Immunology Immunodeficiencies AIDS Vaccine Enzymelinked immunosorbent assays ELISA

Immunology Immunodeficiencies (AIDS) Vaccine Enzyme-linked immunosorbent assays (ELISA) Hypersenstivity Transplantation Dr. P. MURUGAN, M. Sc. , M. Phil. , Ph. D. , Pricipal i/c Department of Biochemistry Bharathidasan University Model. College, Vedharanyam.

Immunodeficiencies (AIDS)
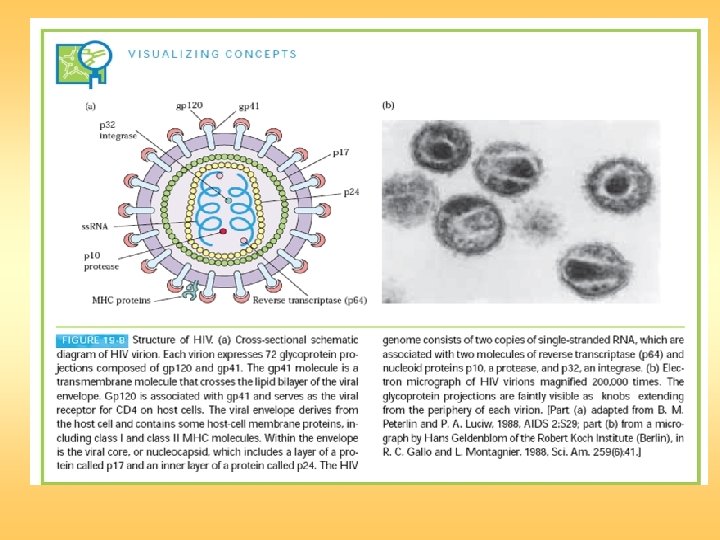
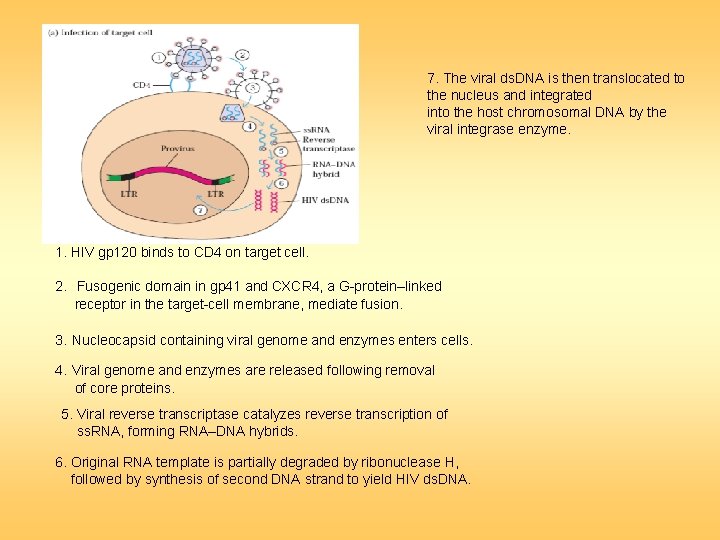
7. The viral ds. DNA is then translocated to the nucleus and integrated into the host chromosomal DNA by the viral integrase enzyme. 1. HIV gp 120 binds to CD 4 on target cell. 2. Fusogenic domain in gp 41 and CXCR 4, a G-protein–linked receptor in the target-cell membrane, mediate fusion. 3. Nucleocapsid containing viral genome and enzymes enters cells. 4. Viral genome and enzymes are released following removal of core proteins. 5. Viral reverse transcriptase catalyzes reverse transcription of ss. RNA, forming RNA–DNA hybrids. 6. Original RNA template is partially degraded by ribonuclease H, followed by synthesis of second DNA strand to yield HIV ds. DNA.

Vaccine
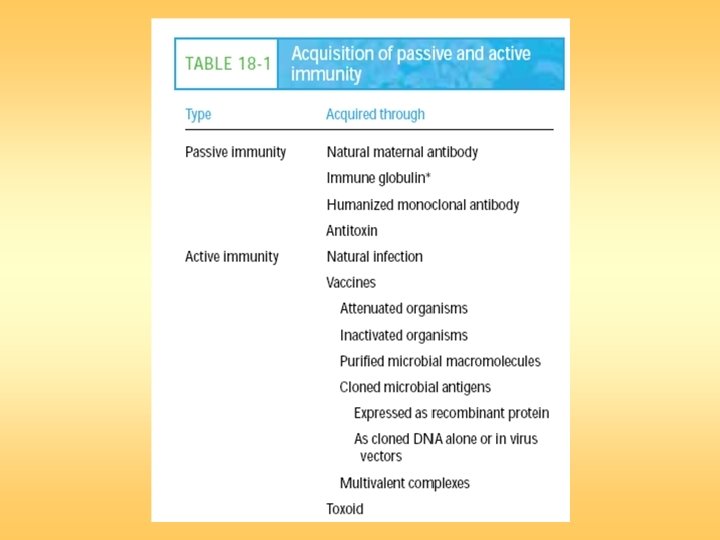
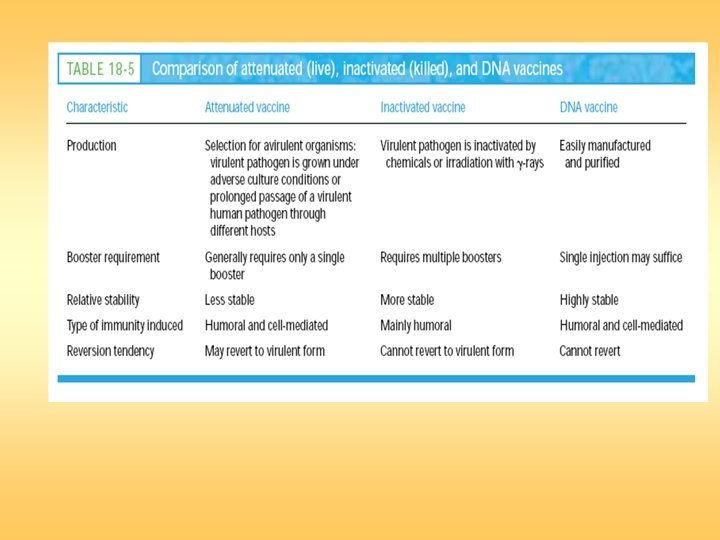
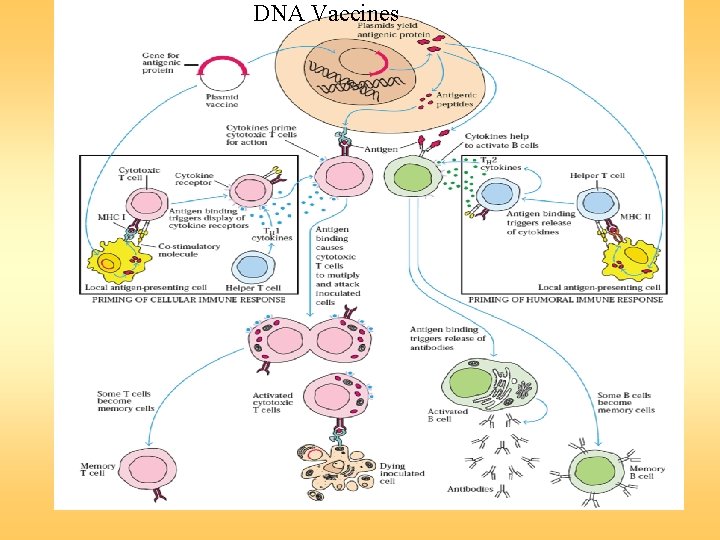
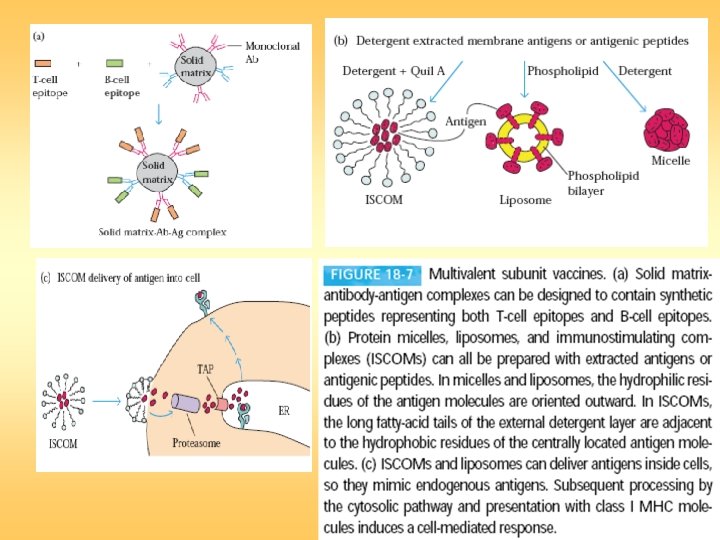
DNA Vaccines
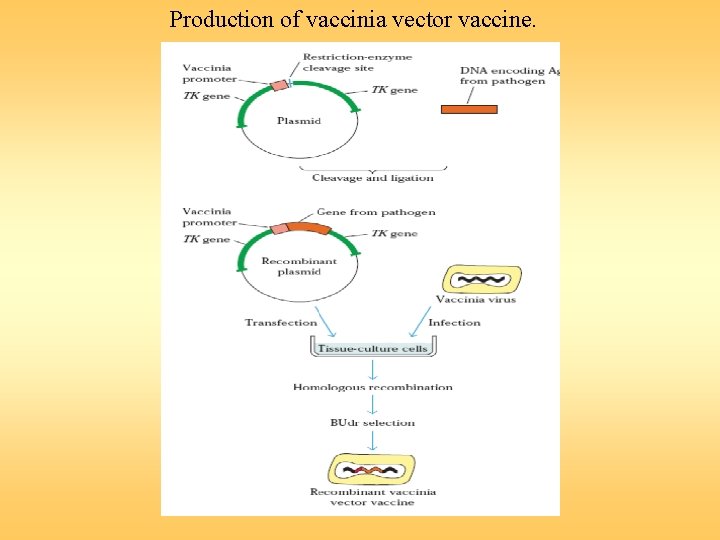
Production of vaccinia vector vaccine.

ELISA zyme-linked immunosorbent assays (ELISA)
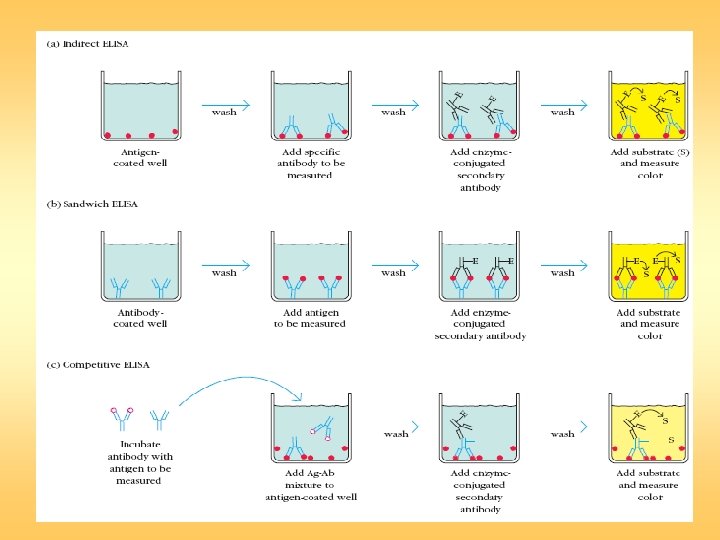
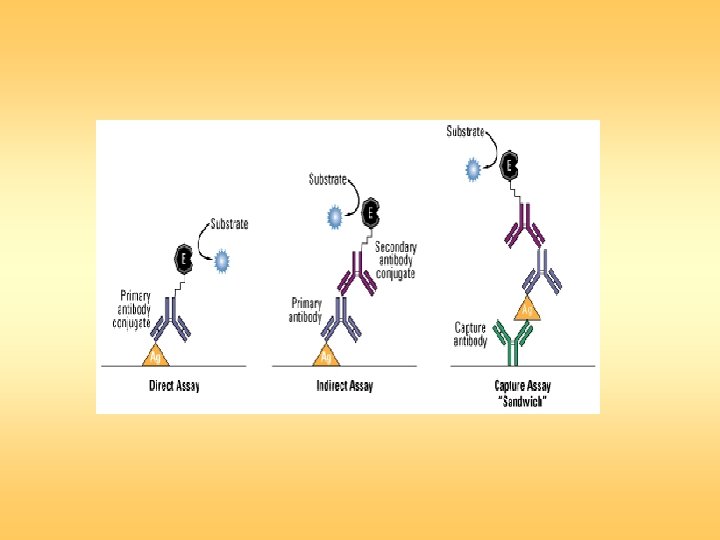
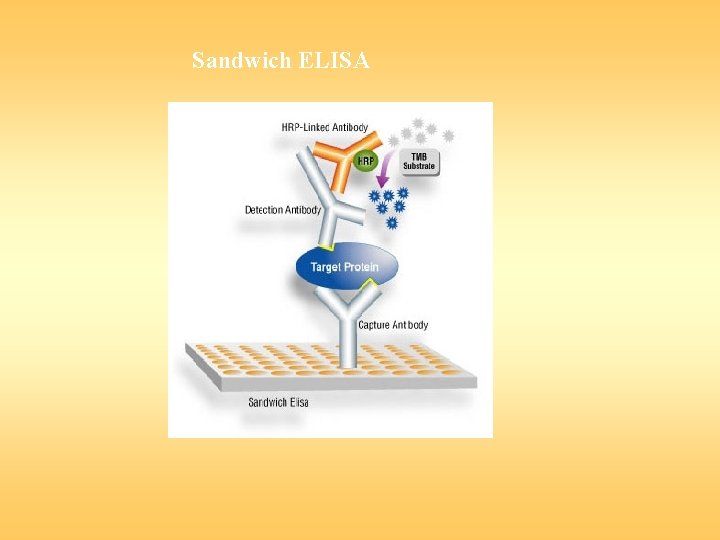
Sandwich ELISA

Fluorescent in situ hybridization

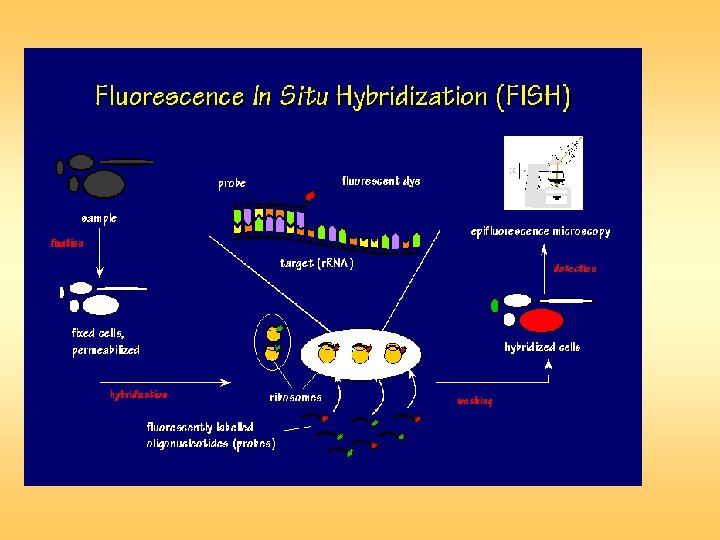
Fluorescent in situ hybridization (FISH) is a powerful technique for detecting RNA or DNA sequences in cells, tissues, and tumors. FISH provides a unique link among the studies of cell biology, cytogenetics, and molecular genetics. In Situ Hybridization (ISH) uses a labeled probe to detect and localize specific RNA or DNA sequences in a tissue or on a chromosome. Fluorescent" means emitting light that comes from a reaction within the emitter. "In situ" refers to the fact that this techniques is done with the chromosomes, cells or tissue in place (in situ) on a microscope slide.

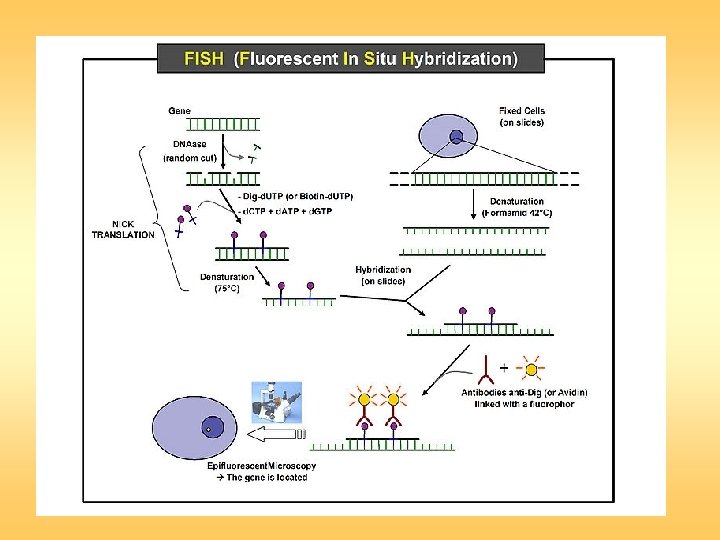
Fluorescent in situ hybridization is a technique in which singlestranded nucleic acids (usually DNA, but RNA may also be used) are permitted to interact so that complexes, or hybrids, are formed by molecules with sufficiently similar, complementary sequences. The method comprises of three basic steps: fixation of a specimen on a microscope slide, hybridization of labeled probe to homologous fragments of genomic DNA, and enzymatic detection of the tagged target hybrids. While probe sequences were initially detected with isotopic reagents, nonisotopic hybridization has become increasingly popular, with fluorescent hybridization now a common choice.

The detection of sequences on the target chromosomes is performed indirectly, commonly with biotinylated or digoxigenin-labeled probes detected via a fluorochromeconjugated detection reagent, such as an antibody conjugated with fluorescein. As a result, the direct visualization of the relative position of the probes is possible. FISH is a very general technique. The differences between the various FISH techniques are usually due to variations in the sequence and labeling of the probes; and how they are used in combination. These few modifications make possible all FISH techniques. Probe size is important because longer probes hybridize more specifically than shorter probes.

Medical applications In medicine, FISH can be used to form a diagnosis, to evaluate prognosis, or to evaluate remission of a disease, such as cancer. FISH can also be used to detect diseased cells more easily than standard Cytogenetic methods. FISH, does not require living cells and can be quantified automatically, a computer counts the fluorescent dots present. FISH can be used to detect directly the presence of the microbial pathogens on small samples of patient's tissue. FISH can also be used to compare the genomes of two biological species, to deduce evolutionary relationships.

Hypersensitive Reactions
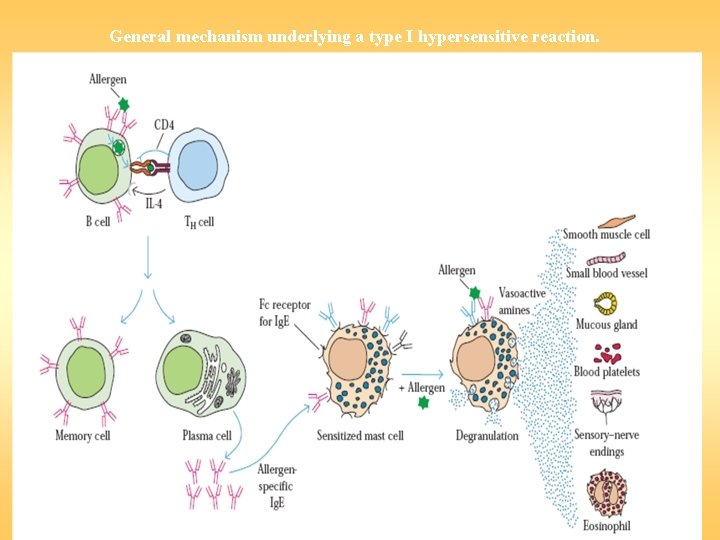
General mechanism underlying a type I hypersensitive reaction.
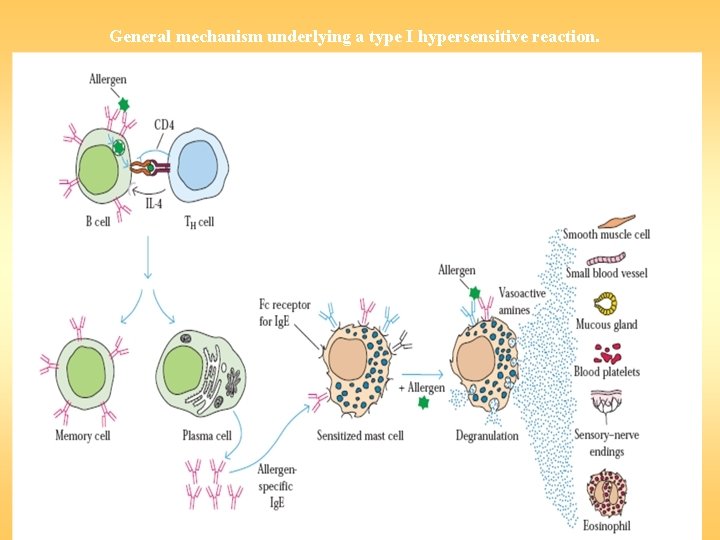
General mechanism underlying a type I hypersensitive reaction.
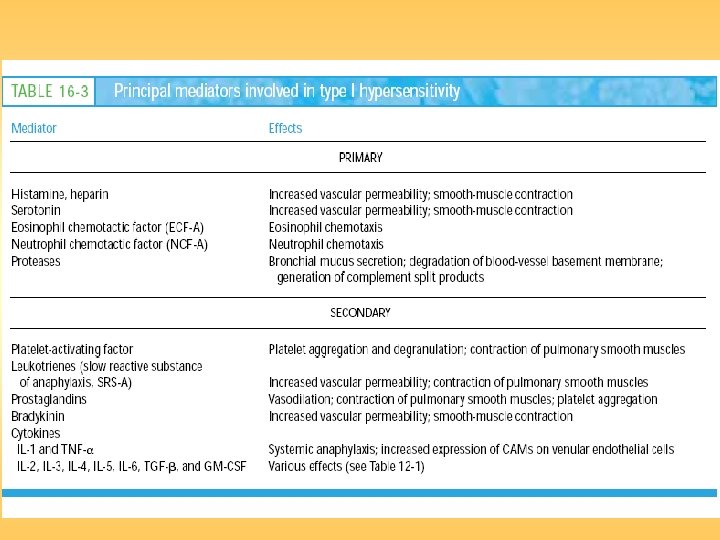
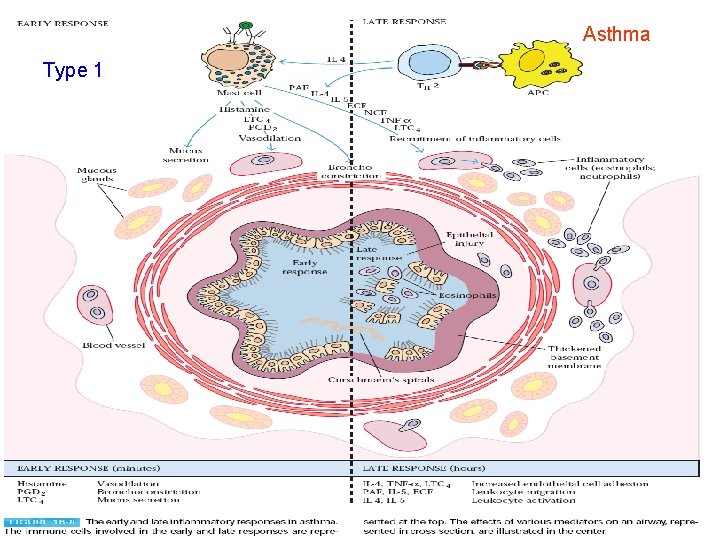
Asthma Type 1
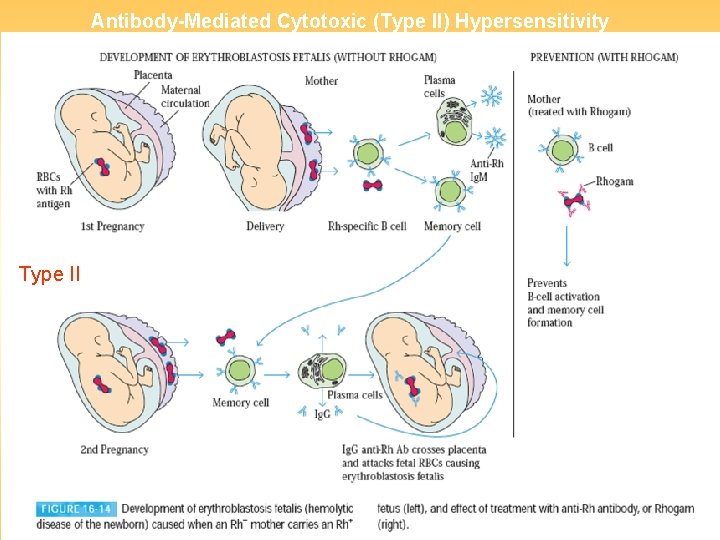
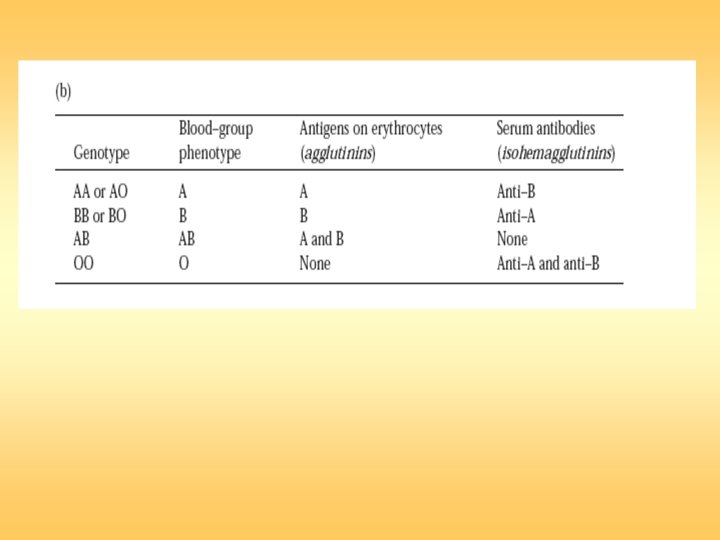
Antibody-Mediated Cytotoxic (Type II) Hypersensitivity Type II
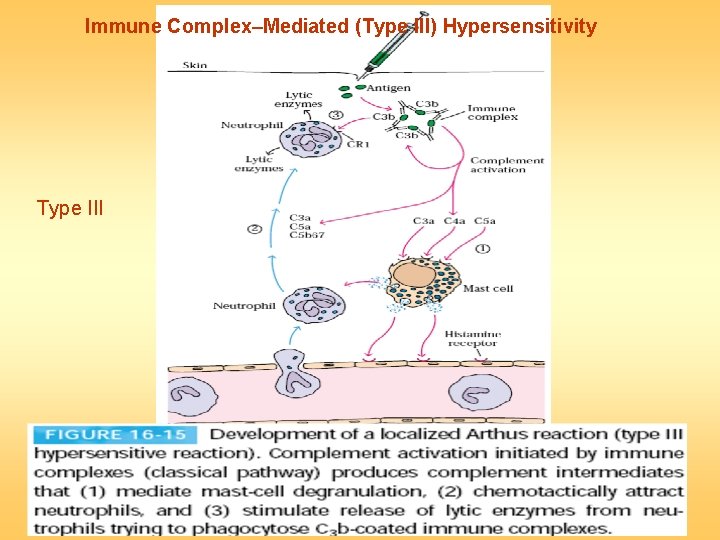
Immune Complex–Mediated (Type III) Hypersensitivity Type III
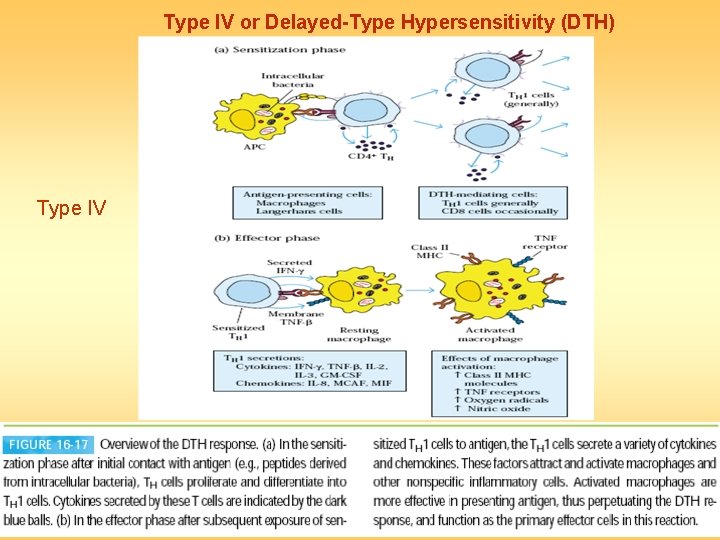
Type IV or Delayed-Type Hypersensitivity (DTH) Type IV
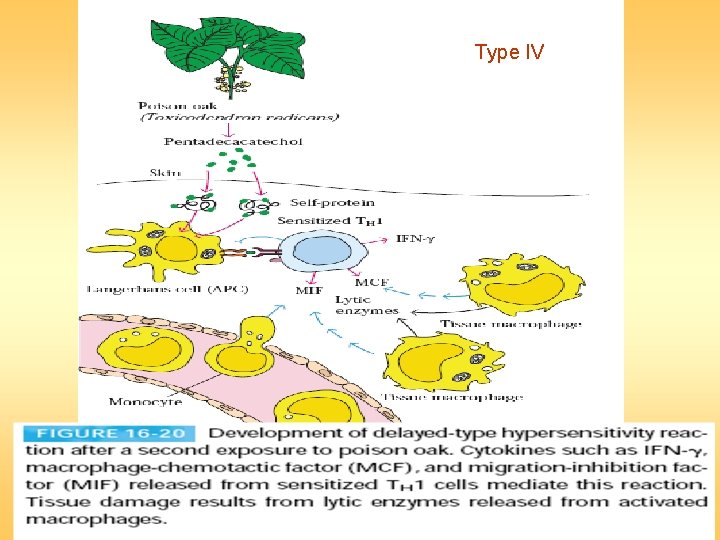
Type IV

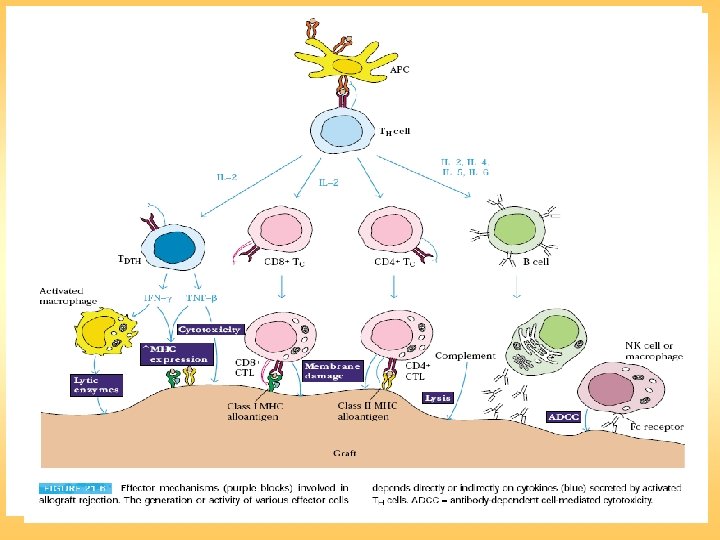
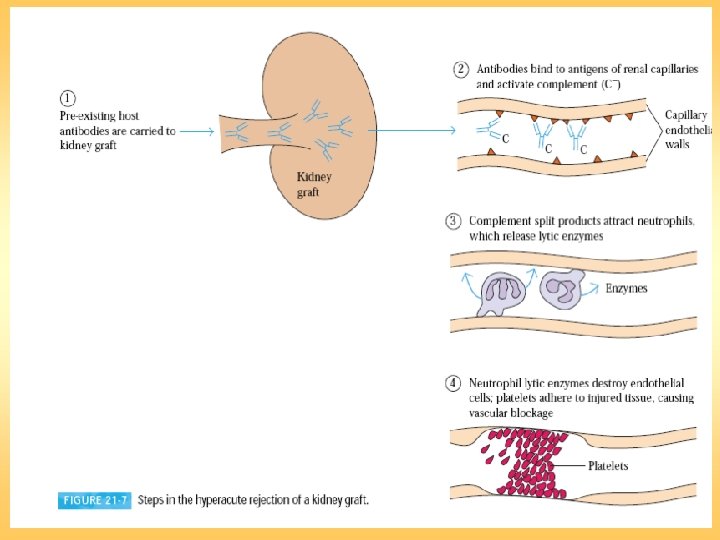
Transplantation
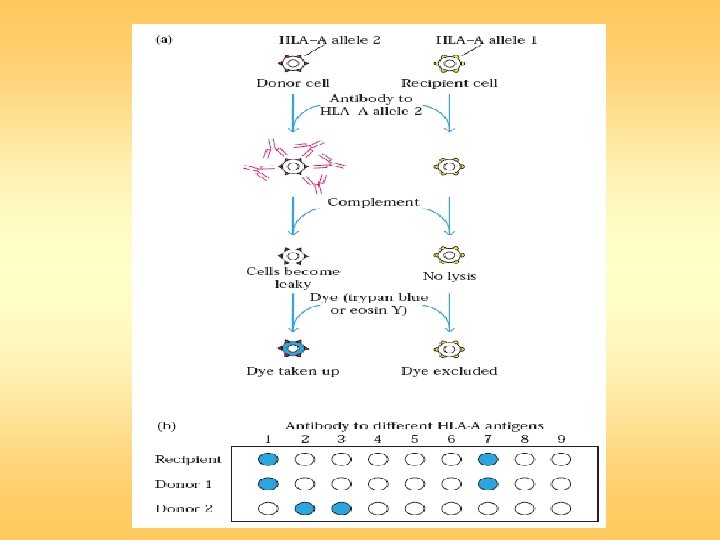
- Slides: 37