Inferring Phylogenies from LCA distances back to the

Inferring Phylogenies from LCA distances (back to the basics of distance-based phylogenetic reconstruction) Ilan Gronau Shlomo Moran Technion, Israel 1 PLGW 01 - September 2007.
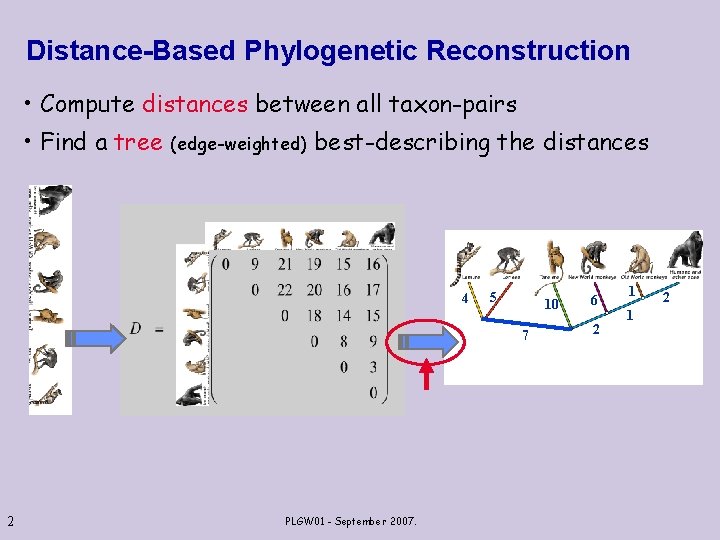
Distance-Based Phylogenetic Reconstruction • Compute distances between all taxon-pairs • Find a tree (edge-weighted) best-describing the distances 4 5 10 7 2 PLGW 01 - September 2007. 6 2 1 1 2

Distance-Based Reconstruction 4 5 10 7 Basics (Sanity check): 6 2 1 Reconstruction algorithms should be consistent, i. e. reconstruct the true tree from accurate (ie, additive) distances. Essential Extras: ØRobustness to noise : Reconstruct the correct tree (or parts of it) given noisy distances. Ø Efficiency: 3 Low time/space complexity. PLGW 01 - September 2007.

Neighbor Joining Methods An agglomerative clustering approach: • A taxon-pair i, j is chosen and replaced by a new taxon v • i, j are connected to new taxon v (i. e. are cherries in the reconstructed tree) • 4 Method recursively applied on reduced matrix PLGW 01 - September 2007.

The Two Basic Components of NJ Methods At each iteration the algorithm performs: 1. Selection: select neighboring taxons consistency: if input is additive, selected taxa are cherries in the corresponding tree 2. Reduction: compute distances from the new taxon consistency: the reduced matrix should fit the reduced tree. usually can be achieved in more than one way 5 PLGW 01 - September 2007.

Saitou & Nei’s NJ Algorithm (1987) Saitou & Nei’s selection criterion Ø Robustness: Considered highly reliable in practice Ø Time complexity - θ(n 3) Ø ~13, 000 citations (Science Citation Index) Ø Implemented in numerous phylogenetic packages Questions: Ø What makes Saitou&Nei’s neighbor selection criterion so good? Ø Is there any simpler consistent neighbor-selection criterion? 6 PLGW 01 - September 2007.
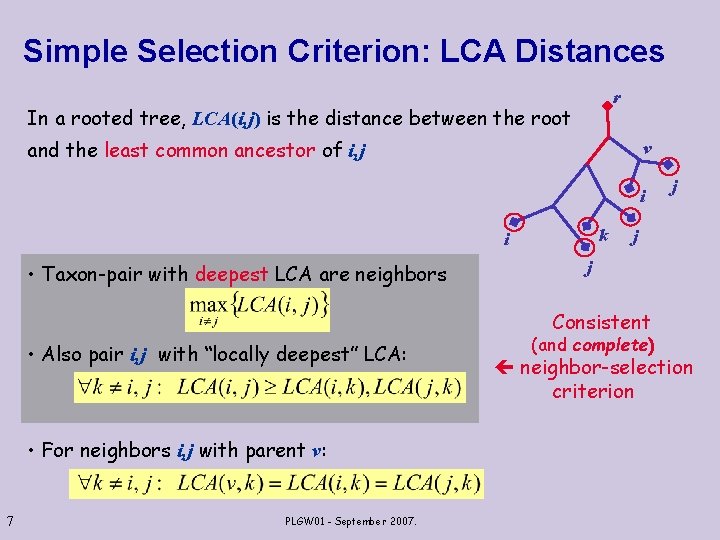
Simple Selection Criterion: LCA Distances r In a rooted tree, LCA(i, j) is the distance between the root and the least common ancestor of i, j v i k i • Taxon-pair with deepest LCA are neighbors j j j Consistent • Also pair i, j with “locally deepest” LCA: • For neighbors i, j with parent v: 7 PLGW 01 - September 2007. (and complete) neighbor-selection criterion
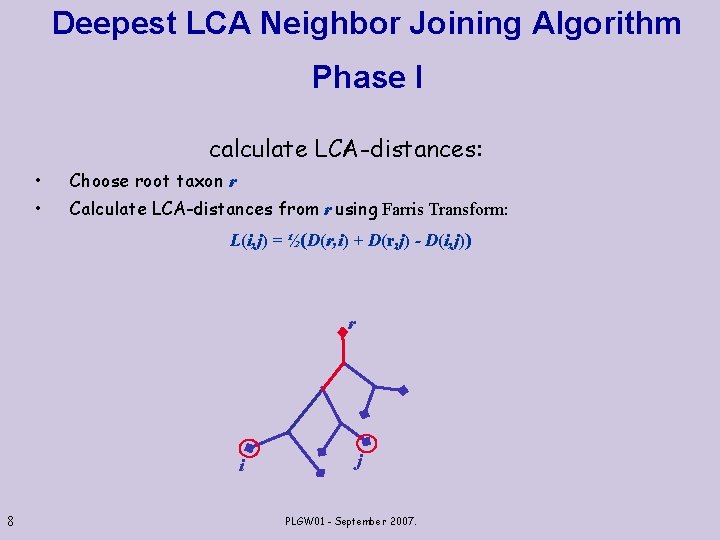
Deepest LCA Neighbor Joining Algorithm Phase I calculate LCA-distances: • Choose root taxon r • Calculate LCA-distances from r using Farris Transform: L(i, j) = ½(D(r, i) + D(r, j) - D(i, j)) r i 8 j PLGW 01 - September 2007.

Deepest LCA Neighbor Joining Algorithm Phase II n -1 neighbor-joining iterations At each iteration: • • • Selection: Choose taxon pair i, j , s. t. L(i, j) = maxi’≠j’{ L(i’, j’) } Connect i, j to new taxon v Reduction: Replace i, j with new taxon v, and reduce L: For k≠v, L(v, k) = αL(i, k) + (1 -α)L(j, k) (α – reduction parameter, may be re-defined each iteration ) 9 PLGW 01 - September 2007.

Simple and Optimal θ(n 2) Implementation of DLCA • Calculating LCA-distances (the matrix L) • Neighbor joining algorithm: - θ(n 2) time n-1 neighbor joining iterations: - Reduction step takes O(n) time per iteration - Bottleneck is in neighbor selection An amortized θ(n 2) implementation of neighbor selection: Join “locally deepest” pair and not necessarily “globally deepest” pair, using the “Nearest Neighbor Chain” clustering technique [Benzecri 82, Juan 82, Murtagh 84, +] 10 PLGW 01 - September 2007.

DLCA: Intermediate Summary • A simple and intuitive consistent neighbor selection criterion • Implemented in optimal time complexity (faster than NJ) What about the noise ? ! Robustness to noise: We consider two theoretical criteria for robustness: • Reconstruction of “Buneman edges” • Atteson’s “edge-reconstruction radius” 11 PLGW 01 - September 2007.
![Buneman’s Edges [Buneman ’ 71] • An edge e induces a split (P|Q) of Buneman’s Edges [Buneman ’ 71] • An edge e induces a split (P|Q) of](http://slidetodoc.com/presentation_image/f0c0cf3f1a2d1bf75be4fd400f6855ba/image-12.jpg)
Buneman’s Edges [Buneman ’ 71] • An edge e induces a split (P|Q) of the taxon set i e is a “Buneman edge” (w. r. t. Distance matrix P D) iff all taxon-quartet (i, j, k, l) which e “crosses” e (i. e. i, j ∊ P, k, l ∊ Q ) agree with e’s split: D (i, j)+D (k, l) < D (i, k)+D (j, l) , D (i, l)+D (j, k) Q l k “Buneman Robustness criterion”: the algorithm should reconstruct all the Buneman edges. 12 PLGW 01 - September 2007. j
![Atteson’s Edge-Reconstruction radius [Atteson ‘ 99] Edge reconstruction-radius: A has edge-reconstruction radius of ε Atteson’s Edge-Reconstruction radius [Atteson ‘ 99] Edge reconstruction-radius: A has edge-reconstruction radius of ε](http://slidetodoc.com/presentation_image/f0c0cf3f1a2d1bf75be4fd400f6855ba/image-13.jpg)
Atteson’s Edge-Reconstruction radius [Atteson ‘ 99] Edge reconstruction-radius: A has edge-reconstruction radius of ε if for each edge e: If ||D-DT||∞ < ε∙w (e): w(e) Noise≤ εw(e) (for all distances) Then A correctly reconstructs e. Atteson: edge-reconstruction radius ≤ ½ e ØA satisfies Buneman’s criterion A has optimal edge-radius of ½ 13 PLGW 01 - September 2007.

Robustness of NJ and DLCA • NJ: -edge-reconstruction radius = ¼ [Atteson ’ 99, Mihaescu et al ‘ 06] (hence it does not satisfy the Buneman Criterion) • DLCA (using “conservative reductions”): - Satisfies the Buneman Criterion - Hence it has edge-reconstruction radius = ½ By these criteria, DLCA is also more robust than NJ And in Practice…? ? ? 14 PLGW 01 - September 2007.
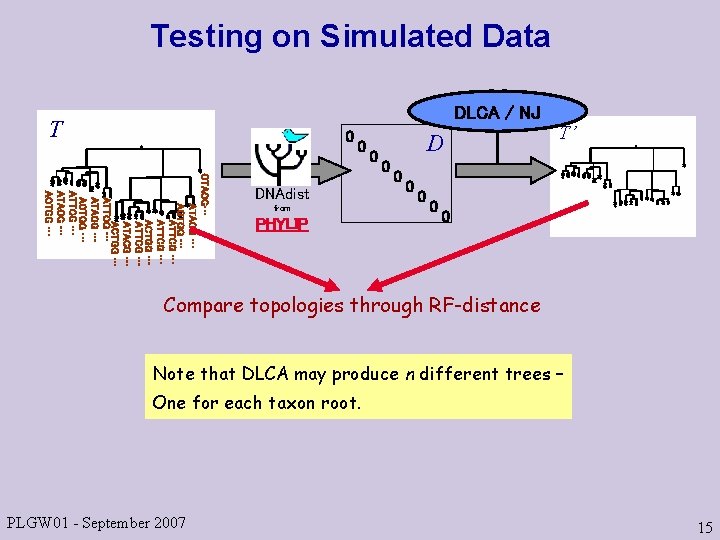
Testing on Simulated Data DLCA / NJ T 0 0 CTACG… ATACG … AGTGG … ATTCG … ACTGG … ATTCG … ATACG … ACTGG … DNAdist from PHYLIP 0 D 0 0 0 T’ 0 Compare topologies through RF-distance Note that DLCA may produce n different trees – One for each taxon root. PLGW 01 - September 2007 15
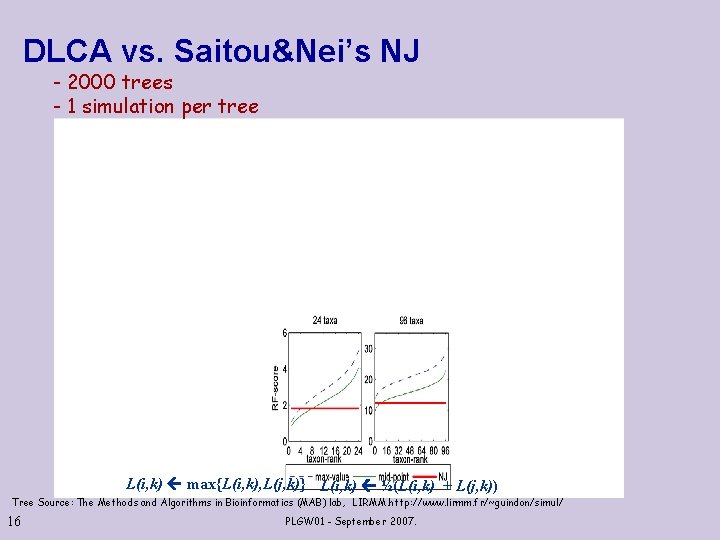
DLCA vs. Saitou&Nei’s NJ - 2000 trees - 1 simulation per tree L(i, k) max{L(i, k), L(j, k)} L(i, k) ½(L(i, k) + L(j, k)) Tree Source: The Methods and Algorithms in Bioinformatics (MAB) lab, LIRMM. http: //www. lirmm. fr/~guindon/simul/ 16 PLGW 01 - September 2007.

Robustness of DLCA – a Summary DLCA is superior to NJ by Buneman&Atteson criteria, but (on the average) is inferior to NJ on simulated data. Where lies the reason for this “conflict”? Take another look at Saitou &Nei selection criterion 17 PLGW 01 - September 2007.

Saitou & Nei’s Selection Criterion… … expressed by LCA distances i. e. , NJ tends to selects taxon-pairs with average deepest lca • Averaging “smoothes” noise • Averaging does not affect worst-case noise (The bound 1/4 on the reconstruction radius of NJ uses an highly improbable scenario) 18 PLGW 01 - September 2007.

Future Directions Use pivotal nature of DLCA to achieve better results: • Pre-processing: use “good” taxa as roots • Post-processing: return “best” tree among n possible outputs. Find robustness criteria which explain the robustness of NJ: • Instead of considering worst-case noise (as Atteson’s criterion), consider stochastic noise. 19 PLGW 01 - September 2007.

For more information… • "Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances" (JCB 14(1) pp. 1 -15 , 2007) • "Optimal Implementations of UPGMA and Other Common Clustering Algorithms” (to Appear in IPL) • Our websites: www. cs. technion. ac. il/~ilangr www. cs. technion. ac. il/~moran 20 PLGW 01 - September 2007.

Thank You PLGW 01 - September 2007 21
- Slides: 21