Improving peptide identification for tandem mass spectrometry by

Improving peptide identification for tandem mass spectrometry by incorporating translatomics information Chuan-Le Xiao (肖传乐) 中山大学眼科学国家重点�� 室 E-mail: xiaochuanle@126. com
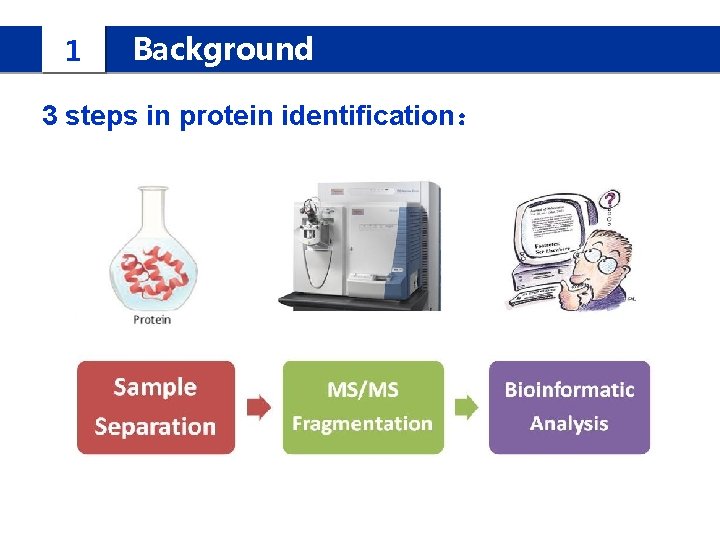
1 Background 3 steps in protein identification:
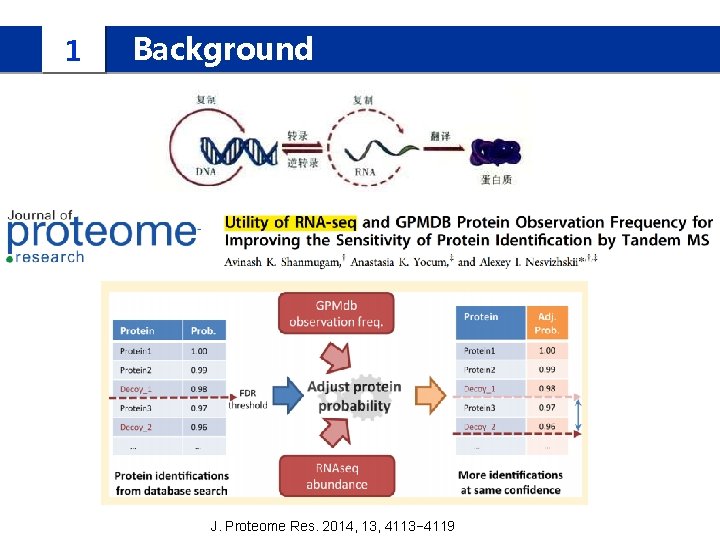
1 Background J. Proteome Res. 2014, 13, 4113− 4119
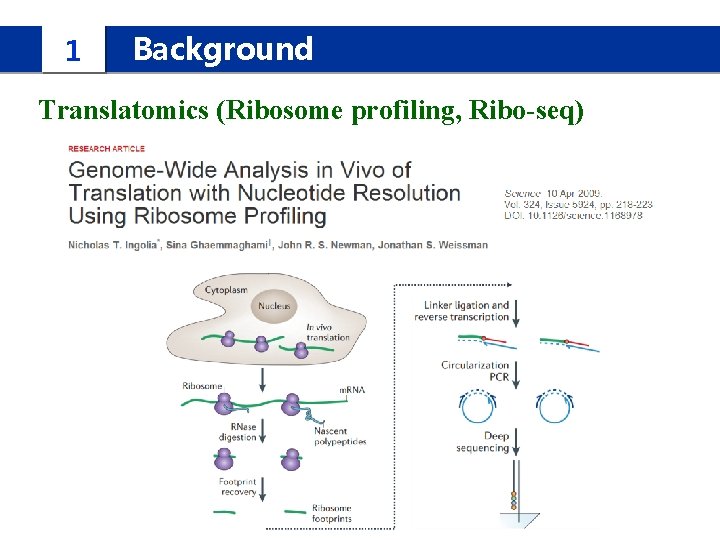
1 Background Translatomics (Ribosome profiling, Ribo-seq)
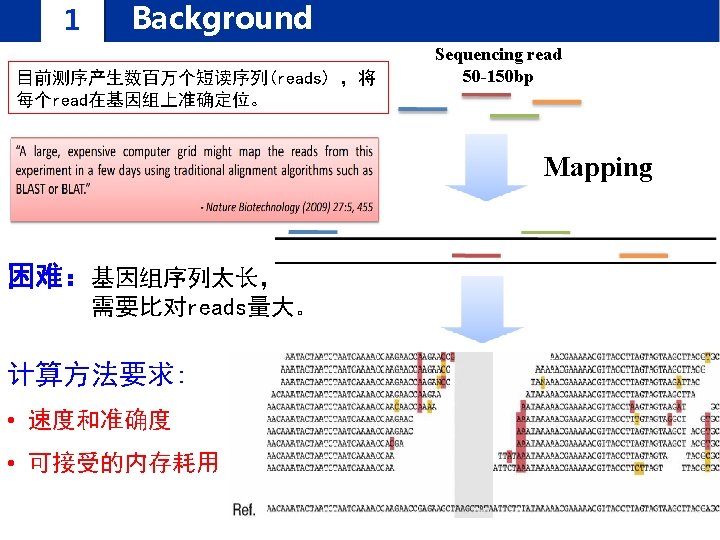
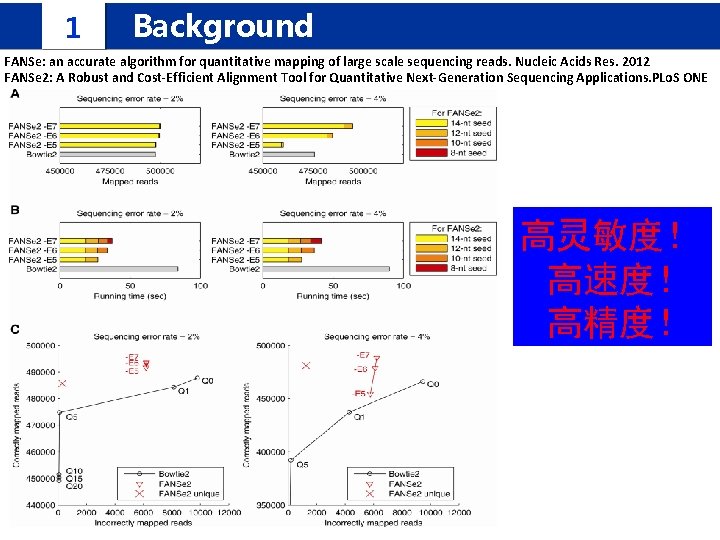
1 Background FANSe: an accurate algorithm for quantitative mapping of large scale sequencing reads. Nucleic Acids Res. 2012 FANSe 2: A Robust and Cost-Efficient Alignment Tool for Quantitative Next-Generation Sequencing Applications. PLo. S ONE 高灵敏度! 高速度! 高精度!
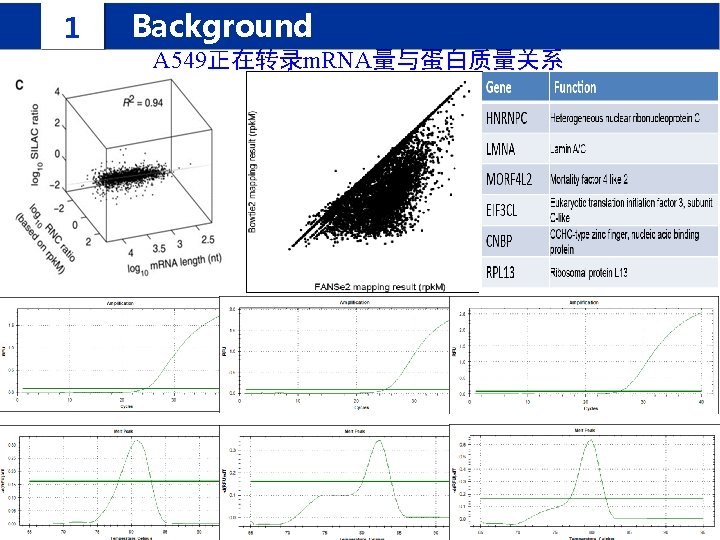
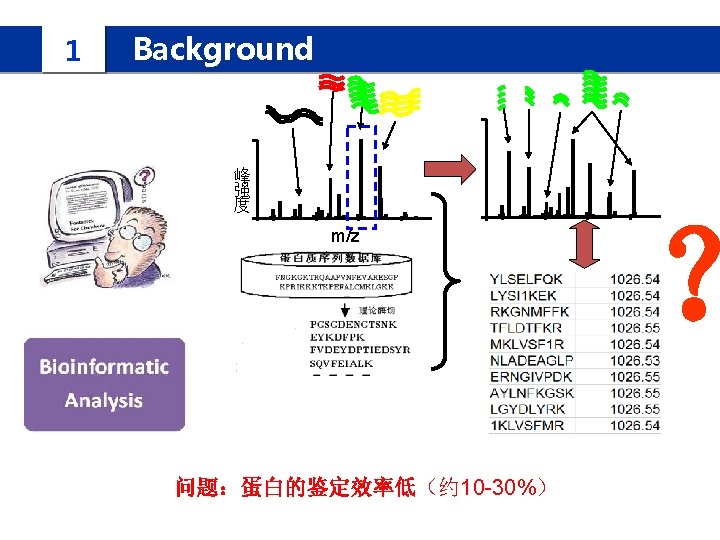
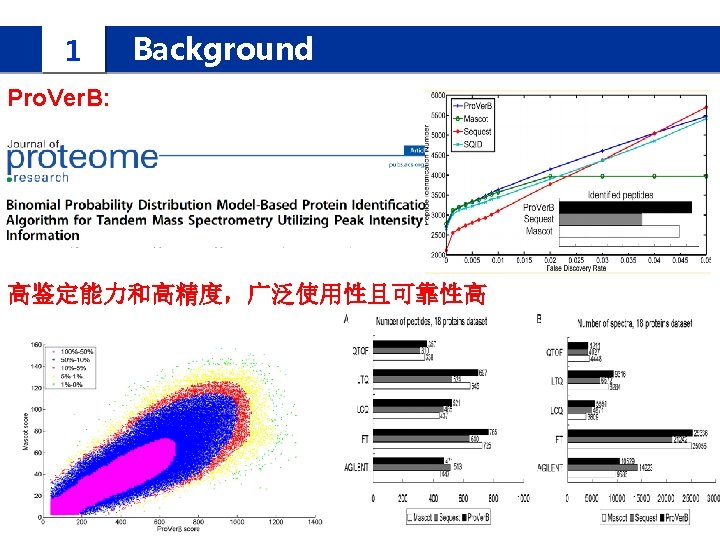
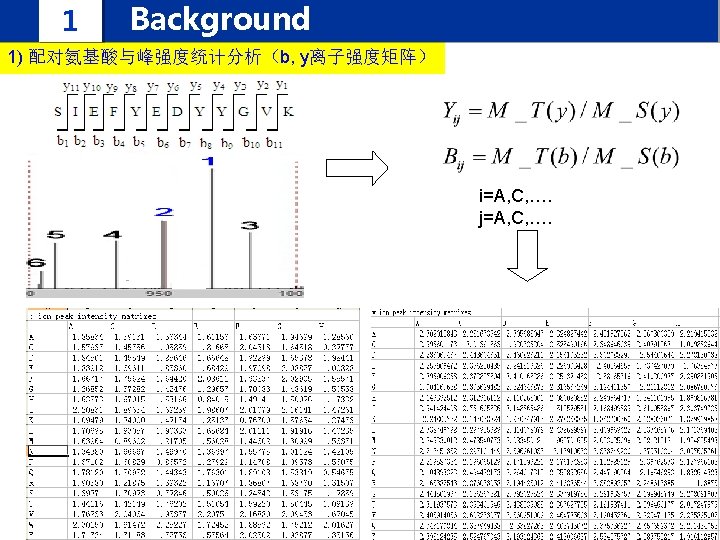
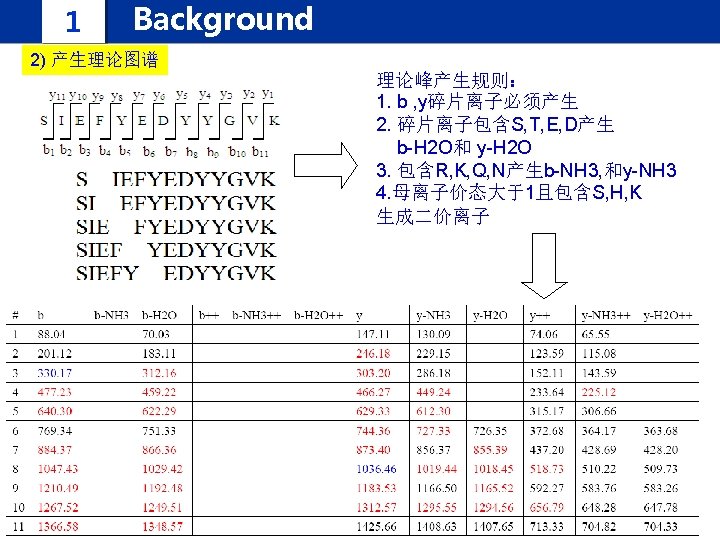

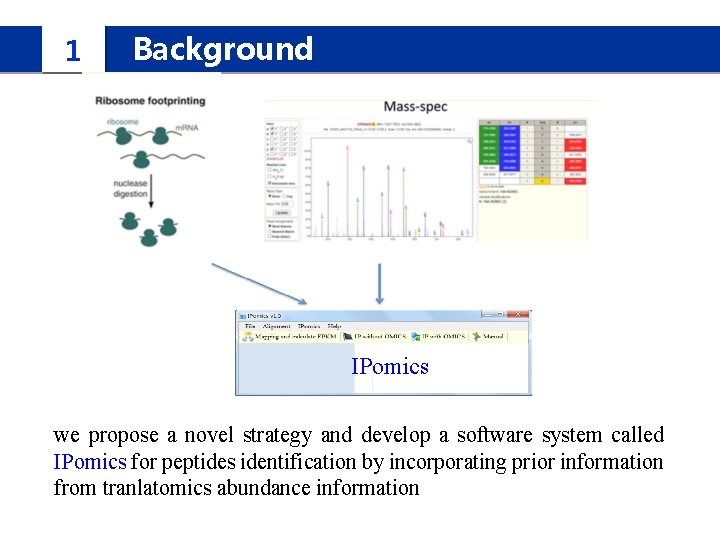
1 Background IPomics we propose a novel strategy and develop a software system called IPomics for peptides identification by incorporating prior information from tranlatomics abundance information
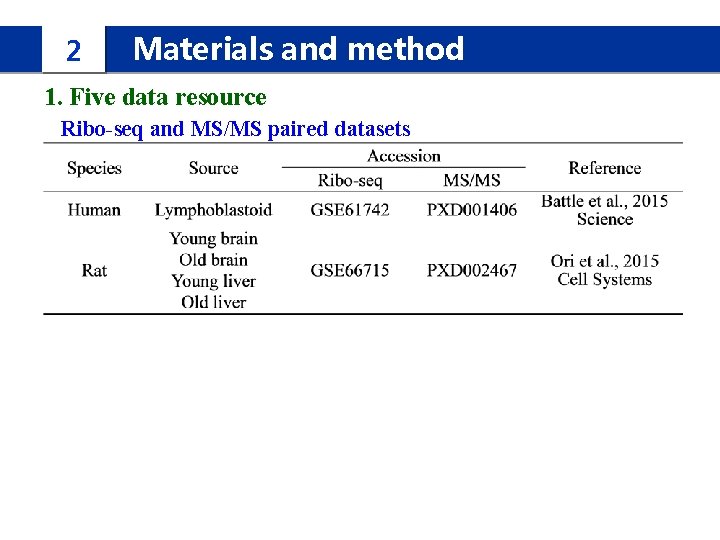
2 Materials and method 1. Five data resource Ribo-seq and MS/MS paired datasets
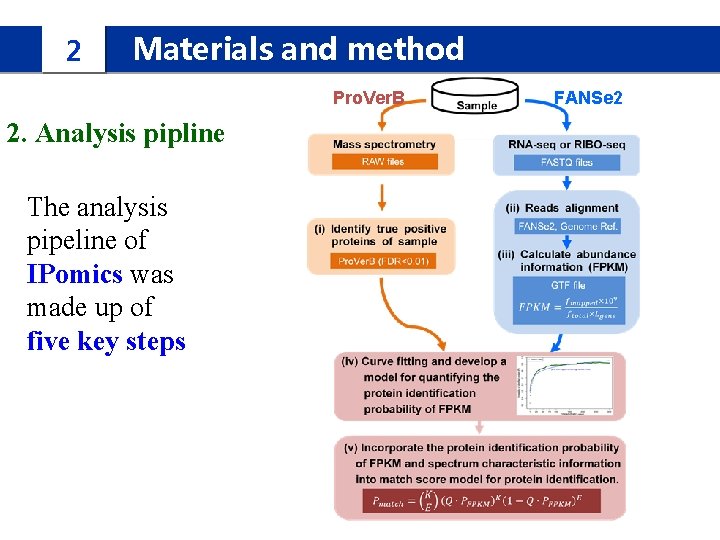
2 Materials and method Pro. Ver. B 2. Analysis pipline The analysis pipeline of IPomics was made up of five key steps FANSe 2

3 RESULTS 1. The prior information of FPKM for protein identification 2. The incorporation of tranlatomic FPKM in scoring model 3. Comparison of IPomics with Mascot, OMSSA, X!Tandem and Pfind 4. Computational validation with SILAC and Tyrosine phosphorylation datasets
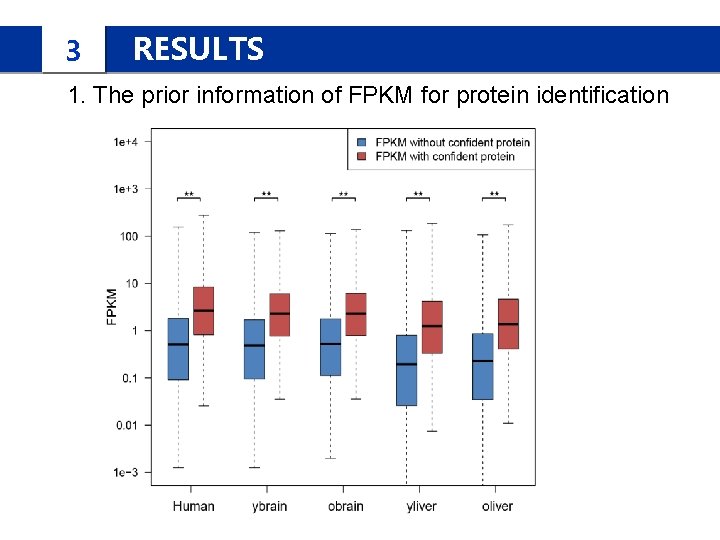
3 RESULTS 1. The prior information of FPKM for protein identification
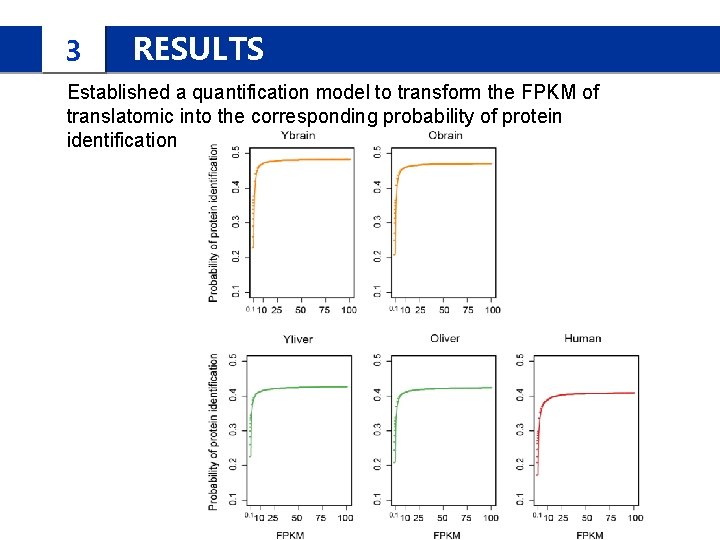
3 RESULTS Established a quantification model to transform the FPKM of translatomic into the corresponding probability of protein identification
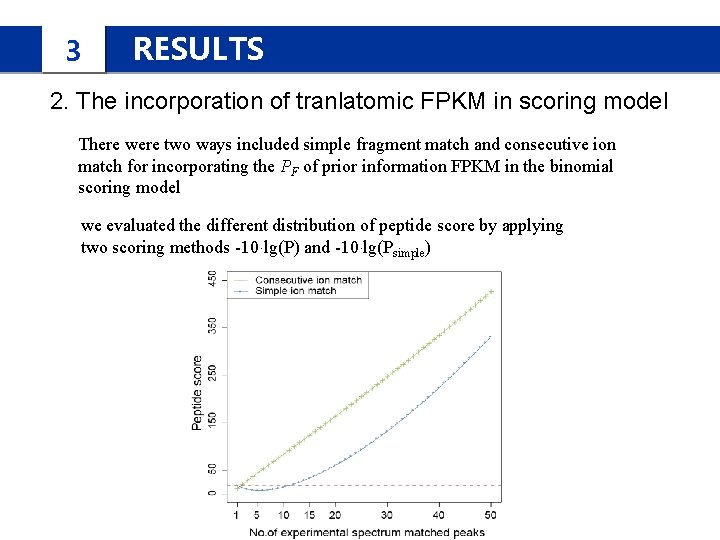
3 RESULTS 2. The incorporation of tranlatomic FPKM in scoring model There were two ways included simple fragment match and consecutive ion match for incorporating the PF of prior information FPKM in the binomial scoring model we evaluated the different distribution of peptide score by applying two scoring methods -10·lg(P) and -10·lg(Psimple)
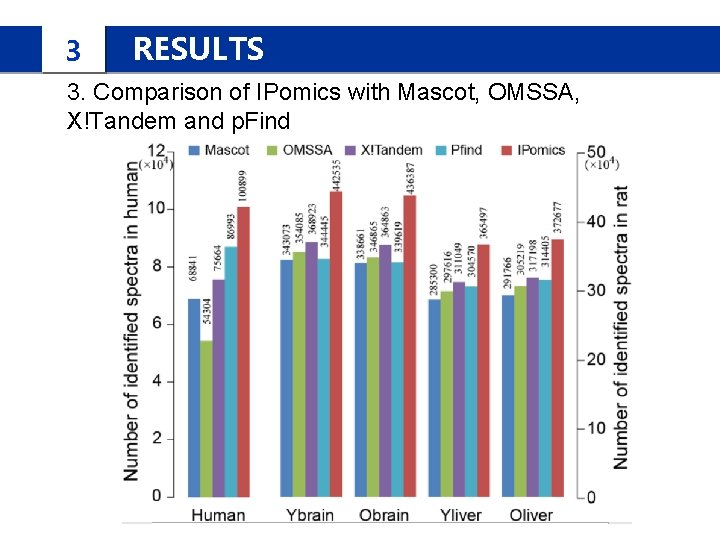
3 RESULTS 3. Comparison of IPomics with Mascot, OMSSA, X!Tandem and p. Find
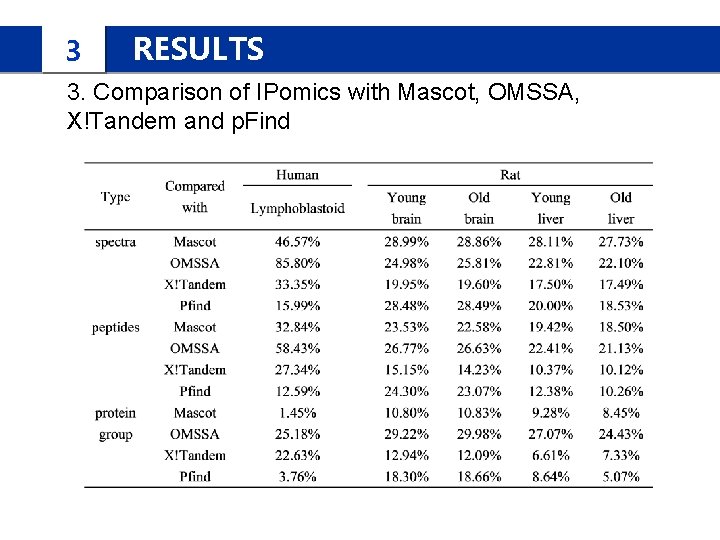
3 RESULTS 3. Comparison of IPomics with Mascot, OMSSA, X!Tandem and p. Find
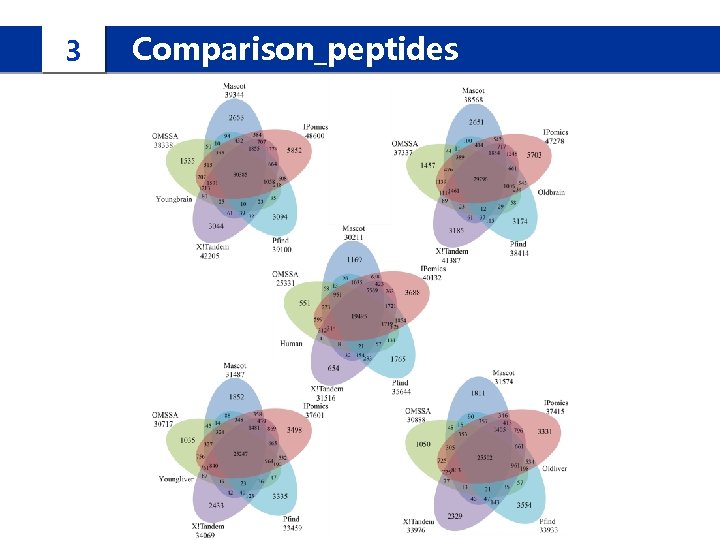
3 Comparison_peptides
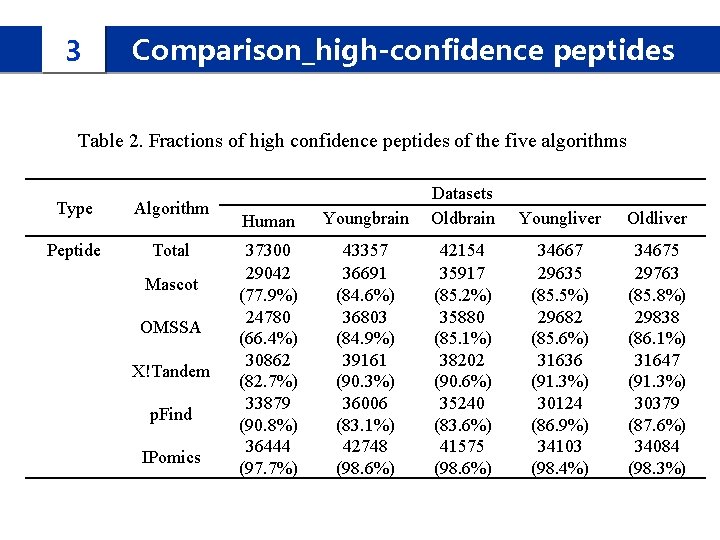
3 Comparison_high-confidence peptides Table 2. Fractions of high confidence peptides of the five algorithms Type Algorithm Peptide Total Mascot OMSSA X!Tandem p. Find IPomics Human Youngbrain Datasets Oldbrain 37300 29042 (77. 9%) 24780 (66. 4%) 30862 (82. 7%) 33879 (90. 8%) 36444 (97. 7%) 43357 36691 (84. 6%) 36803 (84. 9%) 39161 (90. 3%) 36006 (83. 1%) 42748 (98. 6%) 42154 35917 (85. 2%) 35880 (85. 1%) 38202 (90. 6%) 35240 (83. 6%) 41575 (98. 6%) Youngliver Oldliver 34667 29635 (85. 5%) 29682 (85. 6%) 31636 (91. 3%) 30124 (86. 9%) 34103 (98. 4%) 34675 29763 (85. 8%) 29838 (86. 1%) 31647 (91. 3%) 30379 (87. 6%) 34084 (98. 3%)
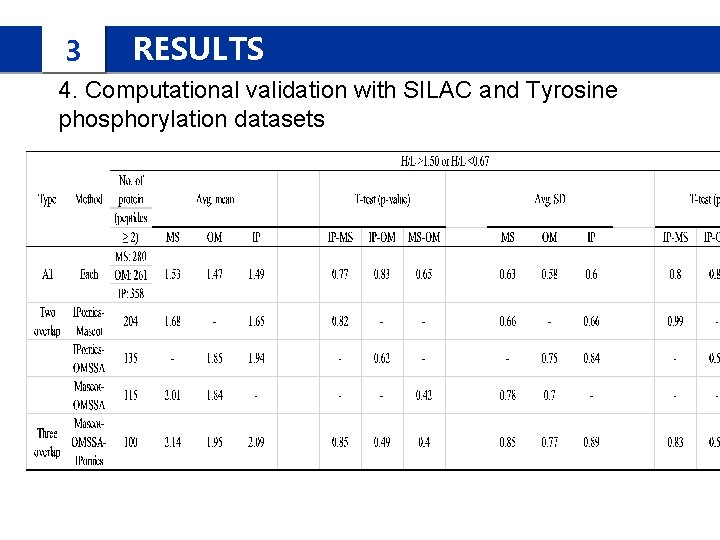
3 RESULTS 4. Computational validation with SILAC and Tyrosine phosphorylation datasets
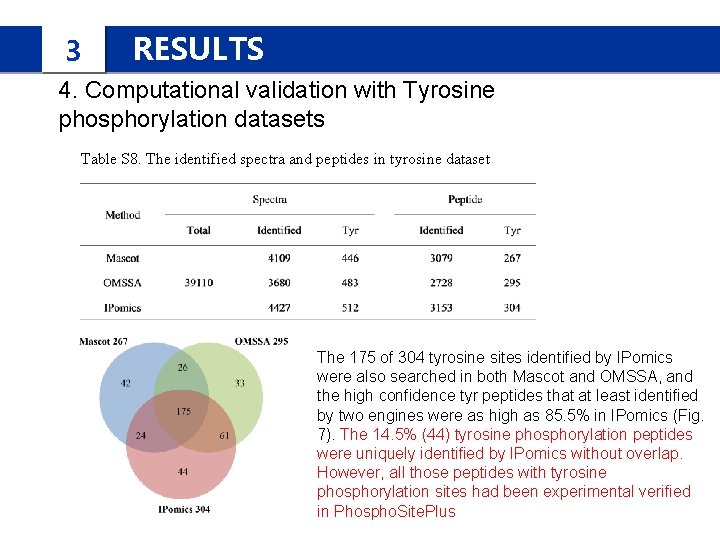
3 RESULTS 4. Computational validation with Tyrosine phosphorylation datasets Table S 8. The identified spectra and peptides in tyrosine dataset The 175 of 304 tyrosine sites identified by IPomics were also searched in both Mascot and OMSSA, and the high confidence tyr peptides that at least identified by two engines were as high as 85. 5% in IPomics (Fig. 7). The 14. 5% (44) tyrosine phosphorylation peptides were uniquely identified by IPomics without overlap. However, all those peptides with tyrosine phosphorylation sites had been experimental verified in Phospho. Site. Plus

Thanks!
- Slides: 26