Informatics tools for nextgeneration sequence analysis Gabor Marth

Informatics tools for next-generation sequence analysis Gabor Marth Boston College Biology Next-Generation Sequencing Mini. Symposium CHOP Philadelphia, PA April 6, 2009

New sequencing technologies…

… offer vast throughput 100 Gb Illumina/Solexa, AB/SOLi. D sequencers (10 -30 Gb in 25 -100 bp reads) bases per machine run 10 Gb 1 Gb Roche/454 pyrosequencer (100 -400 Mb in 200 -450 bp reads) 100 Mb 10 Mb ABI capillary sequencer 1 Mb 10 bp 100 bp read length 1, 000 bp
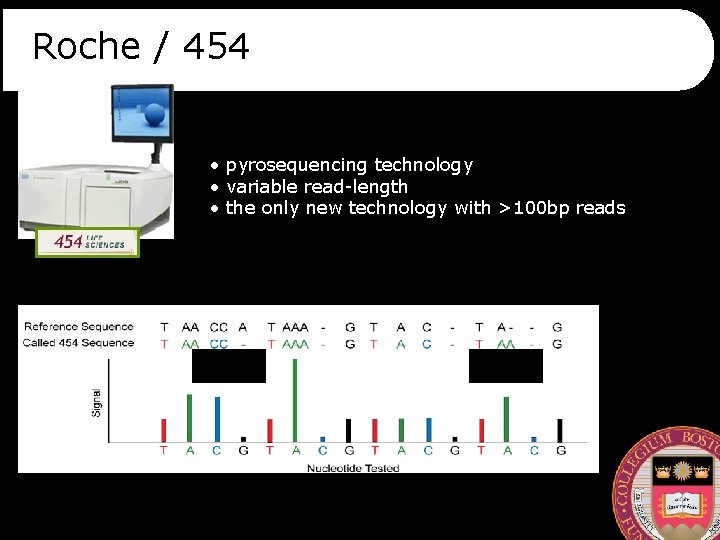
Roche / 454 • pyrosequencing technology • variable read-length • the only new technology with >100 bp reads

Illumina / Solexa • fixed-length short-read sequencer • very high throughput • read properties are very close to traditional capillary sequences
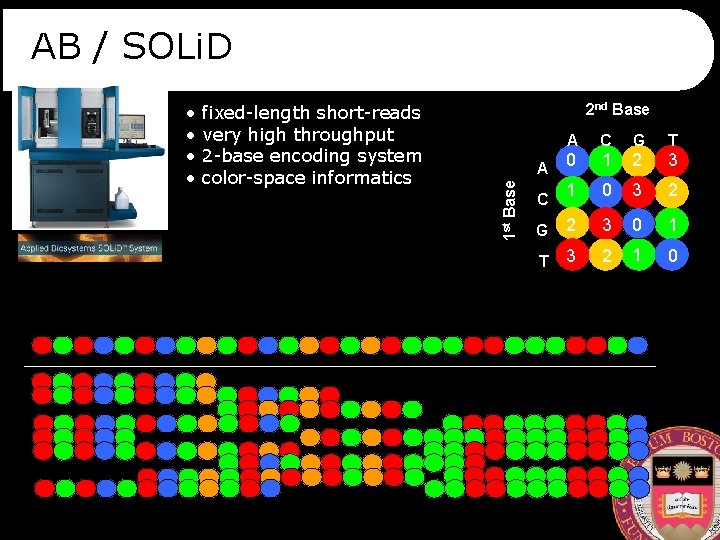
AB / SOLi. D fixed-length short-reads very high throughput 2 -base encoding system color-space informatics 2 nd Base A C G T 0 1 2 3 1 0 3 2 G 2 3 0 1 T 3 2 1 0 A 1 st Base • • C
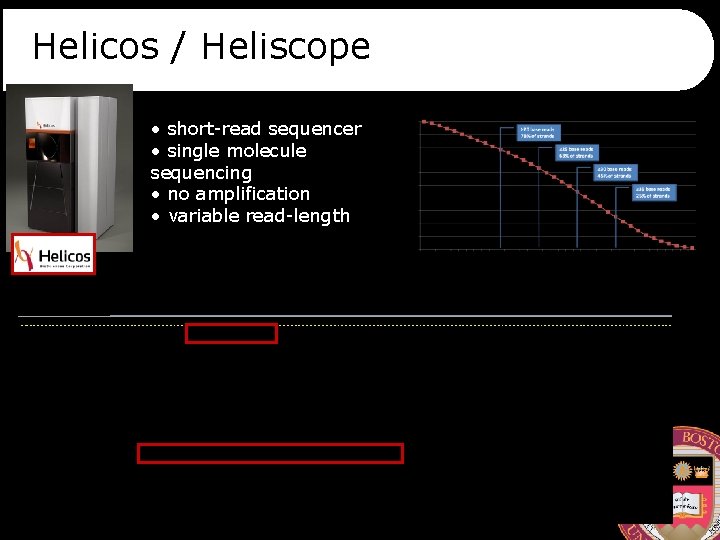
Helicos / Heliscope • short-read sequencer • single molecule sequencing • no amplification • variable read-length
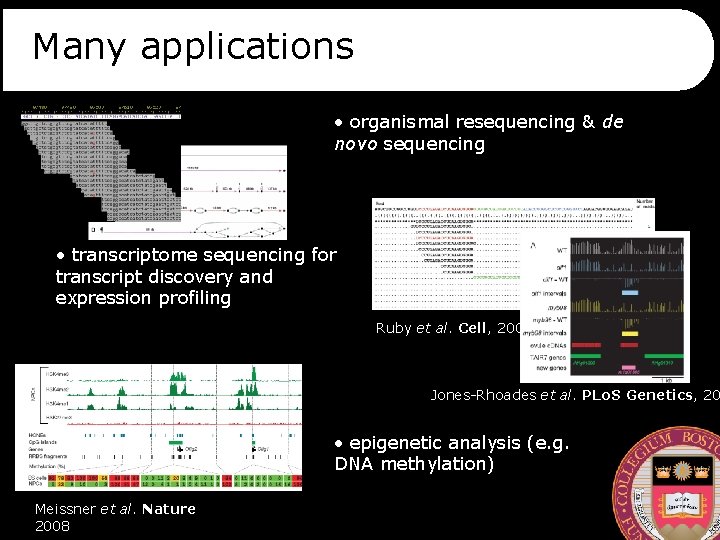
Many applications • organismal resequencing & de novo sequencing • transcriptome sequencing for transcript discovery and expression profiling Ruby et al. Cell, 2006 Jones-Rhoades et al. PLo. S Genetics, 20 • epigenetic analysis (e. g. DNA methylation) Meissner et al. Nature 2008

Data characteristics
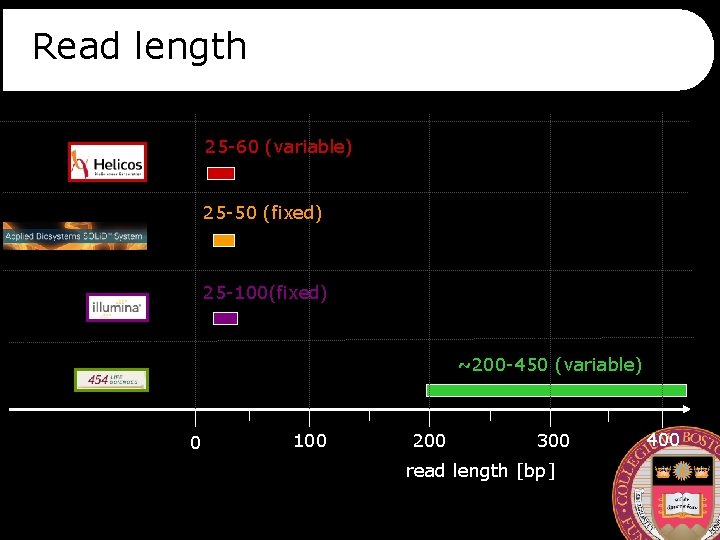
Read length 25 -60 (variable) 25 -50 (fixed) 25 -100(fixed) ~200 -450 (variable) 0 100 200 300 read length [bp] 400
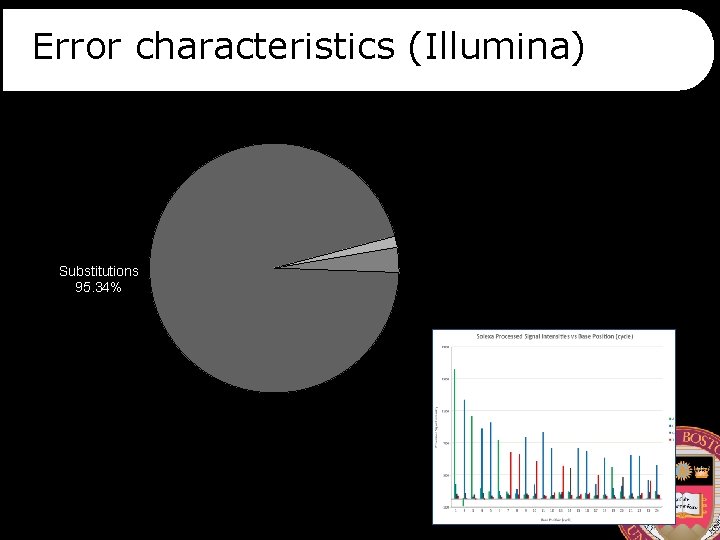
Error characteristics (Illumina) Insertions 1. 43% Substitutions 95. 34% Deletions 3. 23%
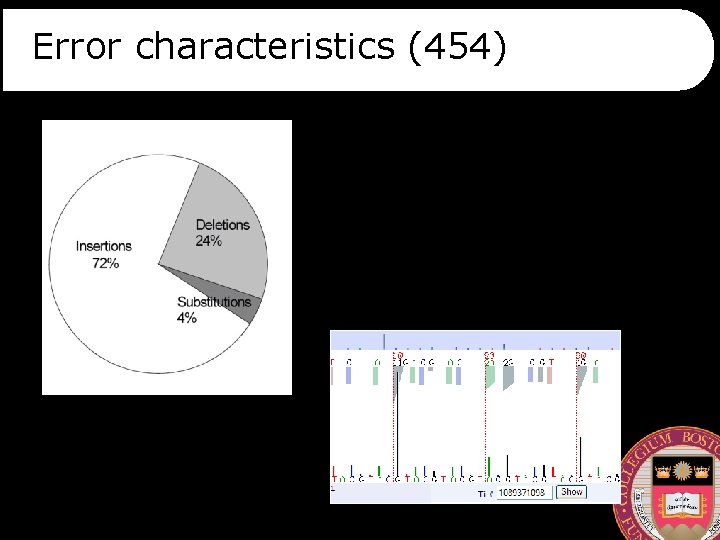
Error characteristics (454)
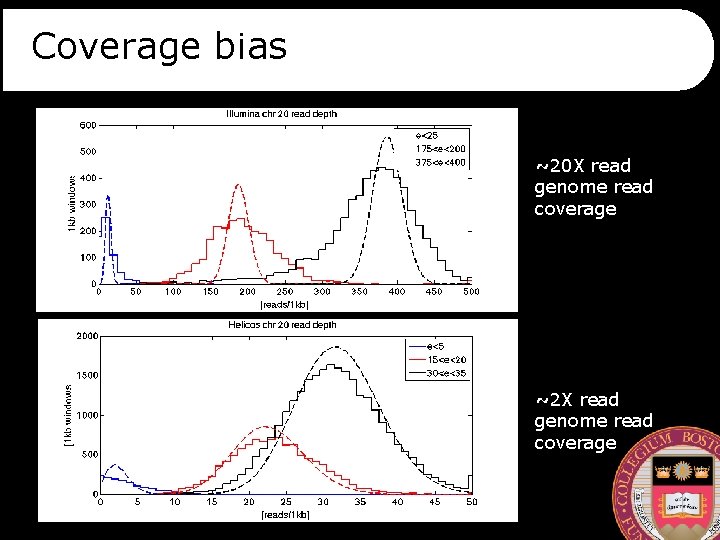
Coverage bias ~20 X read genome read coverage ~2 X read genome read coverage

Genome resequencing
Complete human genomes
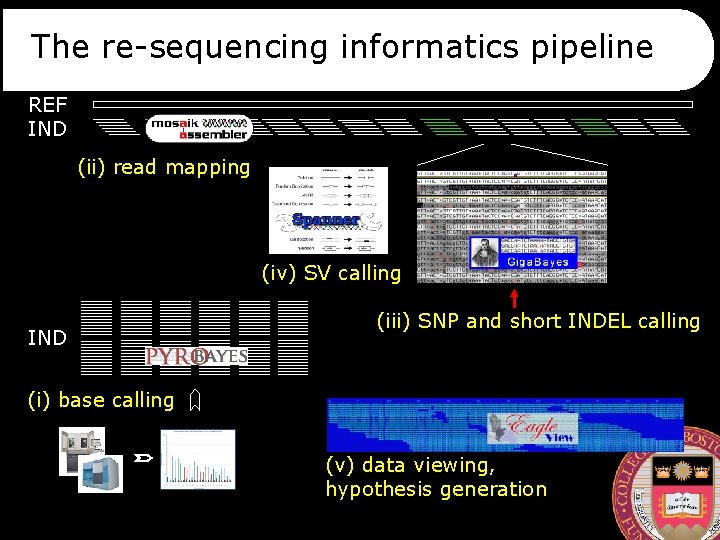
The re-sequencing informatics pipeline REF IND (ii) read mapping (iv) SV calling IND (iii) SNP and short INDEL calling (i) base calling (v) data viewing, hypothesis generation

Read mapping

… is like a jigsaw puzzle …you get the pieces… 2. Read mapping … and they give you the picture on the box Big and Unique pieces are easier to place than others…
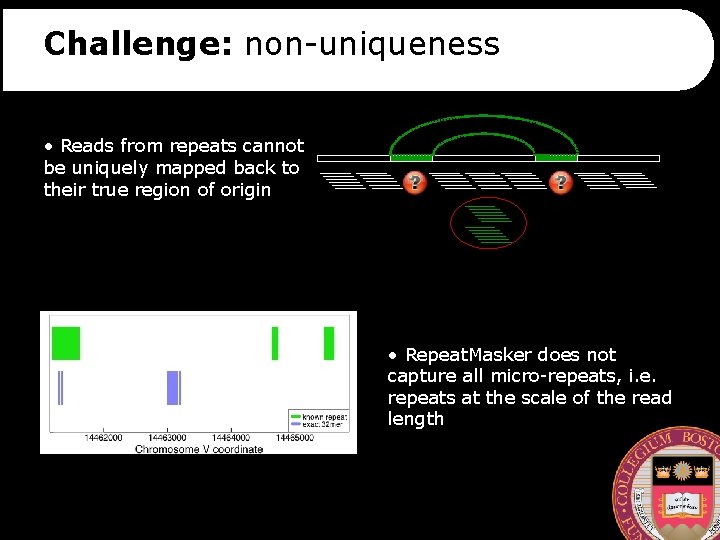
Challenge: non-uniqueness • Reads from repeats cannot be uniquely mapped back to their true region of origin • Repeat. Masker does not capture all micro-repeats, i. e. repeats at the scale of the read length
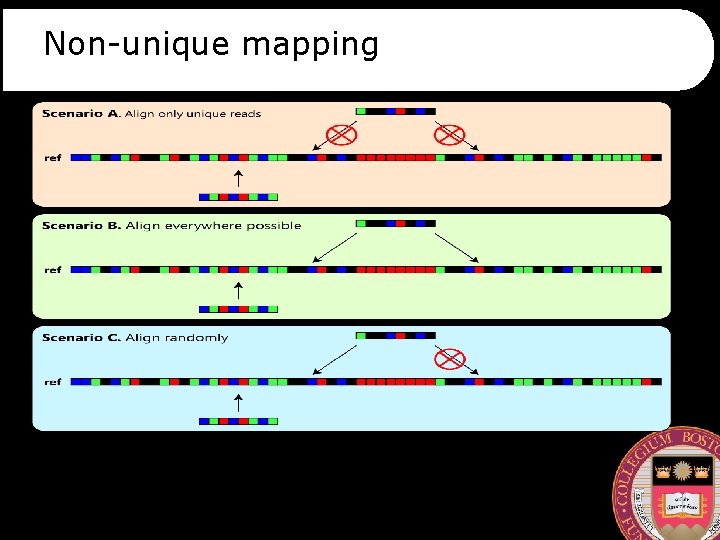
Non-unique mapping

SE short-read alignments are error-prone 0. 35%
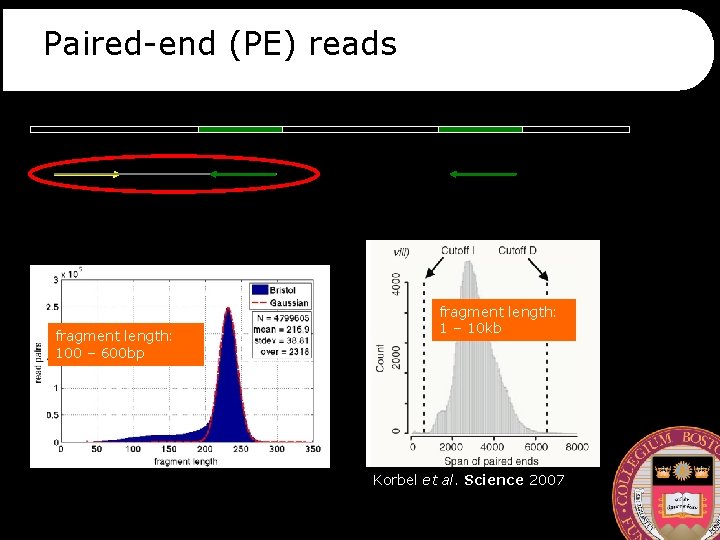
Paired-end (PE) reads fragment length: 100 – 600 bp fragment length: 1 – 10 kb Korbel et al. Science 2007
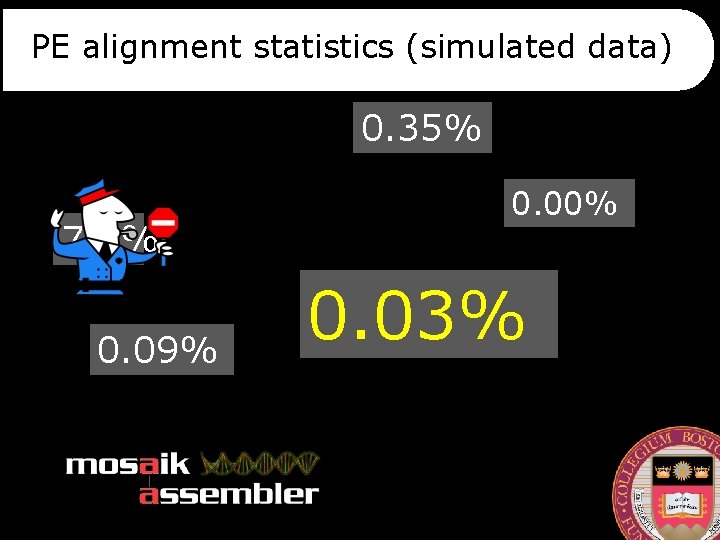
PE alignment statistics (simulated data) 0. 35% 7. 6% 0. 09% 0. 00% 0. 03%
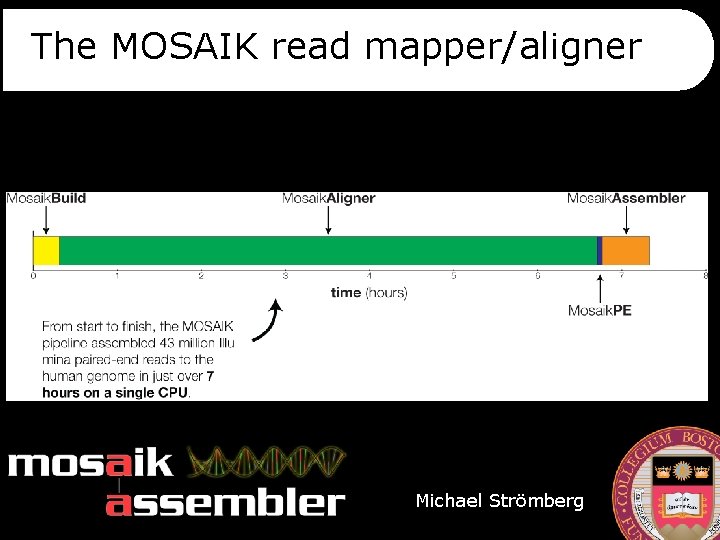
The MOSAIK read mapper/aligner Michael Strömberg
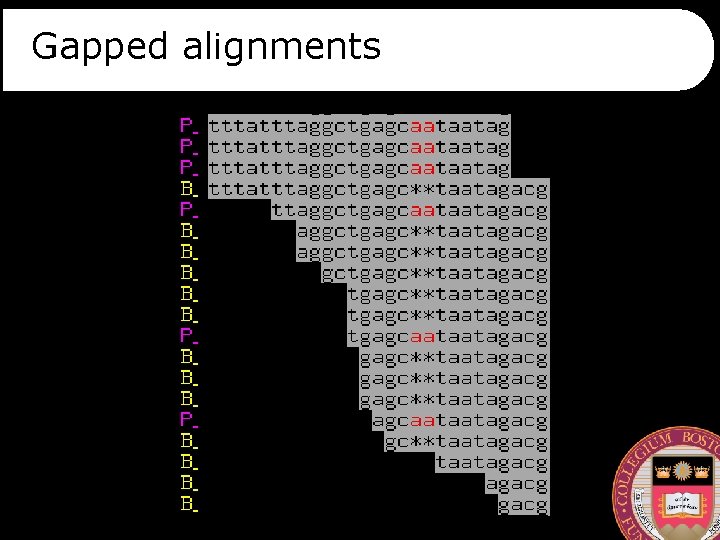
Gapped alignments
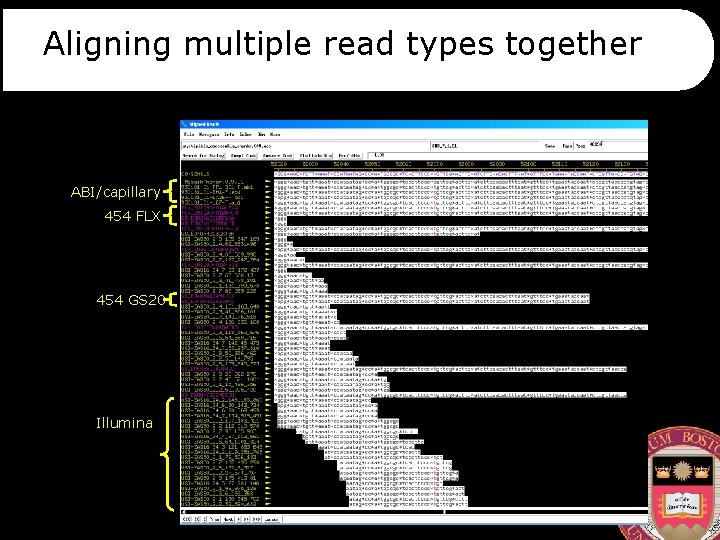
Aligning multiple read types together ABI/capillary 454 FLX 454 GS 20 Illumina

SNP / short-INDEL discovery

Polymorphism detection sequencing error polymorphism

Allele calling in multi-individual data P(B 1=aacc|G 1=aa) P(B 1=aacc|G 1=cc) P(B 1=aacc|G 1=ac) -----a---------a---------c----- P(Bi=aaaac|Gi=aa) P(Bi=aaaac|Gi=cc) P(Bi=aaaac|Gi=ac) -----c---------c----- P(Bn=cccc|Gn=aa) P(Bn=cccc|Gn=cc) P(Bn=cccc|Gn=ac) “genotype likelihoods” P(SNP) P(G 1=aa|B 1=aacc; Bi=aaaac; Bn= cccc) P(G 1=cc|B 1=aacc; Bi=aaaac; Bn= cccc) P(G 1=ac|B 1=aacc; Bi=aaaac; Bn= cccc) Prior(G 1, . . , Gi, . . , Gn) -----a---------c----- P(Gi=aa|B 1=aacc; Bi=aaaac; Bn= cccc) P(Gi=cc|B 1=aacc; Bi=aaaac; Bn= cccc) P(Gi=ac|B 1=aacc; Bi=aaaac; Bn= cccc) P(Gn=aa|B 1=aacc; Bi=aaaac; Bn= cccc) P(Gn=cc|B 1=aacc; Bi=aaaac; Bn= cccc) P(Gn=ac|B 1=aacc; Bi=aaaac; Bn= cccc) “genotype probabilities”

SNP calling in deep sample sets Population Samples Reads Allele detection
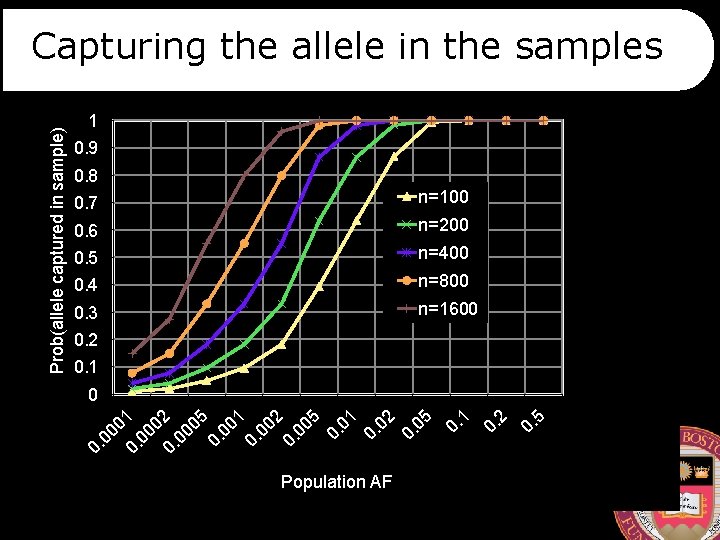
1 0. 9 0. 8 0. 7 n=100 0. 6 n=200 0. 5 n=400 0. 4 n=800 0. 3 n=1600 0. 2 0. 1 Population AF 5 0. 2 0. 1 0. 05 0. 02 0. 01 0. 5 00 2 0. 00 0. 1 00 0. 05 00 02 0. 00 01 0 0. Prob(allele captured in sample) Capturing the allele in the samples

The ability to call rare alleles aatgtagta. Agtacctac aatgtagta. Cgtacctac aatgtagta. Agtacctac aatgtagta. Agtacctac reads Q 30 Q 40 Q 50 Q 60 1 0. 01 0. 5 2 0. 82 1. 0 3 1. 0
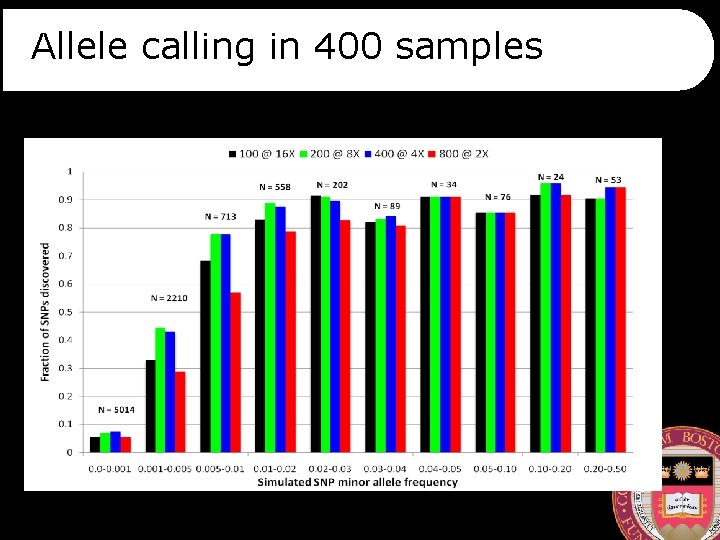
Allele calling in 400 samples
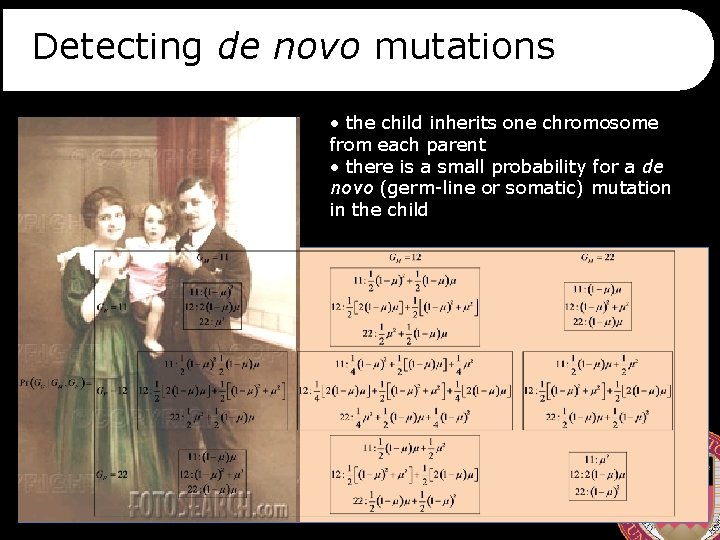
Detecting de novo mutations • the child inherits one chromosome from each parent • there is a small probability for a de novo (germ-line or somatic) mutation in the child
Capture sequencing
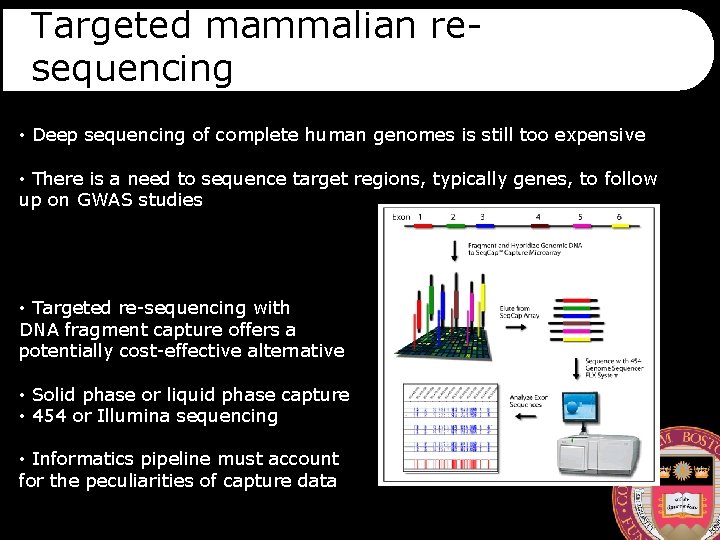
Targeted mammalian resequencing • Deep sequencing of complete human genomes is still too expensive • There is a need to sequence target regions, typically genes, to follow up on GWAS studies • Targeted re-sequencing with DNA fragment capture offers a potentially cost-effective alternative • Solid phase or liquid phase capture • 454 or Illumina sequencing • Informatics pipeline must account for the peculiarities of capture data
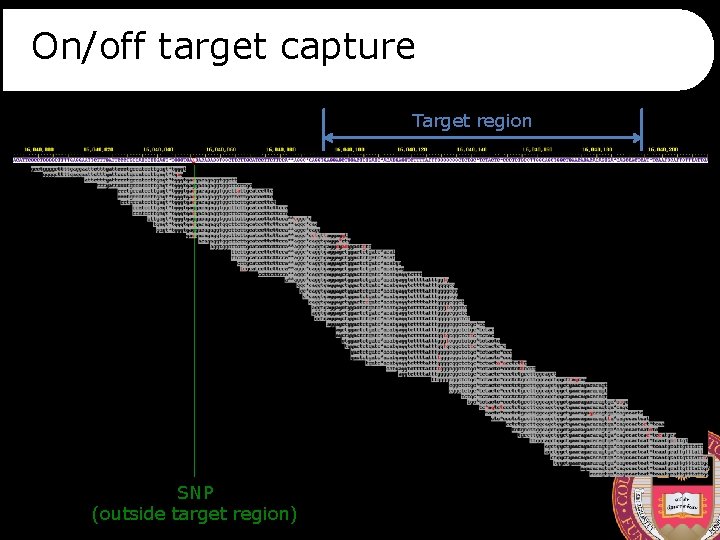
On/off target capture ref allele*: 45% Target region non-ref allele*: 54% SNP (outside target region)
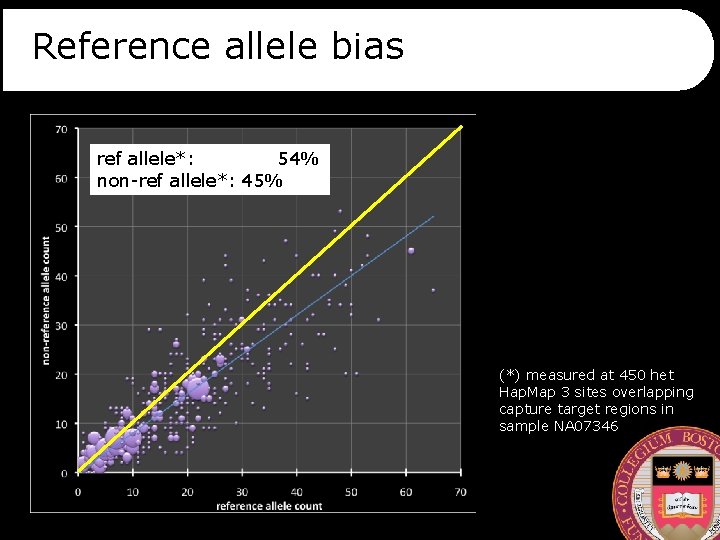
Reference allele bias ref allele*: 54% non-ref allele*: 45% (*) measured at 450 het Hap. Map 3 sites overlapping capture target regions in sample NA 07346

SNP example Amit Indap
Structural Variation discovery
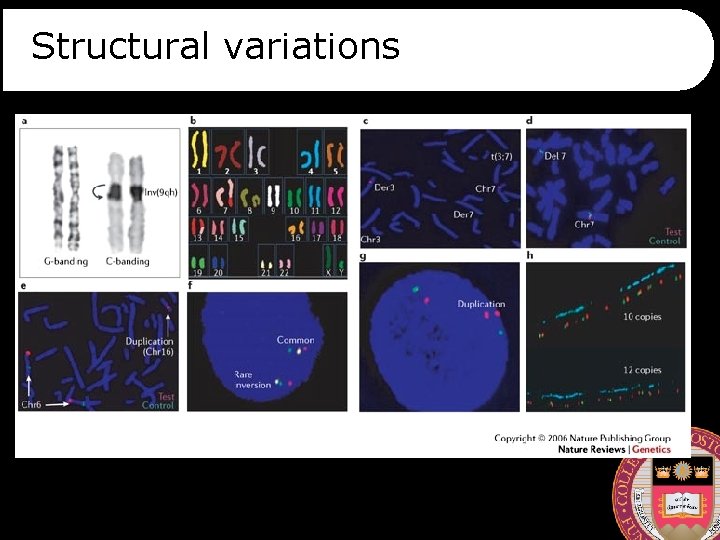
Structural variations
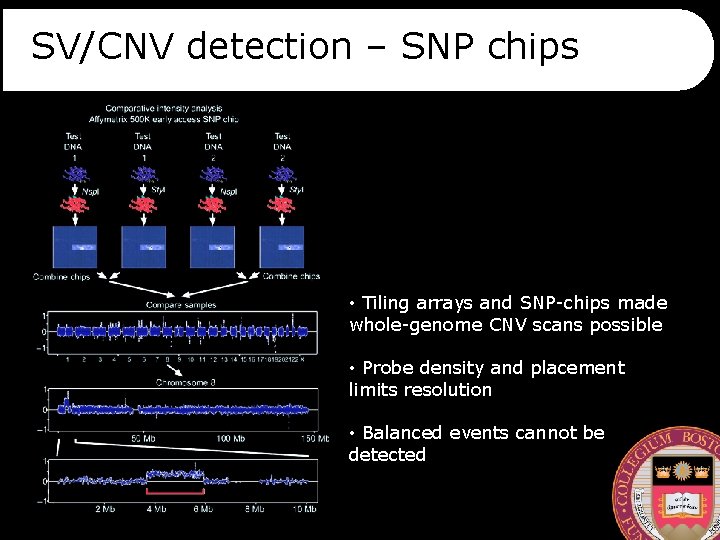
SV/CNV detection – SNP chips • Tiling arrays and SNP-chips made whole-genome CNV scans possible • Probe density and placement limits resolution • Balanced events cannot be detected
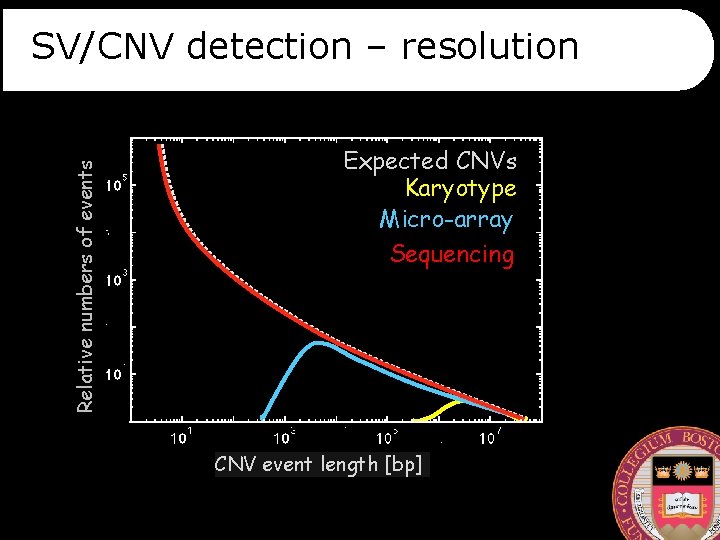
Relative numbers of events SV/CNV detection – resolution Expected CNVs Karyotype Micro-array Sequencing CNV event length [bp]
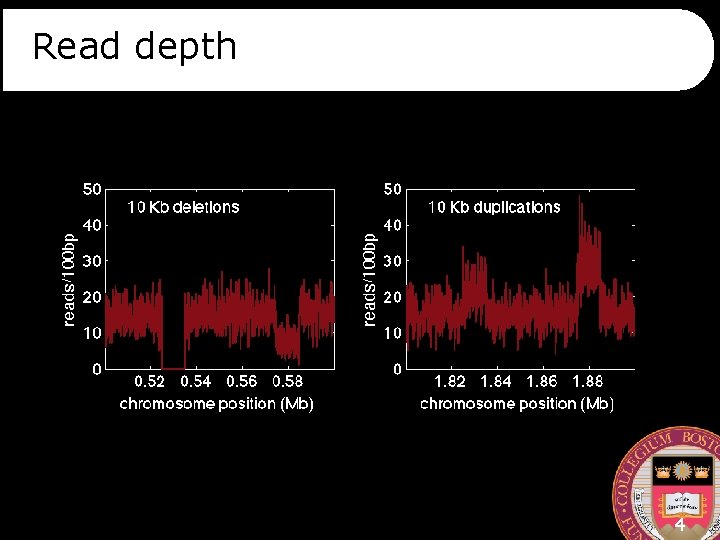
Read depth 4
![CNV events found using RD Chromosome 2 Position [Mb] CNV events found using RD Chromosome 2 Position [Mb]](http://slidetodoc.com/presentation_image/cba8e698ad88a64a131937d05c69fa60/image-45.jpg)
CNV events found using RD Chromosome 2 Position [Mb]
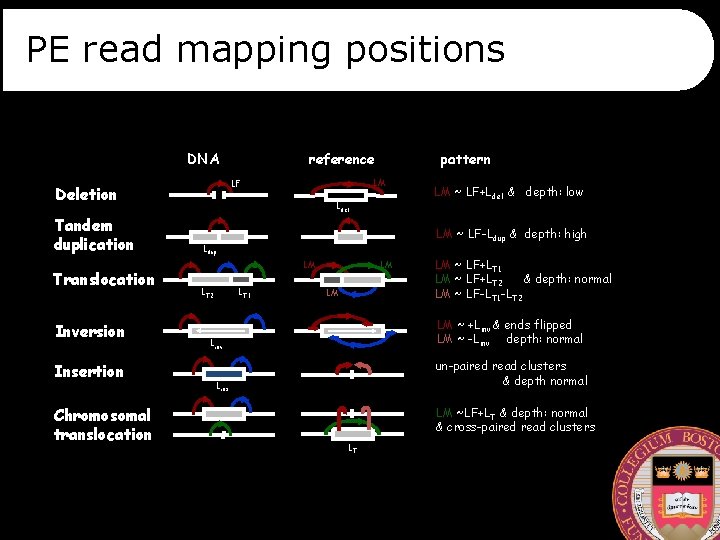
PE read mapping positions DNA Translocation Inversion Insertion Chromosomal translocation pattern LM LF Deletion Tandem duplication reference Ldel LM ~ LF-Ldup & depth: high Ldup LM LT 2 LM ~ LF+Ldel & depth: low LT 1 LM LM LM ~ LF+LT 1 LM ~ LF+LT 2 & depth: normal LM ~ LF-LT 1 -LT 2 LM ~ +Linv & ends flipped LM ~ -Linv depth: normal Linv un-paired read clusters & depth normal Lins LM ~LF+LT & depth: normal & cross-paired read clusters LT
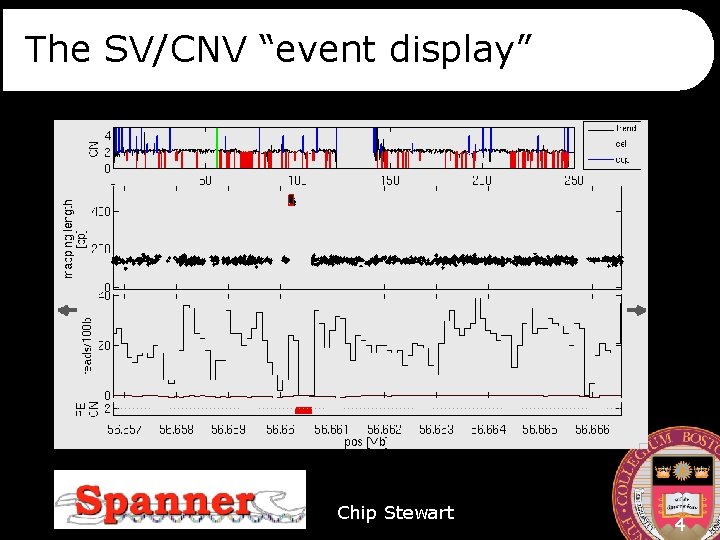
The SV/CNV “event display” Chip Stewart 4
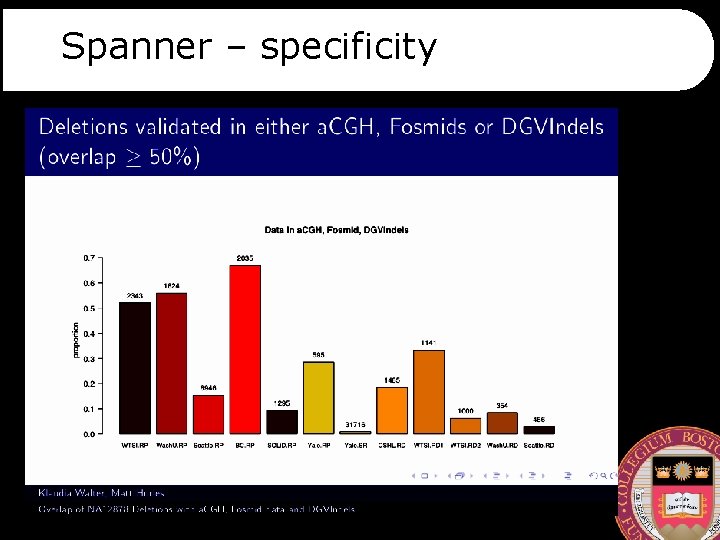
Spanner – specificity

Data standards
Data types with standard formats SRF/FASTQ GLF SAM/BAM

Transcriptome sequencing
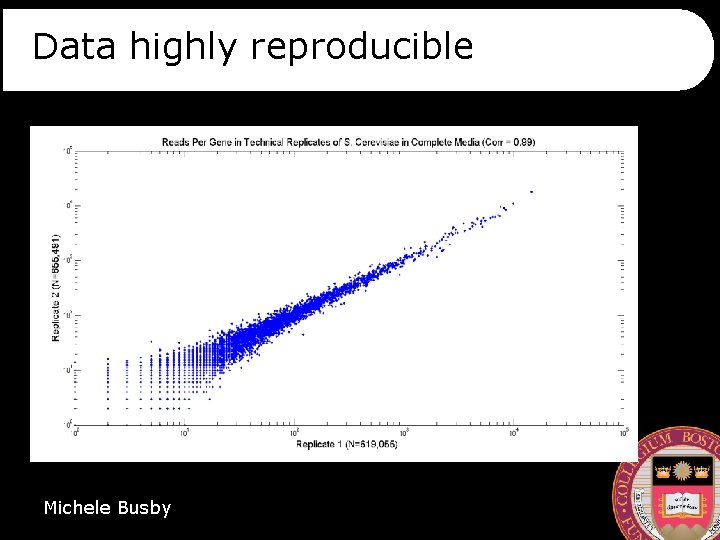
Data highly reproducible Michele Busby
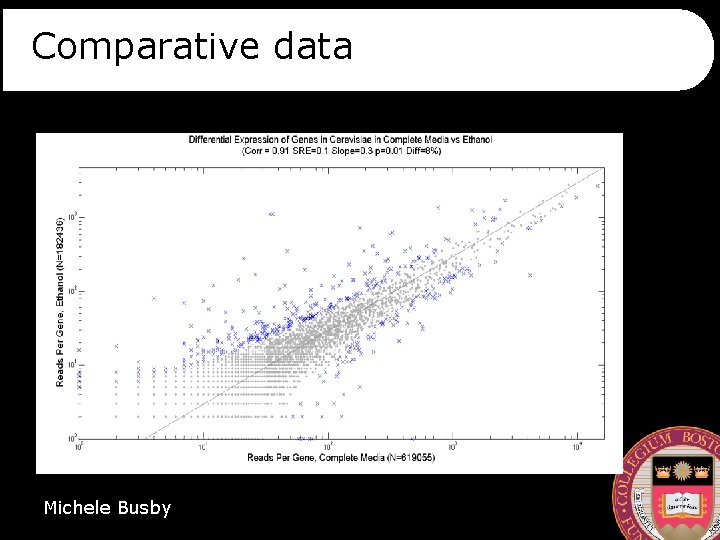
Comparative data Michele Busby
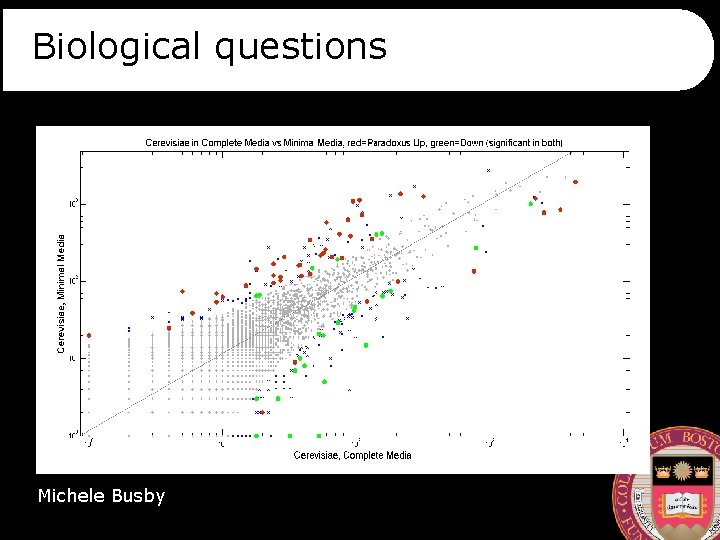
Biological questions Michele Busby
Our software tools for next-gen data http: //bioinformatics. bc. edu/marthlab/Software_Release

Credits Elaine Mardis Andy Clark Aravinda Chakravarti Doug Smith Michael Egholm Scott Kahn Francisco de la Vega Patrice Milos John Thompson

Lab Several postdoc positions are available!

Mutational profiling

Chemical mutagenesis
Mutational profiling: deep 454/Illumina/SOLi. D data Pichia stipitis reference sequence Image from JGI web site • Pichia stipitis converts xylose to ethanol (bio-fuel production) • one mutagenized strain had high conversion efficiency • determine which mutations caused this phenotype • 15 MB genome: 454, Illumina, and SOLi. D reads • 14 true point mutations in the entire genome 10 -15 X genome coverage required
- Slides: 60