Implications of partitioned genetic diversity for linkage disequilibrium

"Implications of partitioned genetic diversity for linkage disequilibrium mapping in elite UK cereal germplasm". Donal O’Sullivan SGC Meeting, JIC, 6 -7 th April 2006

Purpose To explore the prudent use of ‘populations’ of elite cereal varieties as LD mapping panels Why use ‘elite’ varieties? • Most familiar and obvious material • Reasonable levels of diversity present • Relevant to current markets • Obtainable in quantity • Extensive ‘historic series’ of robust field data for most relevant phenotypes

Partitioning in 500+ Gediflux Barley varieties

Association mapping in wheat: proof of principle • Gediflux data set: 499 genotyped varieties. 73 SSAPs, 42 SSRs, 72 NBS 1 B 1 R, pinb haplotypes • Historic trial data: 193 varieties with 18 phenotypes (incomplete ) yield +/- treated, hardness (113 lines) Lodging, disease, etc. • Use pinb as a candidate with known phenotypic effect • Use SSRs for structured association • Analyse using “Structure” and “Strat” • Use SSAPs for genomic control • Analyse trait by logistic regression
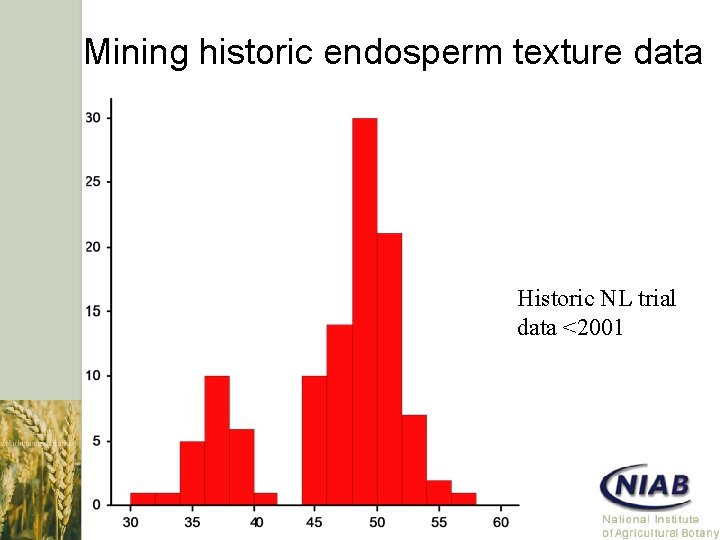
Mining historic endosperm texture data Historic NL trial data <2001

Structured association Structure: burn-in 1 million iterations 1 million No. populations (K) 8 No. replicate runs 2
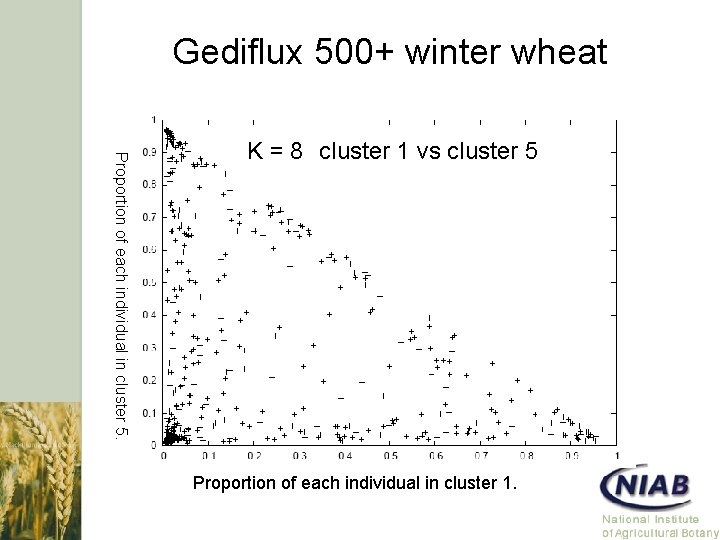
Gediflux 500+ winter wheat Proportion of each individual in cluster 5. K = 8 cluster 1 vs cluster 5 Proportion of each individual in cluster 1.
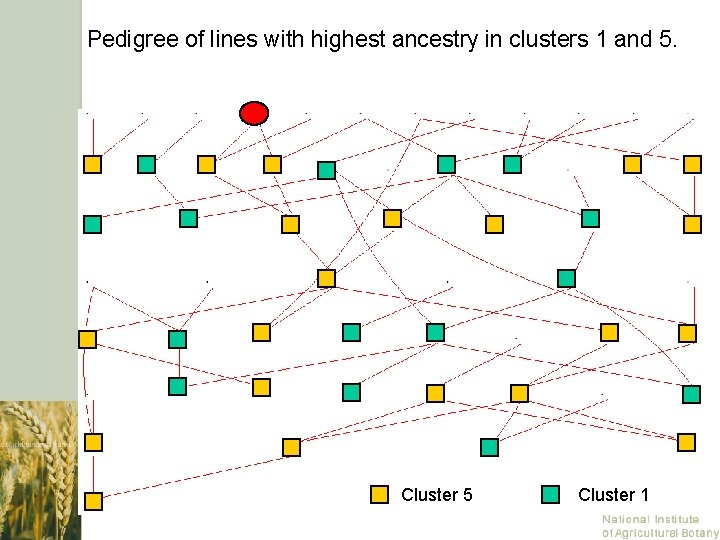
Pedigree of lines with highest ancestry in clusters 1 and 5. Cluster 5 Cluster 1
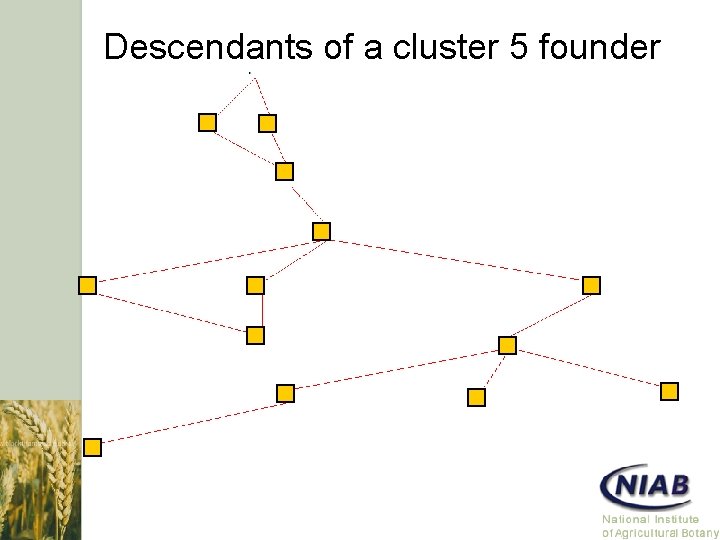
Descendants of a cluster 5 founder

Parentage of lines for clusters 1 and 5. parents progeny cluster 1 cluster 5 13 3 1 19 odds ratio p-value 82 0. 00001 Cluster membership is genetic!

Association test using STRAT / structure Pinb and hardness test chi sq p-value Assuming no structure in population 44. 44 0 Corrected (run a) 21. 89 0 Corrected (run b) 21. 42 0

STRAT and structure - QC Hardness and 55 SSAP markers, p-values <0. 05 Test Assuming no pop. structure No. <0. 05 14 Adjusted, run a 6 Adjusted, run b 6 Expected 3 May be under correcting.
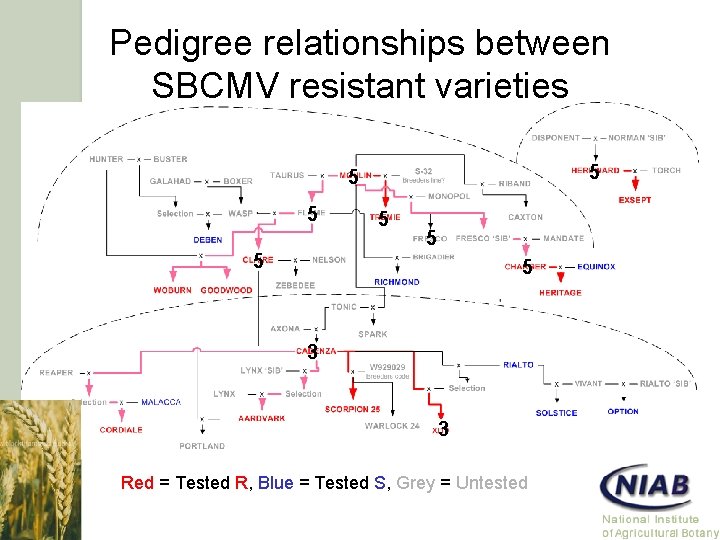
Pedigree relationships between SBCMV resistant varieties 5 5 5 5 3 3 Red = Tested R, Blue = Tested S, Grey = Untested

Genomic Control • Method: use multilocus genotype data to detect and correct for stratification • Premise: admixture operates over the whole genome but LD operates locally at short scales • 18 traits • 58 SSAP • 1044 logistic regression analyses
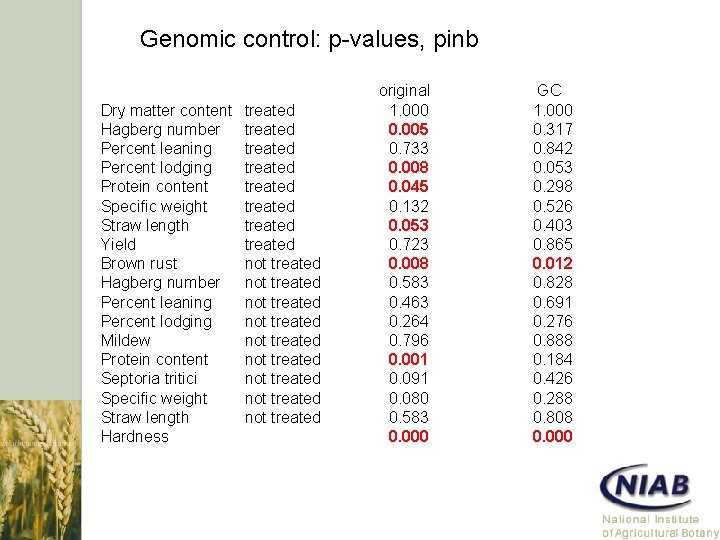
Genomic control: p-values, pinb Dry matter content Hagberg number Percent leaning Percent lodging Protein content Specific weight Straw length Yield Brown rust Hagberg number Percent leaning Percent lodging Mildew Protein content Septoria tritici Specific weight Straw length Hardness treated treated not treated not treated not treated original 1. 000 0. 005 0. 733 0. 008 0. 045 0. 132 0. 053 0. 723 0. 008 0. 583 0. 463 0. 264 0. 796 0. 001 0. 091 0. 080 0. 583 0. 000 GC 1. 000 0. 317 0. 842 0. 053 0. 298 0. 526 0. 403 0. 865 0. 012 0. 828 0. 691 0. 276 0. 888 0. 184 0. 426 0. 288 0. 808 0. 000

Genomic control: QC Test markers across all traits No. of tests 1044 P-value <0. 05 OBS original 313 OBS GC 34 EXP 52 May be overcorrecting

Conclusions • Population structure may be evident e. g. spring-winter/row number divide or less so – Carry out LD mapping within major sub-groups • UK winter wheat shows cryptic population structure which groups varieties consistent with known pedigree • Genomic control and/or structured association both effective in detecting known associations and reducing false +ves to realistic levels • Roll on new phenotype and genotype data!

Acknowledgements • NIAB – – Fiona Leigh John Law Ian Mackay Wayne Powell • Gediflux Partners – – – Barley Marion Roeder (IPK) Wheat Rob Koebner, Simon Orford (JIC) Martin Ganal (Trait Genetics)
- Slides: 18