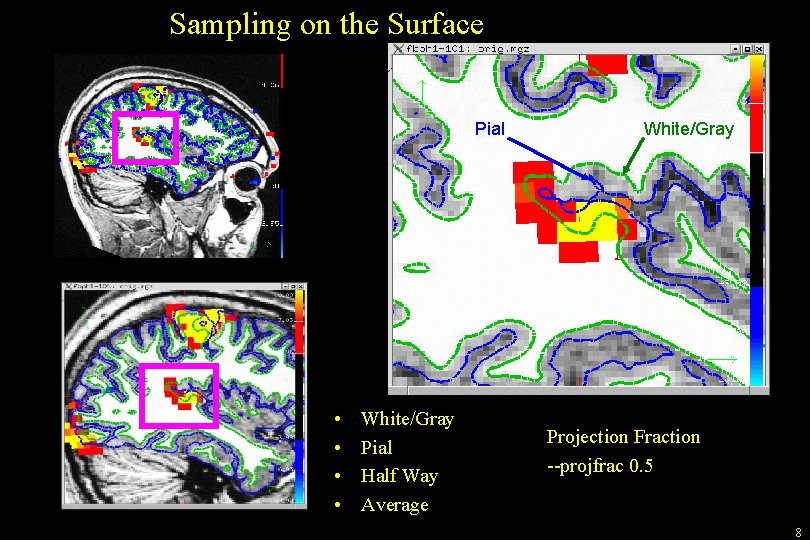
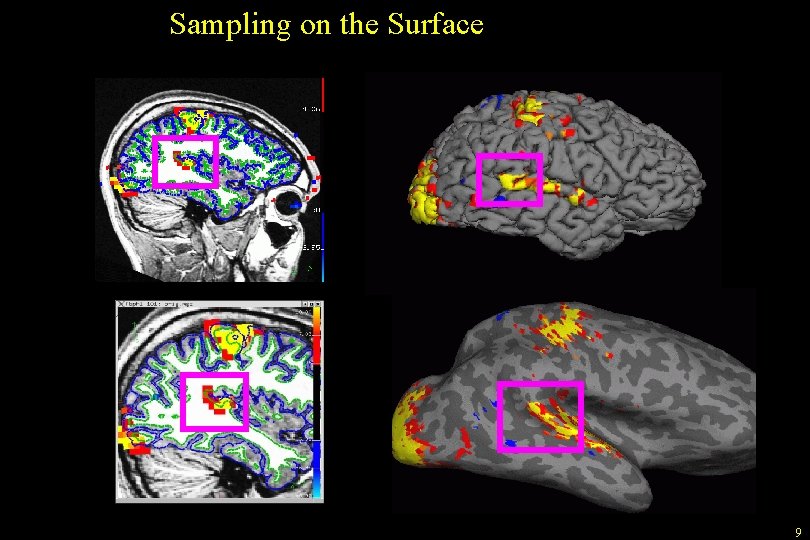
Sampling on the Surface Pial • • White/Gray Pial Half Way Average White/Gray Projection Fraction --projfrac 0. 5 8

Sampling on the Surface 9

Surface-based f. MRI Group Analysis 10

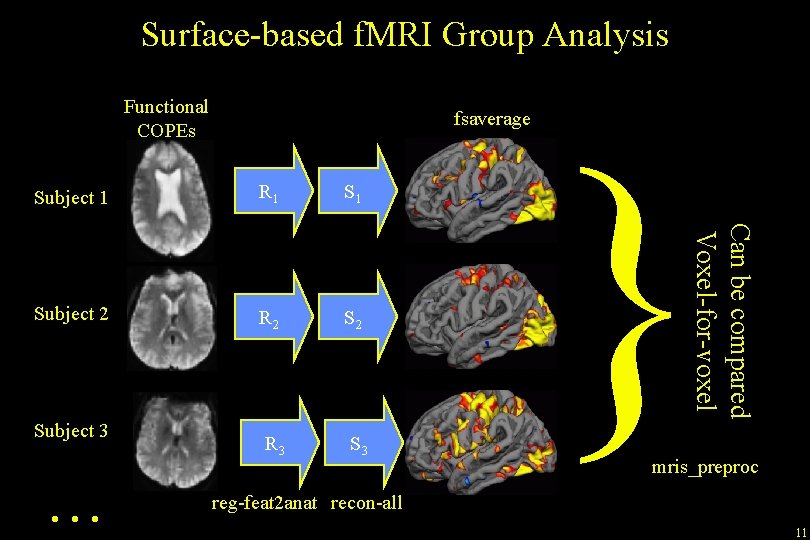
Surface-based f. MRI Group Analysis Functional COPEs fsaverage R 1 Subject 2 R 2 S 2 R 3 Subject 3 … } Can be compared Voxel-for-voxel Subject 1 mris_preproc reg-feat 2 anat recon-all 11

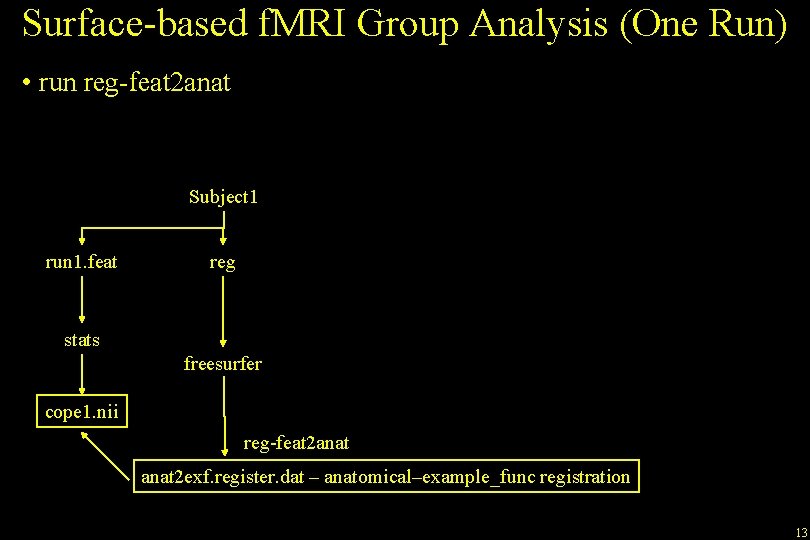
Surface-based f. MRI Group Analysis (One Run) • Single Run • Analyze with FEAT, No smoothing • COPEs are in native functional space Subject 1 run 1. feat stats cope 1. nii 12

Surface-based f. MRI Group Analysis (One Run) • run reg-feat 2 anat Subject 1 run 1. feat reg stats freesurfer cope 1. nii reg-feat 2 anat 2 exf. register. dat – anatomical–example_func registration 13

Surface-based Group f. MRI Analysis (One Run) mris_preproc --out lh. cope 1. nii. gz --target fsaverage --hemi lh --iv bert. feat/stats/cope 1. nii. gz bert. feat/reg/freesurfer/anat 2 exf. register. dat --iv greg. feat/stats/cope 1. nii. gz greg. feat/reg/freesurfer/anat 2 exf. register. dat --iv sally. feat/stats/cope 1. nii. gz sally. feat/reg/freesurfer/anat 2 exf. register. dat --iv pat. feat/stats/cope 1. nii. gz pat. feat/reg/freesurfer/anat 2 exf. register. dat Volumes Registrations • lh. cope 1. mgh – stack of subjects • Can map any functional data, eg, zstat, fzstat, cope, etc • fsaverage – defines common space (spherical surface) • Group analysis same as with a thickness study: • surface smoothing, mri_glmfit, mri_glmfi-sim 14

Surface-based f. MRI Group Analysis (>1 Run) • Multiple Runs • Analyze each run with FEAT, No smoothing • COPEs are in native functional space Subject 1 run 1. feat run 2. feat stats cope 1. nii 15

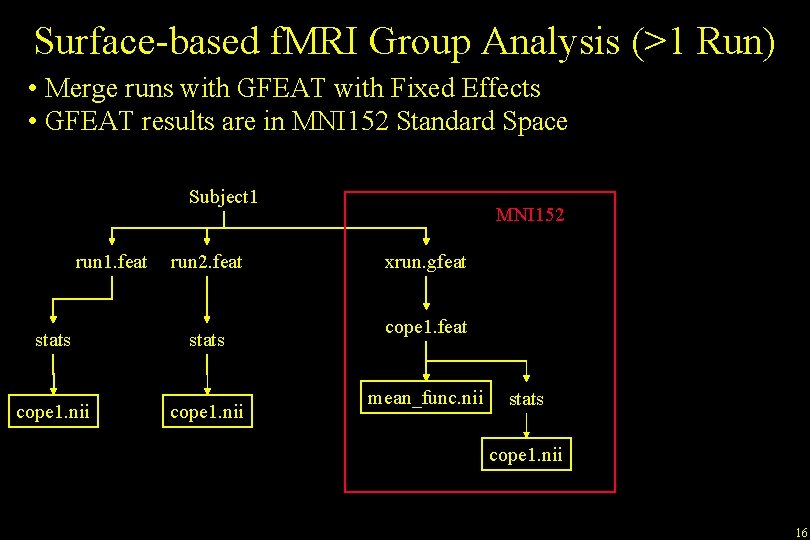
Surface-based f. MRI Group Analysis (>1 Run) • Merge runs with GFEAT with Fixed Effects • GFEAT results are in MNI 152 Standard Space Subject 1 run 1. feat stats cope 1. nii run 2. feat stats cope 1. nii MNI 152 xrun. gfeat cope 1. feat mean_func. nii stats cope 1. nii 16

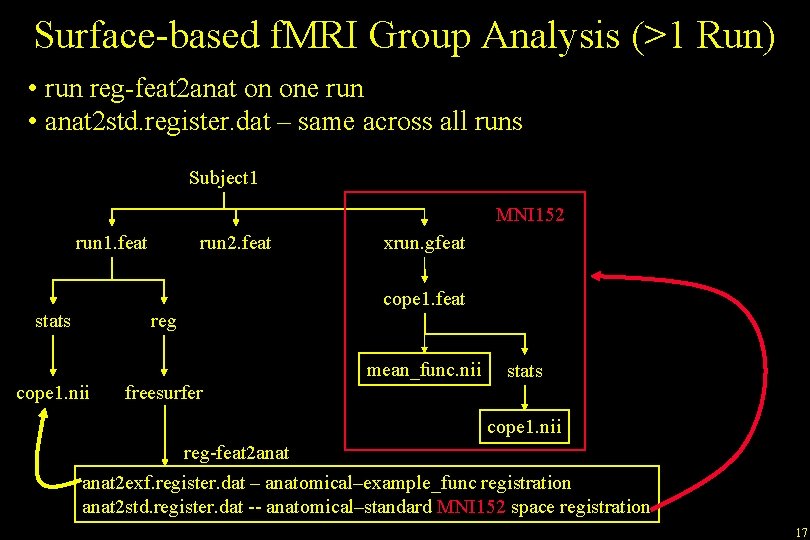
Surface-based f. MRI Group Analysis (>1 Run) • run reg-feat 2 anat on one run • anat 2 std. register. dat – same across all runs Subject 1 MNI 152 run 1. feat stats run 2. feat reg xrun. gfeat cope 1. feat mean_func. nii cope 1. nii freesurfer stats cope 1. nii reg-feat 2 anat 2 exf. register. dat – anatomical–example_func registration anat 2 std. register. dat -- anatomical–standard MNI 152 space registration 17

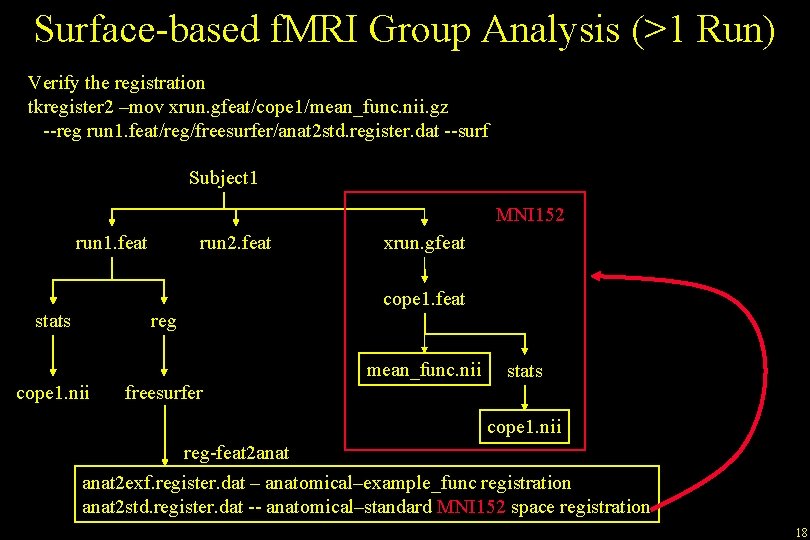
Surface-based f. MRI Group Analysis (>1 Run) Verify the registration tkregister 2 –mov xrun. gfeat/cope 1/mean_func. nii. gz --reg run 1. feat/reg/freesurfer/anat 2 std. register. dat --surf Subject 1 MNI 152 run 1. feat stats run 2. feat reg xrun. gfeat cope 1. feat mean_func. nii cope 1. nii freesurfer stats cope 1. nii reg-feat 2 anat 2 exf. register. dat – anatomical–example_func registration anat 2 std. register. dat -- anatomical–standard MNI 152 space registration 18

Surface-based f. MRI Group Analysis (>1 Run) mris_preproc --out lh. cope 1. mgh --target fsaverage --hemi lh --iv bert. gfeat/cope 1. feat/stats/cope 1. nii. gz bert. feat/reg/freesurfer/anat 2 std. register. dat --iv greg. gfeat/cope 1. feat/stats/cope 1. nii. gz greg. feat/reg/freesurfer/anat 2 std. register. dat --iv sally. feat/cope 1. feat/stats/cope 1. nii. gz sally. feat/reg/freesurfer/anat 2 std. register. dat --iv pat. feat/cope 1. feat/stats/cope 1. nii. gz pat. feat/reg/freesurfer/anat 2 std. register. dat Volumes Registrations • lh. cope 1. mgh – stack of subjects • Can map any functional data, eg, zstat, fzstat, cope, etc • fsaverage – defines common space (spherical surface) • Group analysis same as with a thickness study: • surface smoothing, mri_glmfit, mri_glmfi-sim 19

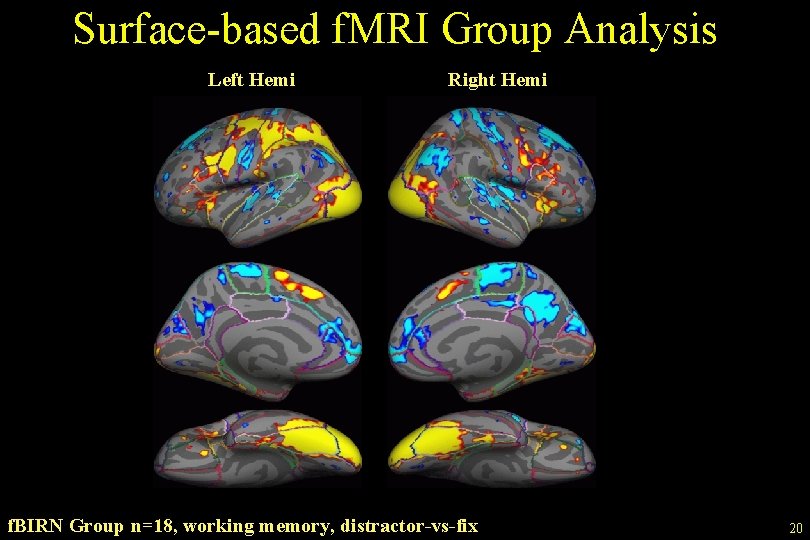
Surface-based f. MRI Group Analysis Left Hemi Right Hemi f. BIRN Group n=18, working memory, distractor-vs-fix 20

Individual Subject Integration Applying Free. Surfer Tools to FSL f. MRI Analysis 21

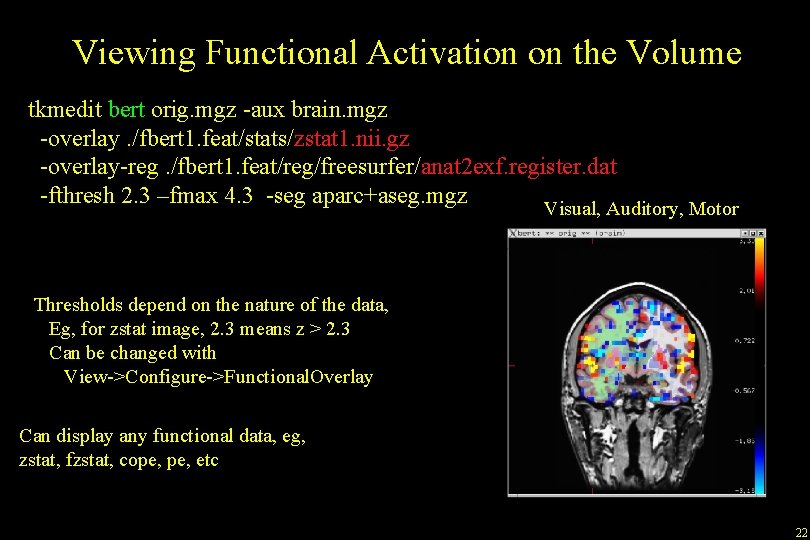
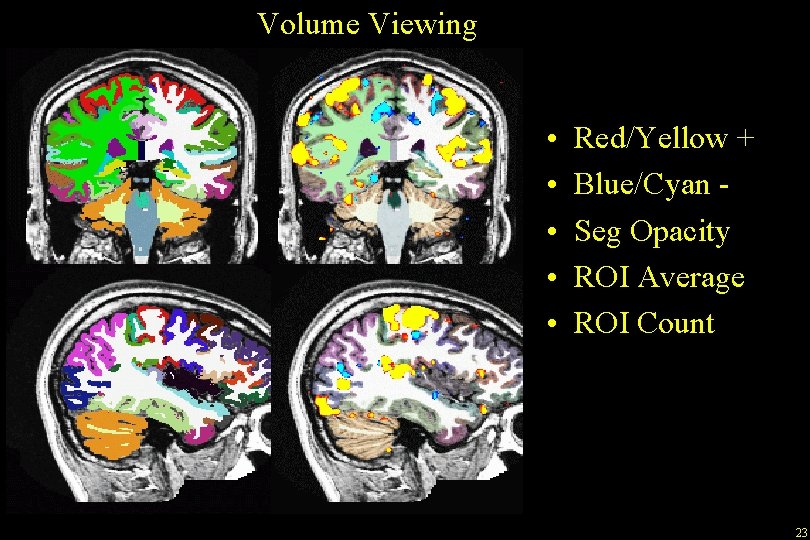
Viewing Functional Activation on the Volume tkmedit bert orig. mgz -aux brain. mgz -overlay. /fbert 1. feat/stats/zstat 1. nii. gz -overlay-reg. /fbert 1. feat/reg/freesurfer/anat 2 exf. register. dat -fthresh 2. 3 –fmax 4. 3 -seg aparc+aseg. mgz Visual, Auditory, Motor Thresholds depend on the nature of the data, Eg, for zstat image, 2. 3 means z > 2. 3 Can be changed with View->Configure->Functional. Overlay Can display any functional data, eg, zstat, fzstat, cope, etc 22

Volume Viewing • • • Red/Yellow + Blue/Cyan Seg Opacity ROI Average ROI Count 23

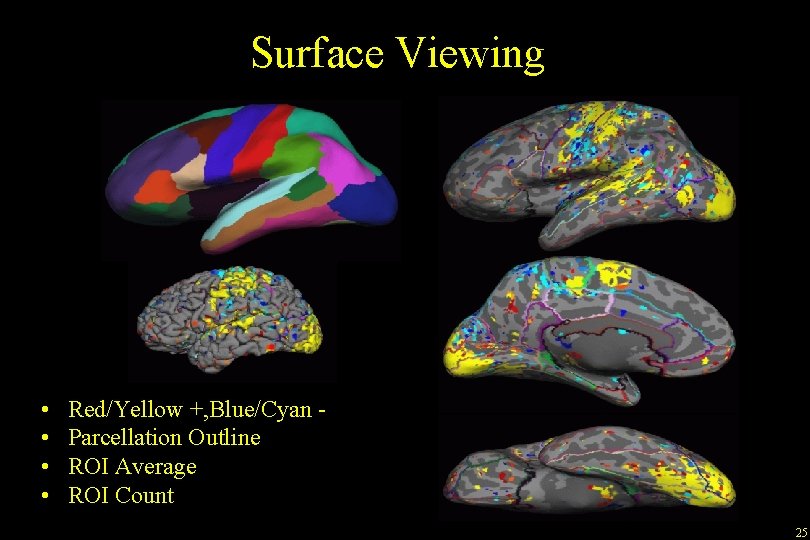
Viewing FEAT Stats on the Surface tksurfer bert rh inflated -overlay. /fbert 1. feat/stats/zstat 1. nii. gz -overlay-reg. /fbert 1. feat/reg/freesurfer/anat 2 exf. register. dat -fthresh 2. 3 -fmid 3. 3 -fslope 1 Visual, Auditory, Motor Can display any functional data, eg, zstat, fzstat, cope, etc 24

Surface Viewing • • Red/Yellow +, Blue/Cyan Parcellation Outline ROI Average ROI Count 25

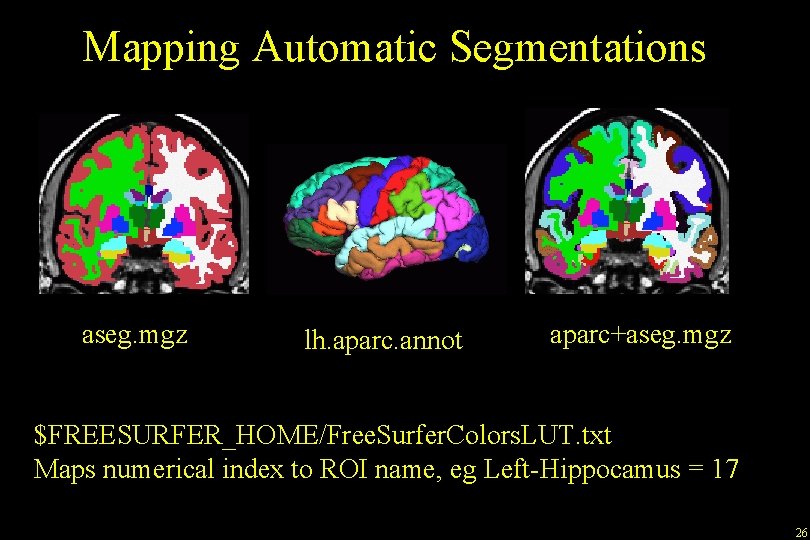
Mapping Automatic Segmentations aseg. mgz lh. aparc. annot aparc+aseg. mgz $FREESURFER_HOME/Free. Surfer. Colors. LUT. txt Maps numerical index to ROI name, eg Left-Hippocamus = 17 26

Mapping Automatic Segmentations aseg 2 feat --feat fbert. feat –aseg aparc+aseg In the functional FOV, creates: fbert 1. feat/reg/freesurfer/aseg+aparc. nii. gz Create Text Summary Table (nvox, mean cope, std cope, etc) mri_segstats --seg fbert 1. feat/reg/freesurfer/aparc+aseg. nii. gz --nonempty --ctab-default --in fbert. feat/stats/cope 1. nii. gz --sum fbert 1. sum. txt Can summarize any functional data, eg, zstat, fzstat, cope, etc 27

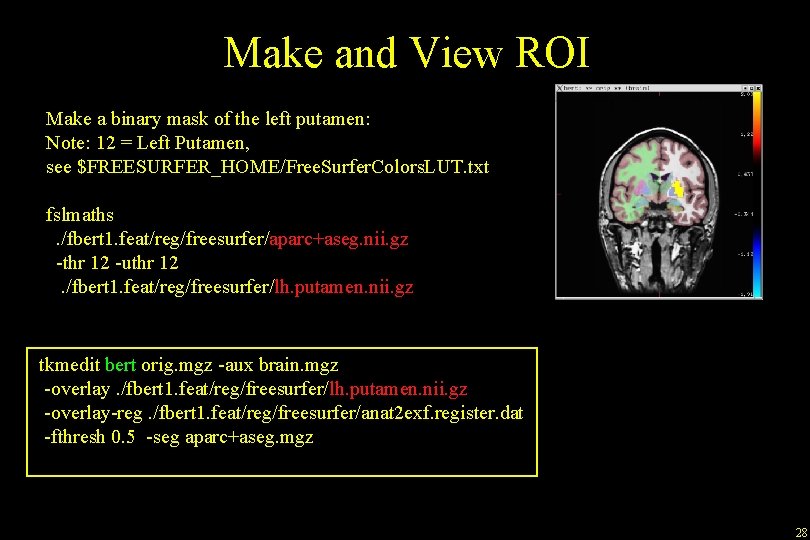
Make and View ROI Make a binary mask of the left putamen: Note: 12 = Left Putamen, see $FREESURFER_HOME/Free. Surfer. Colors. LUT. txt fslmaths. /fbert 1. feat/reg/freesurfer/aparc+aseg. nii. gz -thr 12 -uthr 12. /fbert 1. feat/reg/freesurfer/lh. putamen. nii. gz tkmedit bert orig. mgz -aux brain. mgz -overlay. /fbert 1. feat/reg/freesurfer/lh. putamen. nii. gz -overlay-reg. /fbert 1. feat/reg/freesurfer/anat 2 exf. register. dat -fthresh 0. 5 -seg aparc+aseg. mgz 28

Practical Data • • • Sensory-motor study Blocked Design (15 sec OFF, 15 sec ON) One subject – “bert” Two runs (TR=3, 85 time points) FEAT has been run on both runs (FWHM=5) Combined with GFEAT – FFx – One-Sample Group Mean (OSGM) • Actually, all analysis steps already performed! 29

Practical • • Automatic registration (<5 min) Manual registration View FEAT results on subject’s anatomy (aparc+aseg) View FEAT results with tksurfer Map aparc+aseg to Functional Space Verify GFEAT registration View GFEAT results in volume and on surface “Higher-Level” analysis with mri_glmfit – Cross-Run – Fixed-Effects (FFX), One-Sample Group Mean (osgm) 30

End of Presentation 31

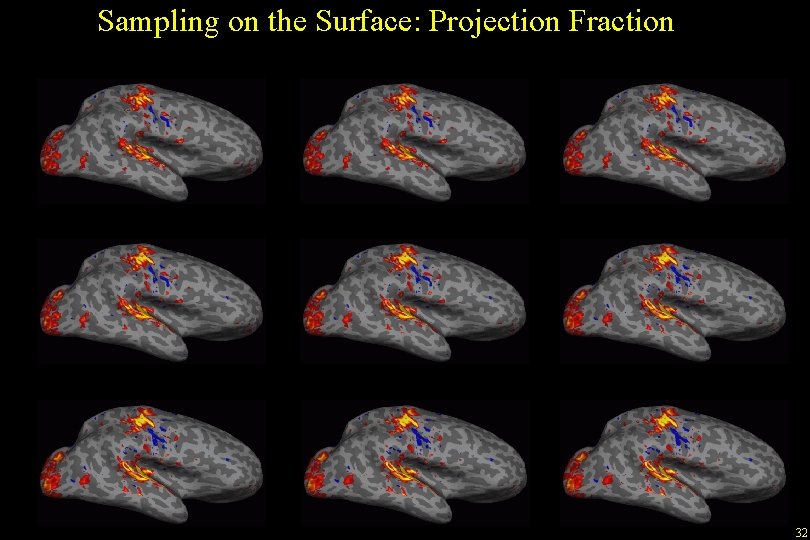
Sampling on the Surface: Projection Fraction -0. 1 0. 0 (white) +0. 1 +0. 3 +0. 5 +0. 7 +0. 9 +1. 0 (pial) +1. 1 32

Step 1: Register Anatomical with Surface Atlas (fsaverage) Native Anatomical Surface Space Surface-based Registration fsaverage Space S recon-all Anatomical Surface in fsaverage Space 33

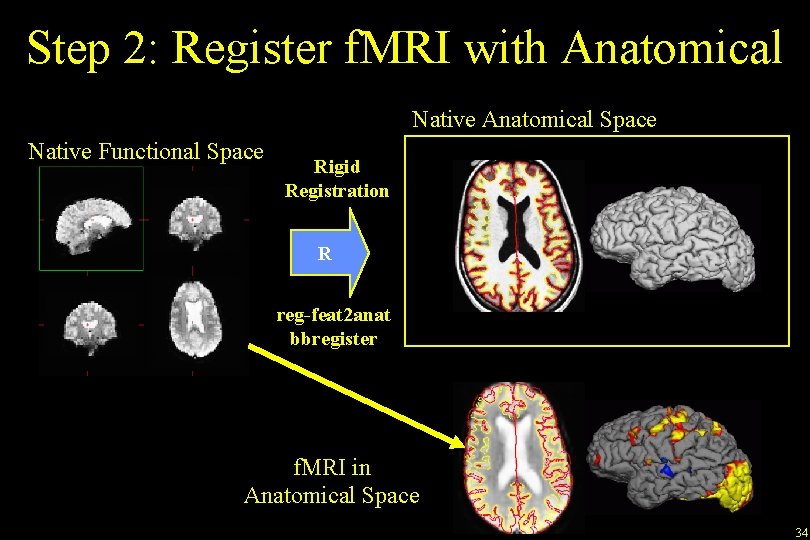
Step 2: Register f. MRI with Anatomical Native Anatomical Space Native Functional Space Rigid Registration R reg-feat 2 anat bbregister f. MRI in Anatomical Space 34

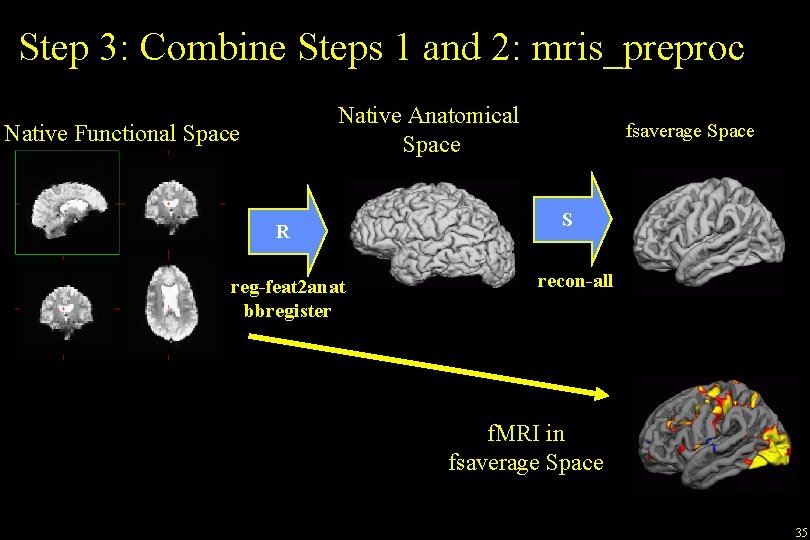
Step 3: Combine Steps 1 and 2: mris_preproc Native Anatomical Space Native Functional Space R reg-feat 2 anat bbregister fsaverage Space S recon-all f. MRI in fsaverage Space 35

Within-Subject, Cross-Run Analysis • Analyze each run with FEAT (data. X. feat) • Combine runs with GFEAT (standard space) • mean_func. nii. gz – avg of example_func in std space • Register each run (not. gfeat) with reg-feat 2 anat. • data. X. feat/reg/freesurfer/anat 2 std. register. dat • All runs (X) should be very close Verify the registration tkregister 2 –mov data. gfeat/mean_func. nii. gz --reg anat 2 std. register. dat --surf Use anat 2 std. register. dat the way you would anat 2 exf. register. dat 36