Improved saccharification efficiency of alfalfa for bioethanol production

Improved saccharification efficiency of alfalfa for bioethanol production Annick Bertrand, Yves Castonguay, Annie Claessens, Jean Cloutier Agriculture and Agri-Food Canada, Québec City

Perennial forage crops Perennial vs annual crops: v Require fewer inputs, v Increase carbon sequestration v Low management v Reduce erosion v Grow on marginal lands

Alfalfa v Low reliance on N fertilizer (legume) v High yield (6 t/ha) v Drought resistant (deep root system) v Amenable to genetic improvement
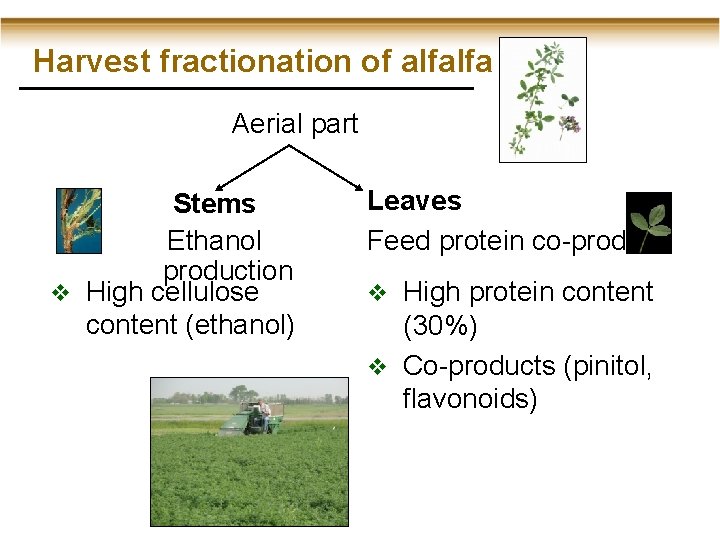
Harvest fractionation of alfalfa Aerial part Stems Ethanol production v High cellulose content (ethanol) Leaves Feed protein co-product v High protein content (30%) v Co-products (pinitol, flavonoids)

Bioethanol production from perennial alfalfa To increase the competitivity of alfalfa as feedstock for ethanol production, there is a need for the development of new genetic resources with: v High sugar content v High stem saccharification • Screening test using commercial enzymes v Value-added co-products

High throughput screening Enzymatic cocktail for stem degradation v Accellerase 1500 (Genencor) Cellulase and xylanase activity (XC and XY additives) v Pectinex 3 XL (Sigma) added for alfalfa Pectinase, cellulase et hemicellulase Near-infrared reflectance spectroscopy v Prediction of physicochemical parameters v v Minimal sample preparation High throughput screening Stem degradability = Enzyme-released glucose

Frequency Enzyme-released glucose (NIRS) D- D+ Enzyme-released glucose v v Duceppe, Bertrand et al. (2010) Efficient high throughput screening Rapidly characterizes CW degradability of 1200 genotypes Large genetic diversity for stem degradability Selection of 20 genotypes with high (D+) and 20 with low (D-) degradability

Intercrossing 20 D+ and 20 Dgenotypes • Three cycles of recurrent divergent selection in two genetic b • Conventional breeding (no GMO)
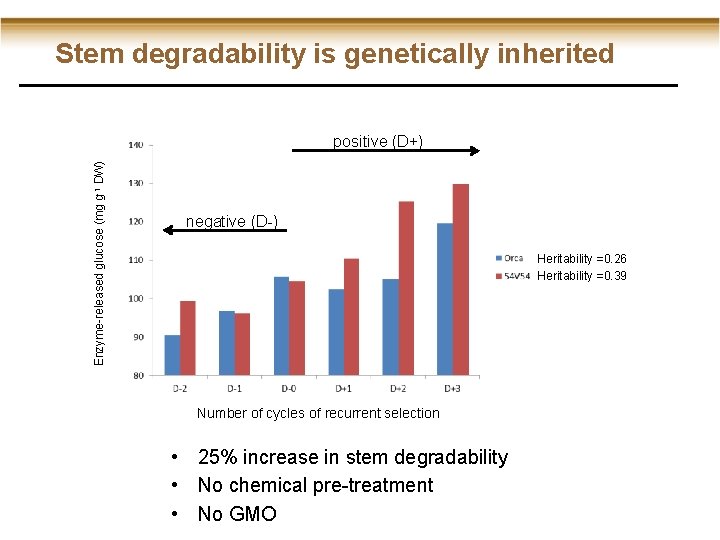
Stem degradability is genetically inherited Enzyme-released glucose (mg g-1 DW) positive (D+) negative (D-) Heritability =0. 26 Heritability =0. 39 Number of cycles of recurrent selection • 25% increase in stem degradability • No chemical pre-treatment • No GMO
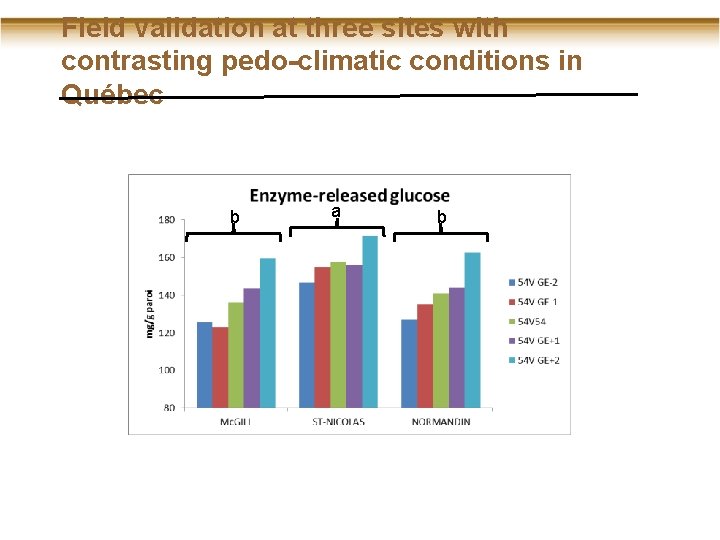
Field validation at three sites with contrasting pedo-climatic conditions in Québec b a b
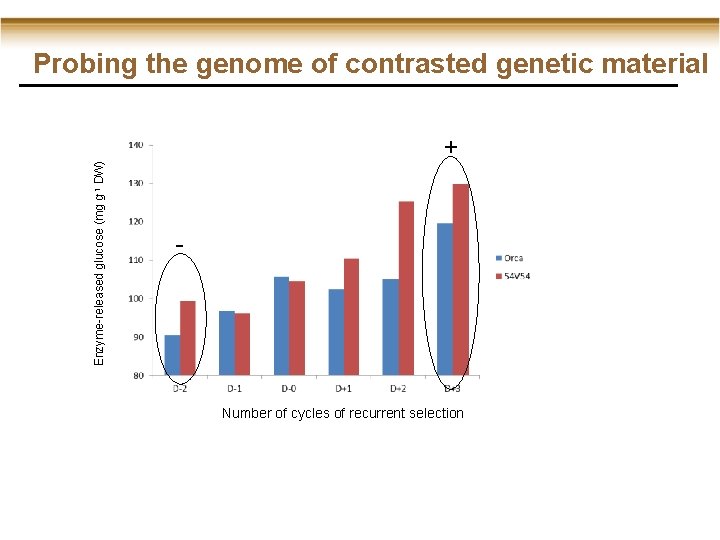
Probing the genome of contrasted genetic material Enzyme-released glucose (mg g-1 DW) + - Number of cycles of recurrent selection

Marker-Assisted Selection (MAS) for higher CW degradability • Use of the unique genetic material generated by recurrent divergent selection • Identification of genomic regions affecting stem degradability • Development of molecular markers to accelerate the identification of highly degradable plants

Random genome amplification v Sequence-related amplified polymorphisms (SRAP) v PCR approach targeting coding sequences v v Base on two primer pairs (17 -18 nucleotides) v One rich in GC content (exons) v One rich in AATT content (introns) Uncover numerous polymorphic regions over the entire genome

Polymorphism analyses v Populations v v v Pools v v v Biomass type : Orca Winter-hardy type: 54 V 54 Genotypes with high (D+) or low (D-) stem degradability Pool of DNA of 50 for each population PCR approach v 42 SRAP primer pairs
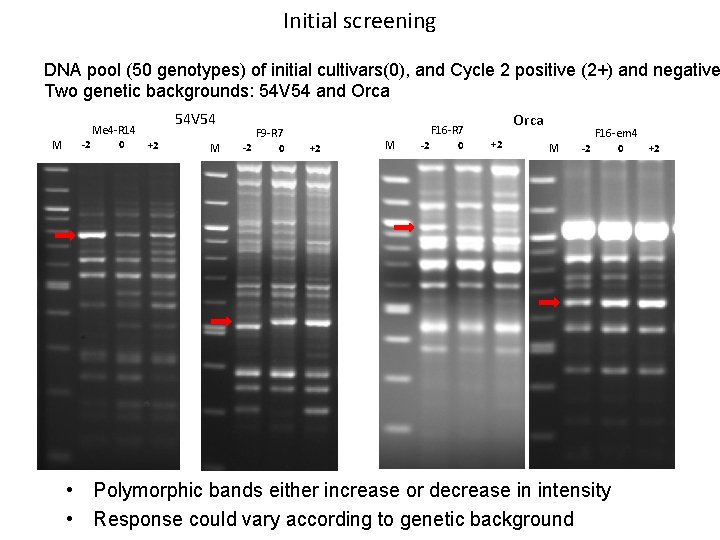
Initial screening DNA pool (50 genotypes) of initial cultivars(0), and Cycle 2 positive (2+) and negative Two genetic backgrounds: 54 V 54 and Orca 54 V 54 Me 4 -R 14 M -2 0 +2 M -2 0 Orca F 16 -R 7 F 9 -R 7 +2 M -2 0 +2 M F 16 -em 4 -2 • Polymorphic bands either increase or decrease in intensity • Response could vary according to genetic background 0 +2
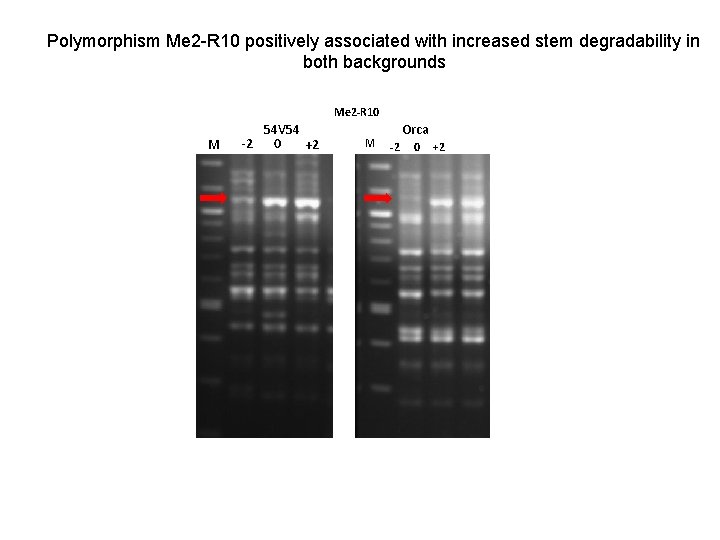
Polymorphism Me 2 -R 10 positively associated with increased stem degradability in both backgrounds Me 2 -R 10 M 54 V 54 -2 0 +2 M Orca -2 0 +2

SRAP primers combination: Me 2 -R 10 54 V 54 Cycle 2 M -2 0 0 +2 +2 N= 2 54 V 54 Cycle 2+ N= 10 Band intensity reflects the number of genotypes with polymorphisms in each population

Selection based on polymorphic markers • DNA extraction in 250 genotypes of each population. • Score of genotypes with six different polymorphic markers in 54 V 54 and Orca (presence or abscence) • Selection of 25 genotypes with more than two polymorphisms • Intercrossing selected genotypes • Proceeded to a second cycle in order to increase the frequency of the selected markers in new populations

Next steps • Stem degradability assessment of progenies after 2 cycles of MAS: to confirm the link between Markers and degradability Impact of MAS on stem degradability • Next generation sequencing (GBS): for genome-wide identification of regions affecting stem degradability

Scientists Annick Bertrand Annie Claessens Yves Castonguay Patrice Audy Research team Plant biochemistry Plant breeding Molecular physiology Molecular biology Post-doctorate Solen Rocher Molecular genetics Research assistants Josée Bourassa Sandra Delaney Jean Cloutier Josée Michaud Marie-Claude Pépin AAFC © 2012 Plant biochemistry Molecular genetics Molecular physiology Plant breeding
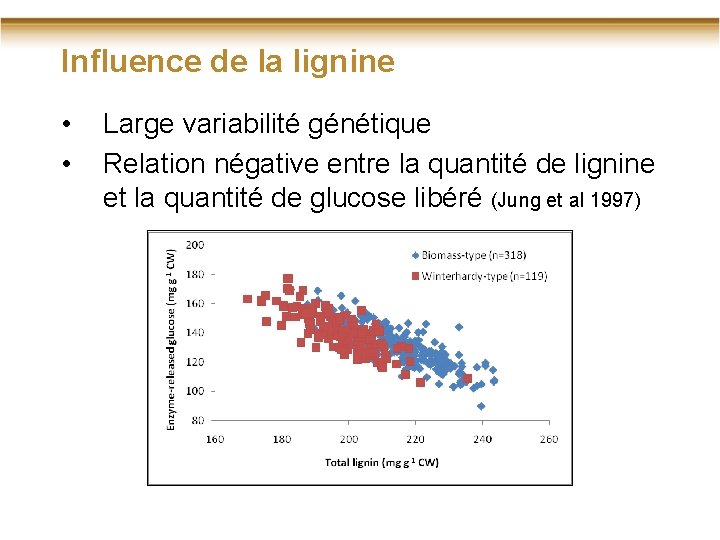
Influence de la lignine • • Large variabilité génétique Relation négative entre la quantité de lignine et la quantité de glucose libéré (Jung et al 1997)

Sélection pour une plus grande dégradabilité de la fibre basée sur les marqueurs moléculaires NOI 2652

Validation de différences dans la fréquence génotypique • Score 45 génotypes des populations 2+ et 2 dans chaque background • 10 marqueurs validés

Screening des deux popualtions • Extraction de l’ADN de 500 génotypes de chaque population • Score de 250 génotypes avec 5 marqueurs dans 54 V 54 et 6 marqueurs dans Orca • 25 génotypes sélectionnés pour les croisements

Prochaines étapes • Croisements polycross de 25 plants de Orca et de 54 V 54 (en cours) • Purification et clonage des séquences SRAP polymorphiques (hiver 2014) • Second cycle de sélection MAS dans les deux backgrounds (Été-automne 2014). • Croisements polycross du cycle 2 à l’hiver 2015.
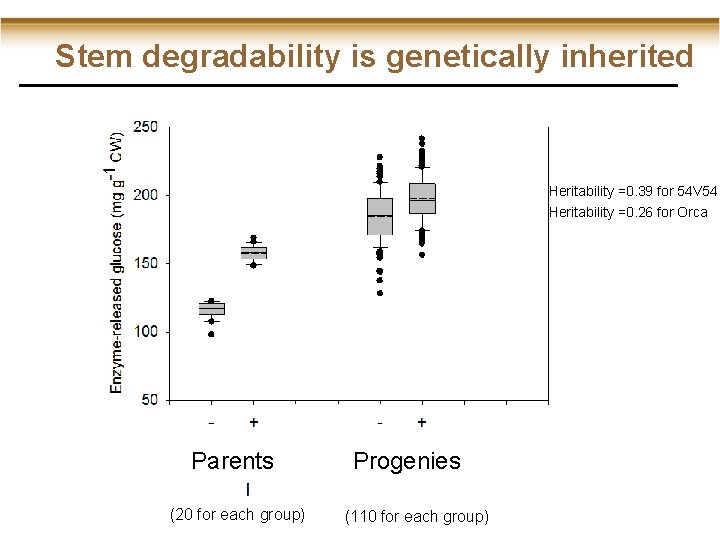
Stem degradability is genetically inherited Heritability =0. 39 for 54 V 54 Heritability =0. 26 for Orca Parents (20 for each group) Progenies (110 for each group)

Context v Over 2 millions ha of agricultural land in forage production in Québec v Knowledge, machinery, the infrastructure to cultivate harvest and store perennial forage crops v Bioenergy represent new opportunities for farmers, processors, and rural communities.
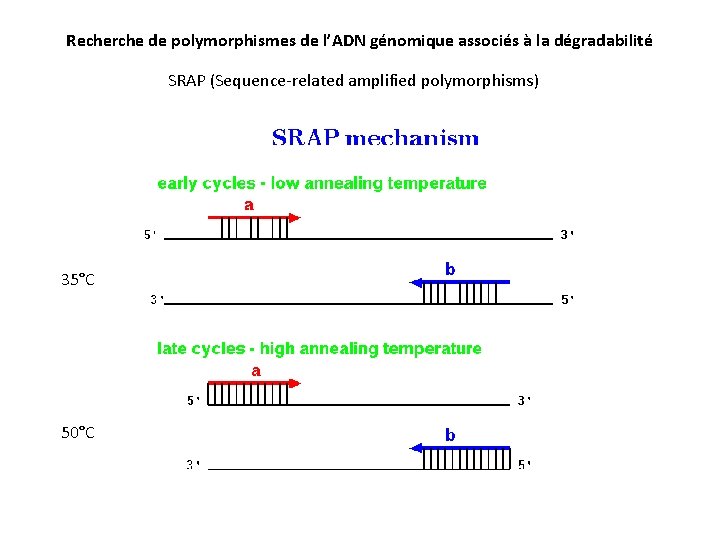
Recherche de polymorphismes de l’ADN génomique associés à la dégradabilité SRAP (Sequence-related amplified polymorphisms) 35°C 50°C
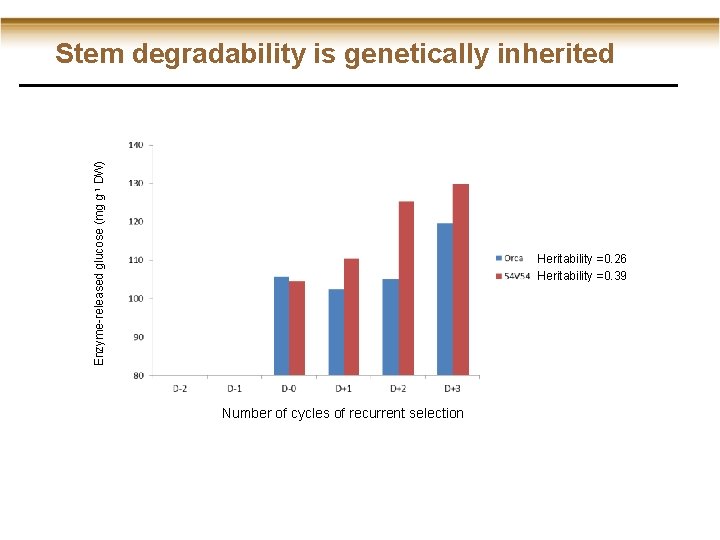
Enzyme-released glucose (mg g-1 DW) Stem degradability is genetically inherited Heritability =0. 26 Heritability =0. 39 Number of cycles of recurrent selection
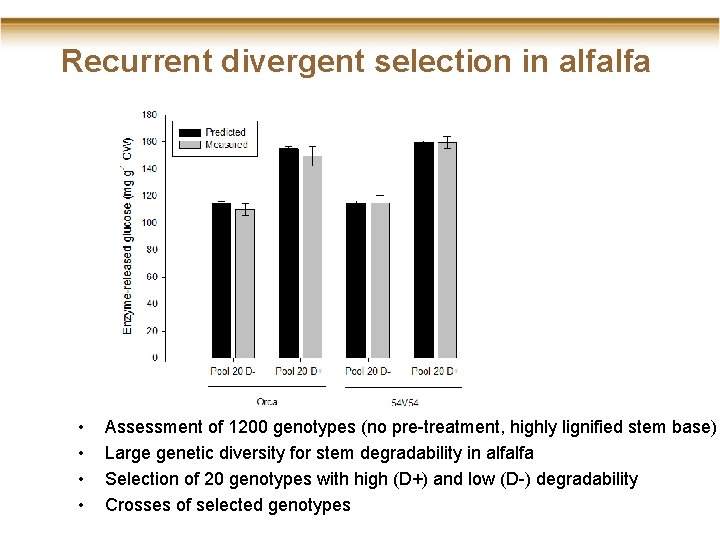
Recurrent divergent selection in alfalfa • • Assessment of 1200 genotypes (no pre-treatment, highly lignified stem base) Large genetic diversity for stem degradability in alfalfa Selection of 20 genotypes with high (D+) and low (D-) degradability Crosses of selected genotypes
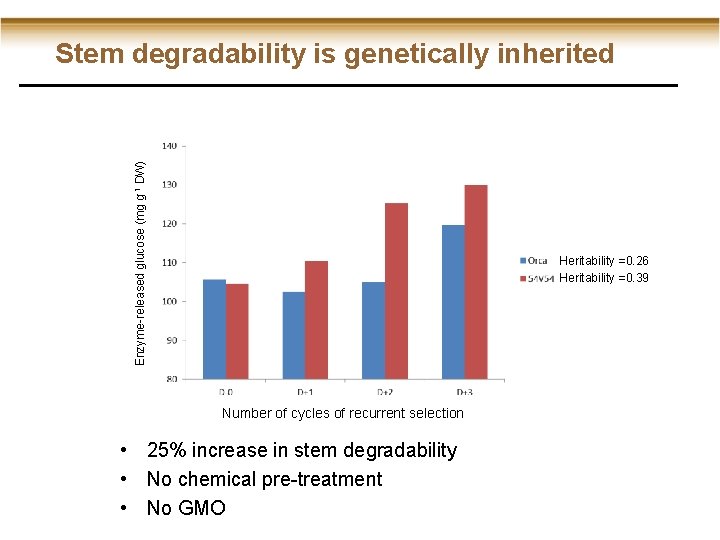
Enzyme-released glucose (mg g-1 DW) Stem degradability is genetically inherited Heritability =0. 26 Heritability =0. 39 Number of cycles of recurrent selection • 25% increase in stem degradability • No chemical pre-treatment • No GMO
- Slides: 31