Insertional mutagenesis in zebrafish rapidly identifies genes essential

Insertional mutagenesis in zebrafish rapidly identifies genes essential for early vertebrate development Arnaldur Hall Freyr Ævarsson Þorkell Guðjónsson Gregory Colling, Adam Amsterdam, Zhaoxia Sun, Marcelo Antonelli, Ernesto Maldonado, Wenbiao Chen, Shawn Burgess, Maryann Haldi, Karen Artzt, Sarah Farrington, Shuh-Yow Lin, Robert M. Nissen & Nancy Hopkins

Why are zebrafish (Danio rerio) ideal models for development and disease research? • Zebrafish are vertebrates. Like humans, they have a backbone. • Zebrafish have features that make them easy to maintain, manipulate, and observe in the lab. • The embryos develop outside the mother's body, so you can have easy access to them. • Zebrafish embryos are transparent. This means you can watch development as it happens in living embryos. • The embryos develop quickly. • You can physically manipulate the embryos.

Introduction • It has been estimated that roughly 800 genes can be mutaded to yield specific or localized defects during development in zebrafish • Approximately 1600 additional genes can be mutaded to yield less specific phenotypes or recurring syndromes • Identification of these 2, 400 genes would contribute significantly to understanding vertebrate development.

Indroduction • More than 500 insertional mutants have been isolated. • First 75 insertional mutants, for which the disrupted genes have been identified are described in this article. • The genes underlying about 50 mutants had been reported when this article was published (june 2002)

Experimental Questions • How many genes are essential for the development of a vertebrate? • What is the role of each gene? • How serious are the defects caused by mutated genes?

Identification of retrovirusinduced mutations Methods • Mutagenesis: Method of insertional mutagenesis for zebrafish was designed, using a Moloney murine leukemia−based retroviral vector as a mutagen. Plus + Minus Integrates into many different sites in mammalian and avian chromosomes Less efficient than chemicals Integrates without rearrangement of its own sequences or significant alterations to host DNA - Seldom, if ever, integrates entirely randomly

Identification of retrovirus-induced mutations • DNA flanking the insert cloned by inverse PCR • If candidate gene was not found small chromosomal walk was used • RT-PCR and RACE was then used to obtain the rest of the c. DNA • To confirm that the correct junction fragment (and gene) have been cloned, linkage analysis was carried out • Primers were designed to amplify different-sized products from chromosomes with or without the putative mutagenic insert in a PCRbased assay

Inverse PCR • Inverse PCR is used to amplify and clone unknown DNA that flanks one end of a known DNA sequence and for which no primers are available. • The technique involves digestion by a restriction enzyme of a preparation of DNA containing the known sequence and its flanking region. • The individual restriction fragments are converted into circles by intramolecular ligation, and the circularized DNA is then used as a template in PCR. • The unknown sequence is amplified by two primers that bind specifically to the known sequence and point in opposite direction.

Identification of retrovirus-induced mutations Methods Genotyping embryos: • embryos were sorted from heterozygous parents into phenotypically wildtype and mutant groups • mutant embryos are homozygous with respect to the mutagenic insertion • almost all of the mutants are recessive-lethals • 24 embryos of each group were genotyped by PCR • Pair of genomic primers flanking the responsible mutagenic viral insertion and a viral-specific primer were used in a single reaction • A viral insertion leads to amplification between the viralspecific primer and one of the genomic primers

Identification of retrovirus-induced mutations • If no recombinants are seen, the insert should lie no further than 3 c. M (2 Mb) from the mutation responsible for the mutant phenotype. • The screen is not perfect • It is possible that a proviral insert could be linked to a mutation, but not be its cause.

Identification of retrovirus-induced mutations • Further evidence that the correct gene had been identified obteined by use of RT-PCR or in situ hybridization • In this screen, all embryos with mutant phenotypes were kept visible in a dissecting microscope by five days postfertilization

Classification of mutant phenotypes • Specific developmental defects • brain, eyes, jaw, arches or cartilages, midline, ear, fins, liver, gut, kidney, muscle, pigment, body shape, motility or touch insensitivity and motility or altered touch sensitivity in combination with some degree of visible morphological defects • General developmental defects • extensive cell death in the central nervous system (CNS), small head and eyes, embryos that show retardation, mutants that have several defects and mutants which fail to inflate the swim bladder

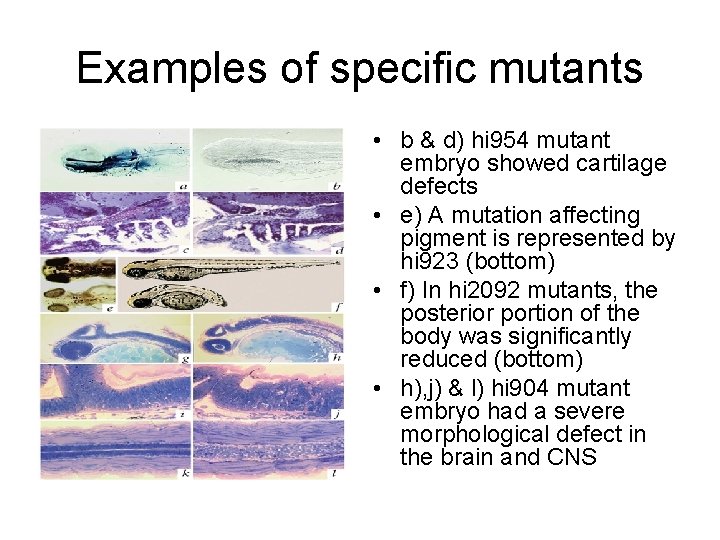
Examples of specific mutants • b & d) hi 954 mutant embryo showed cartilage defects • e) A mutation affecting pigment is represented by hi 923 (bottom) • f) In hi 2092 mutants, the posterior portion of the body was significantly reduced (bottom) • h), j) & l) hi 904 mutant embryo had a severe morphological defect in the brain and CNS
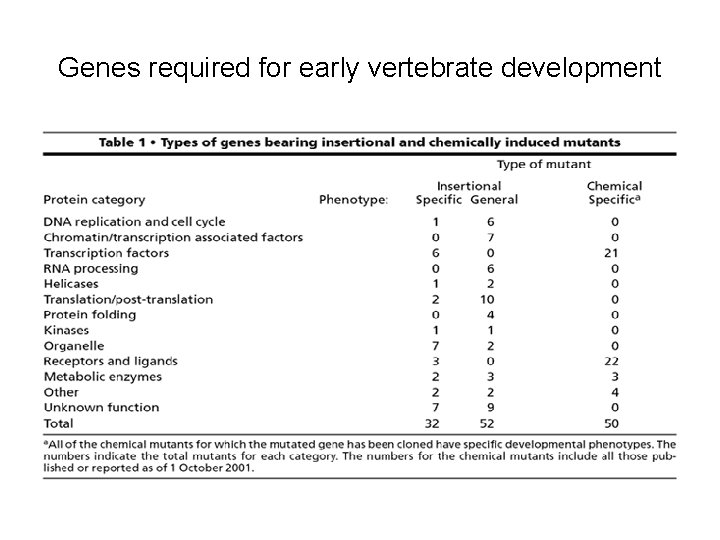
Genes required for early vertebrate development
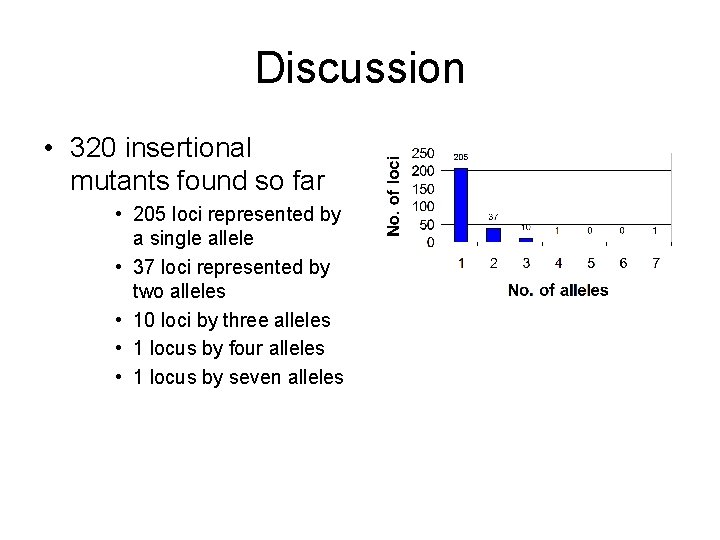
Discussion • 320 insertional mutants found so far • 205 loci represented by a single allele • 37 loci represented by two alleles • 10 loci by three alleles • 1 locus by four alleles • 1 locus by seven alleles

Discussion • The data do not fit a Poisson distribution • The allele distribution in large-scale ENU mutagenesis screens also did not fit a Poisson distribution • => All genes are not mutated with equal frequency with either mutagen • => Retroviruses may be no more biased in the genes they mutate than are chemical mutagens

The end •
- Slides: 18