IGFBP 7 and Its Fuctions IGFBP 7 55

IGFBP 7 and Its Fuctions IGFBP 7的 抗肿瘤效应 浙江大学 德等 来茂
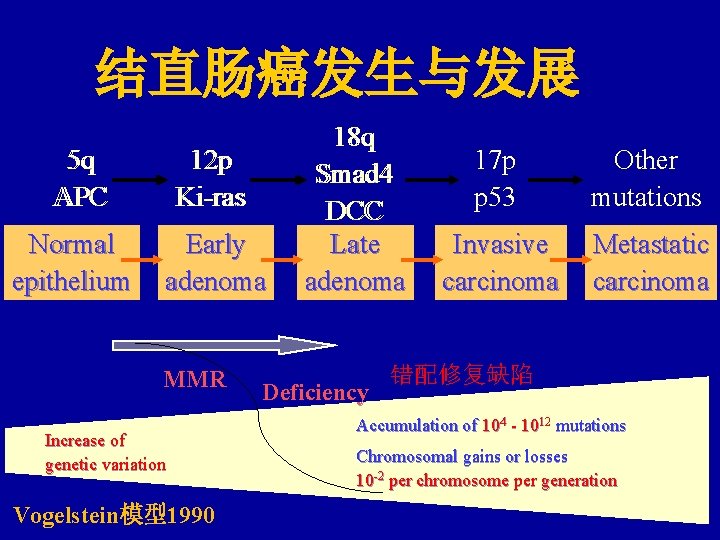
结直肠癌发生与发展 55 qq APC Normal epithelium 12 12 pp Ki-ras Early adenoma MMR Increase of genetic variation Vogelstein模型1990 18 18 qq Smad 4 DCC Late adenoma Deficiency 17 p p 53 Other mutations Invasive carcinoma Metastatic carcinoma 错配修复缺陷 Accumulation of 104 - 1012 mutations Chromosomal gains or losses 10 -2 per chromosome per generation
In 1999, by SSH, We constructed three libraries normal adenoma APC, etc. N cancer P 53, etc. A T
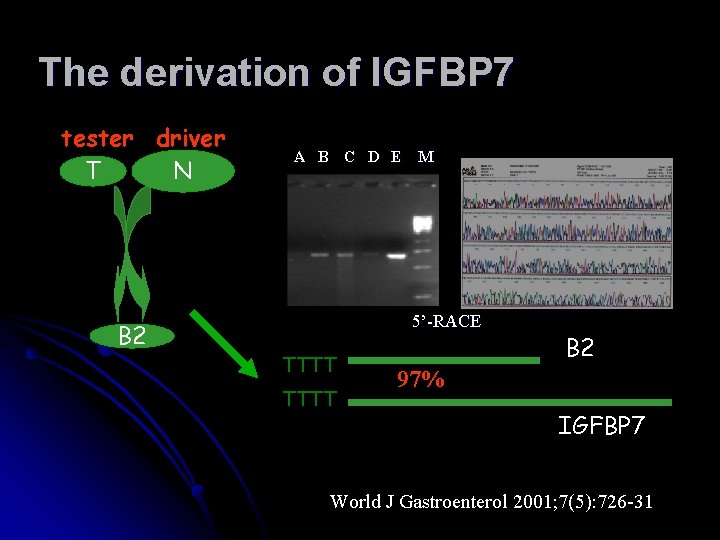
The derivation of IGFBP 7 tester driver N T B 2 A B C D E M 5’-RACE TTTT B 2 97% IGFBP 7 World J Gastroenterol 2001; 7(5): 726 -31
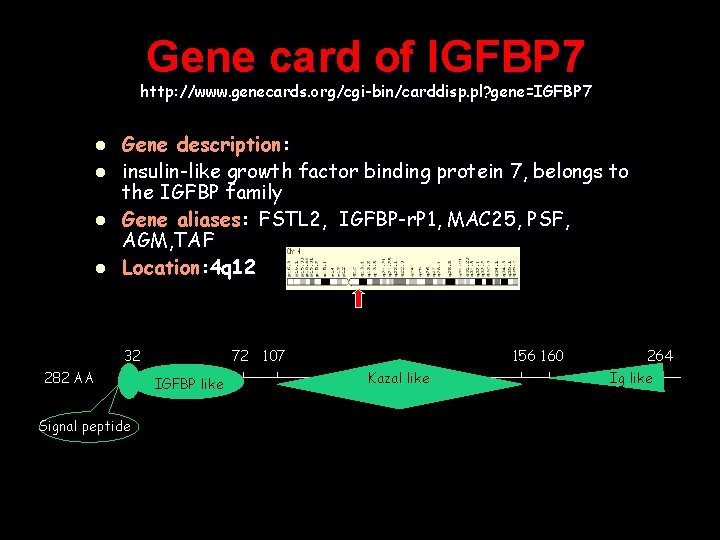
Gene card of IGFBP 7 http: //www. genecards. org/cgi-bin/carddisp. pl? gene=IGFBP 7 l l Gene description: insulin-like growth factor binding protein 7, belongs to the IGFBP family Gene aliases: FSTL 2, IGFBP-r. P 1, MAC 25, PSF, AGM, TAF Location: 4 q 12 32 282 AA Signal peptide 156 160 72 107 IGFBP like Kazal like 264 Ig like

I: IGFBP 7 in colon cancer tissue IGFBP 7在癌组织中的表达
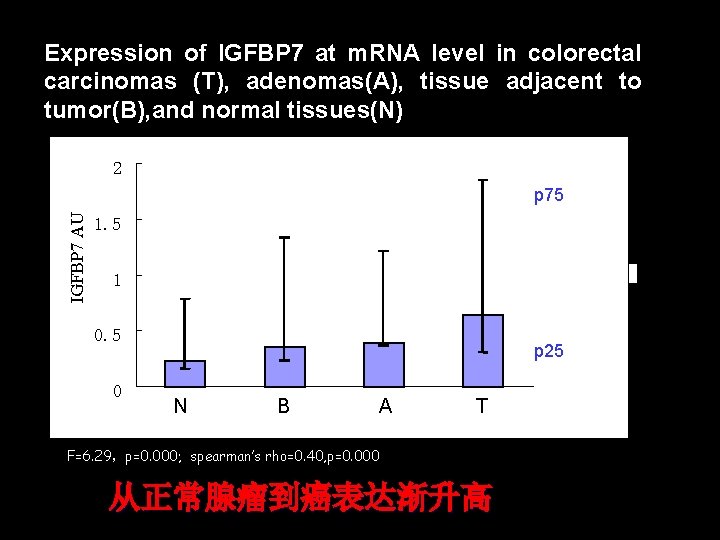
Expression of IGFBP 7 at m. RNA level in colorectal carcinomas (T), adenomas(A), tissue adjacent to tumor(B), and normal tissues(N) 2 IGFBP 7 AU p 75 1. 5 P 25 1 0. 5 N 0 N B B A A T T F=6. 29,p=0. 000; spearman’s rho=0. 40, p=0. 000 从正常腺瘤到癌表达渐升高 p 25
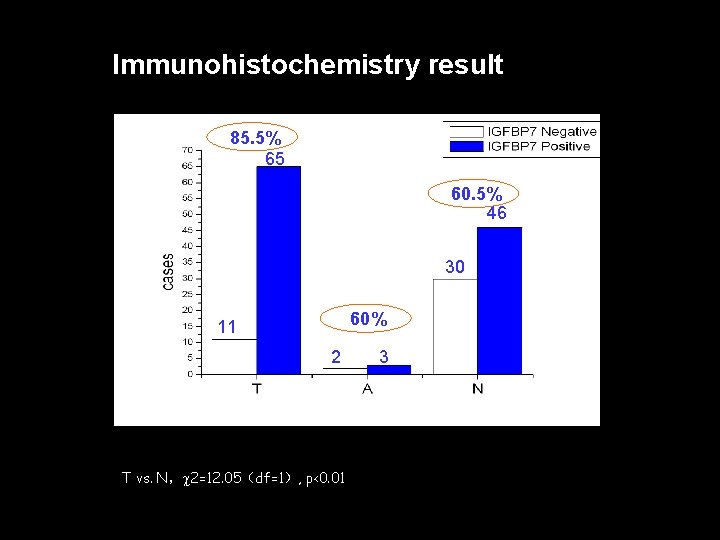
Immunohistochemistry result 85. 5% 65 60. 5% 46 30 60% 11 2 T vs. N, 2=12. 05(df=1), p<0. 01 3
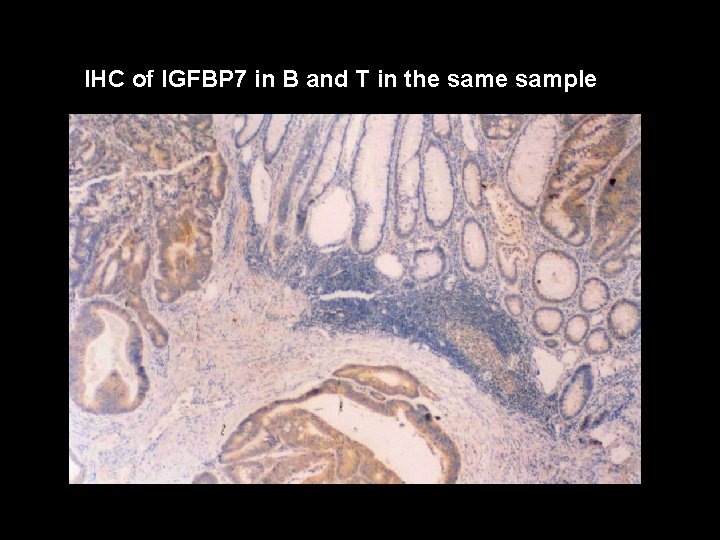
IHC of IGFBP 7 in B and T in the sample
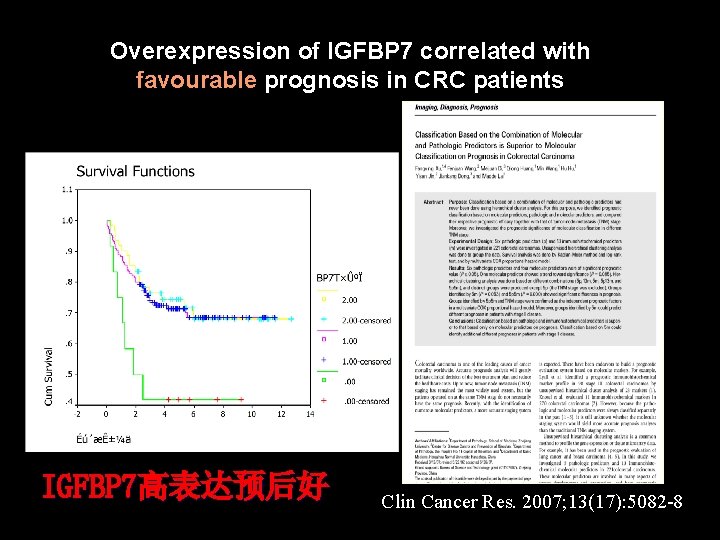
Overexpression of IGFBP 7 correlated with favourable prognosis in CRC patients IGFBP 7高表达预后好 Clin Cancer Res. 2007; 13(17): 5082 -8

IGFBP 7 has a low expression in CRC cell lines Lo. Vo HCT 8 SW 1116 RKO HT 29 COLO 205 Hce 8693 CW 2 SW 620 SW 480 Caco 2 Control Marker GAPDH IGFBP 7

II: The regulation mechanism of IGFBP 7 in colon cancer
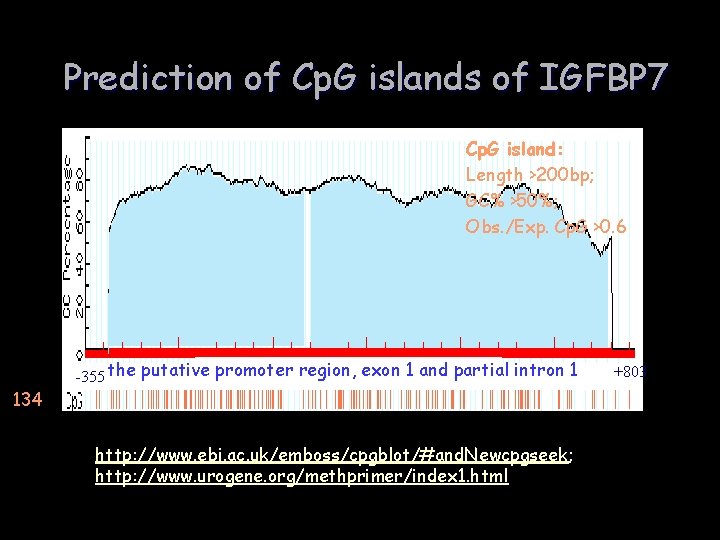
Prediction of Cp. G islands of IGFBP 7 Cp. G island: Length >200 bp; GC% >50%; Obs. /Exp. Cp. G >0. 6 -355 the putative promoter region, exon 1 and partial intron 1 134 http: //www. ebi. ac. uk/emboss/cpgblot/#and. Newcpgseek; http: //www. urogene. org/methprimer/index 1. html +803
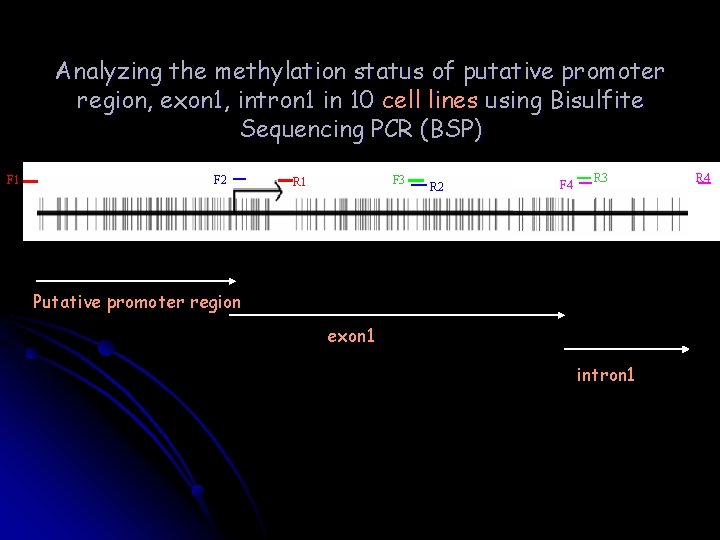
Analyzing the methylation status of putative promoter region, exon 1, intron 1 in 10 cell lines using Bisulfite Sequencing PCR (BSP) F 1 F 2 F 3 R 1 R 2 F 4 R 3 Putative promoter region exon 1 intron 1 R 4
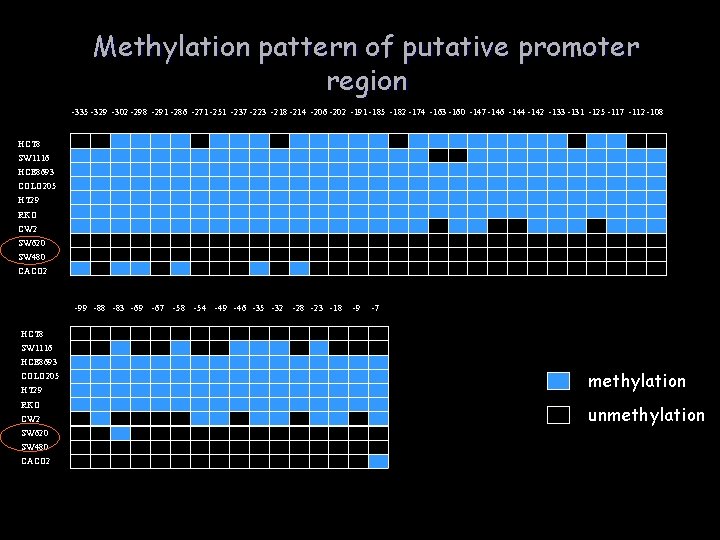
Methylation pattern of putative promoter region -335 -329 -302 -298 -291 -286 -271 -251 -237 -223 -218 -214 -206 -202 -191 -185 -182 -174 -163 -160 -147 -146 -144 -142 -133 -131 -125 -117 -112 -108 HCT 8 SW 1116 HCE 8693 COLO 205 HT 29 RKO CW 2 SW 620 SW 480 CACO 2 -99 -88 -83 -69 -67 -58 -54 -49 -46 -35 -32 -28 -23 -18 -9 -7 HCT 8 SW 1116 HCE 8693 COLO 205 HT 29 RKO CW 2 SW 620 SW 480 CACO 2 methylation unmethylation
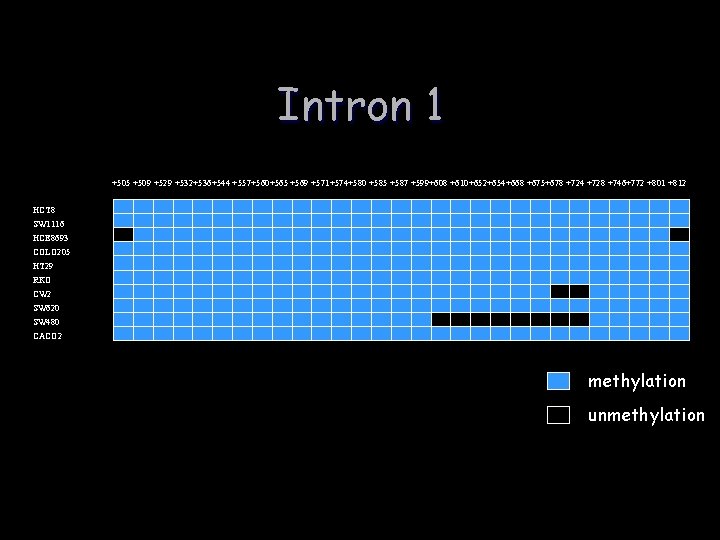
Intron 1 +505 +509 +529 +532+536+544 +557+560+565 +569 +571+574+580 +585 +587 +599+608 +610+652+654+668 +675+678 +724 +728 +746+772 +801 +812 HCT 8 SW 1116 HCE 8693 COLO 205 HT 29 RKO CW 2 SW 620 SW 480 CACO 2 methylation unmethylation
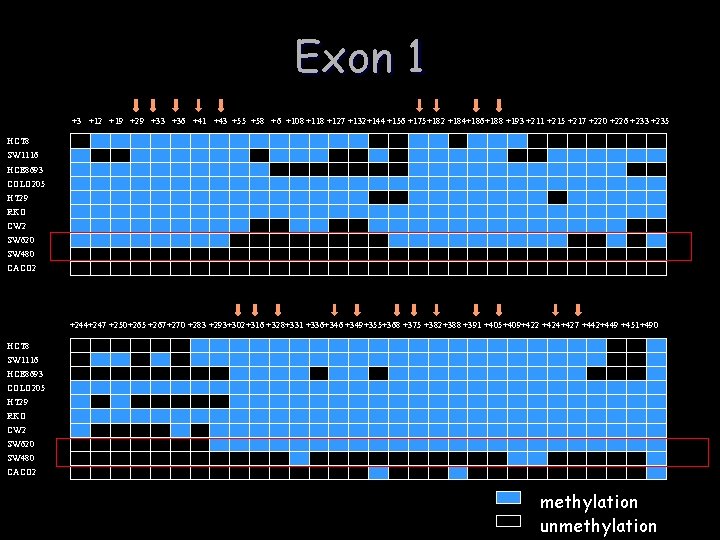
Exon 1 +3 +12 +19 +29 +33 +36 +41 +43 +55 +58 +6 +108 +118 +127 +132+144 +156 +175+182 +184+186+188 +193 +211 +215 +217 +220 +226 +233 +235 HCT 8 SW 1116 HCE 8693 COLO 205 HT 29 RKO CW 2 SW 620 SW 480 CACO 2 +244+247 +250+265 +267+270 +283 +293+302+316 +328+331 +336+346 +349+355+368 +375 +382+388 +391 +405+409+422 +424+427 +442+449 +451+490 HCT 8 SW 1116 HCE 8693 COLO 205 HT 29 RKO CW 2 SW 620 SW 480 CACO 2 methylation unmethylation

Confirmation using MSP CW 2 Hce 8693 SW 1116 M U M U HCT 8 HT 29 RKO COLO 205 SW 620 M U M U M U IGFBP 7(+ SW 480 ) Caco 2 M U Control M U unmethylation M, methylated product; U, unmethylated product; M-P, positive control of methylation; U-P, positive control of unmethylation; C, blank control of PCR system Marker

Cluster analysis depending on the methylation status of exon 1 IGFBP 7+
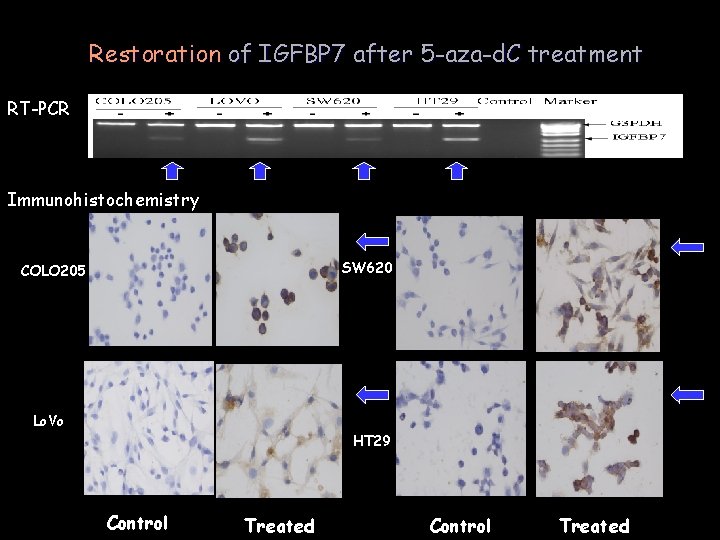
Restoration of IGFBP 7 after 5 -aza-d. C treatment RT-PCR Immunohistochemistry SW 620 COLO 205 Lo. Vo HT 29 Control Treated

Methy-and de. Methy sites of IGFBP-r. P 1 gene 5’-region -,control; treated cells ;Black : methy-per centage Methylation of+,IGFBP 7 exon 1: the key regulation mechanism silencing expression in CRC cells J Pathol. IGFBP 7 2007; 212(1): 83 -90. 外显子 1甲基化抑制基因表达

Aberrant methylation of IGFBP 7 exon 1 in colorectal cancer tissues 在结直肠癌中的异常表达
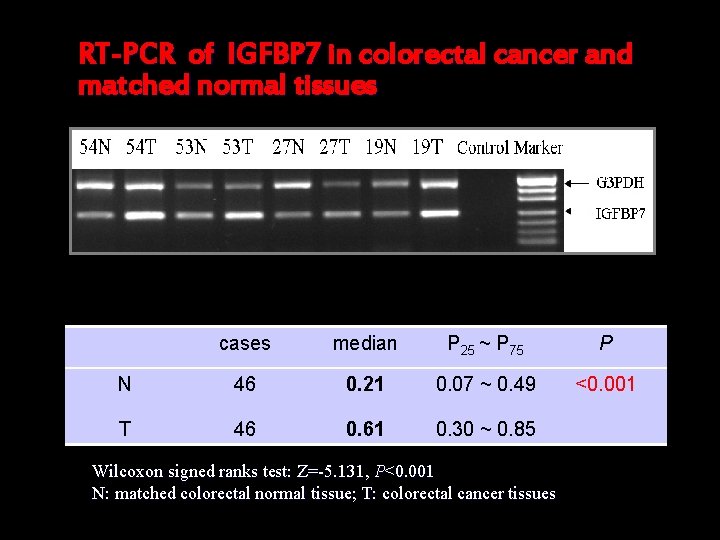
RT-PCR of IGFBP 7 in colorectal cancer and matched normal tissues cases median P 25 ~ P 75 P N 46 0. 21 0. 07 ~ 0. 49 <0. 001 T 46 0. 61 0. 30 ~ 0. 85 Wilcoxon signed ranks test: Z=-5. 131, P<0. 001 N: matched colorectal normal tissue; T: colorectal cancer tissues
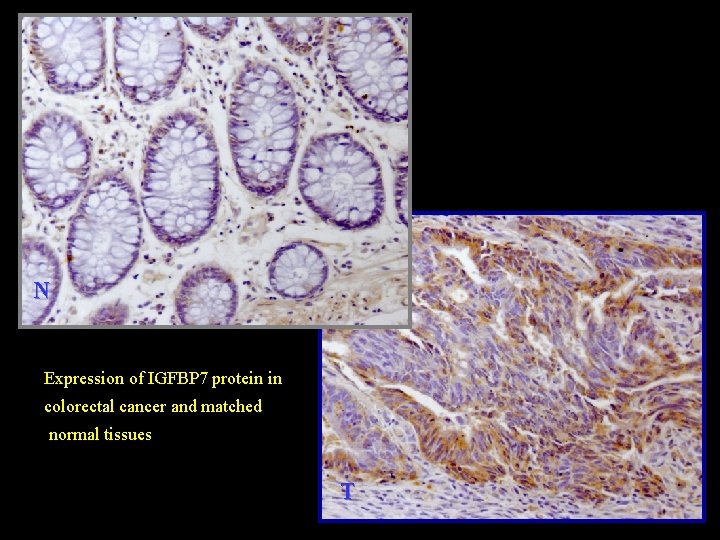
N Expression of IGFBP 7 protein in colorectal cancer and matched normal tissues T
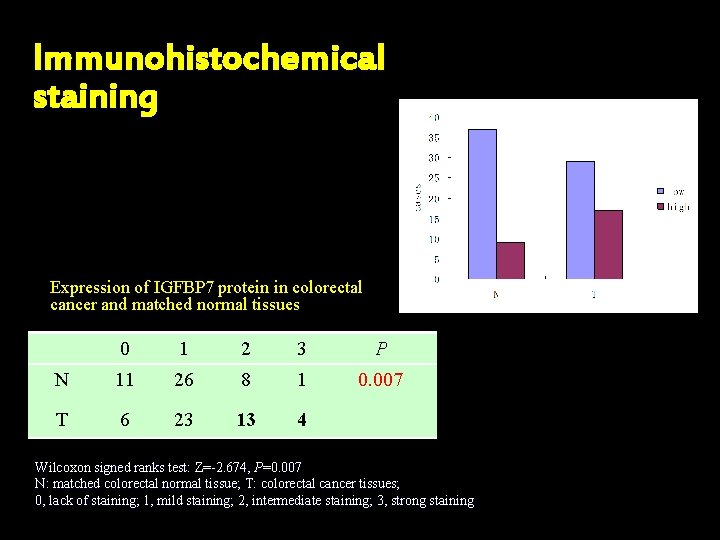
Immunohistochemical staining Expression of IGFBP 7 protein in colorectal cancer and matched normal tissues 0 1 2 3 P N 11 26 8 1 0. 007 T 6 23 13 4 Wilcoxon signed ranks test: Z=-2. 674, P=0. 007 N: matched colorectal normal tissue; T: colorectal cancer tissues; 0, lack of staining; 1, mild staining; 2, intermediate staining; 3, strong staining
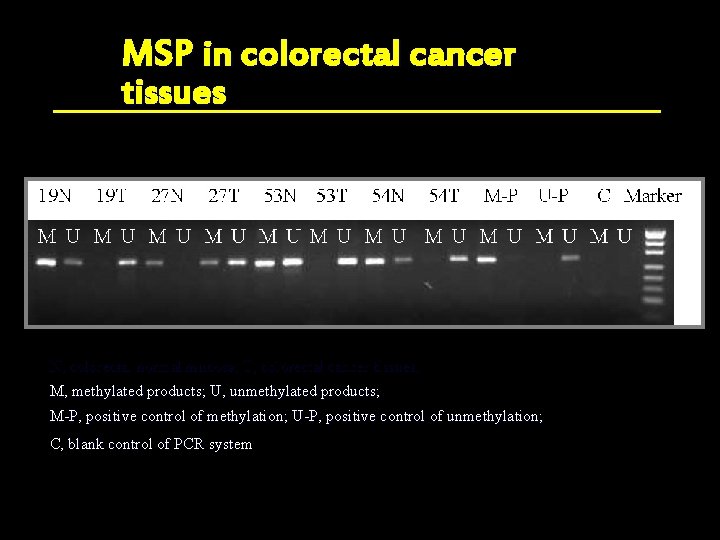
MSP in colorectal cancer tissues N, colorectal normal mucosa; T, colorectal cancer tissues; M, methylated products; U, unmethylated products; M-P, positive control of methylation; U-P, positive control of unmethylation; C, blank control of PCR system
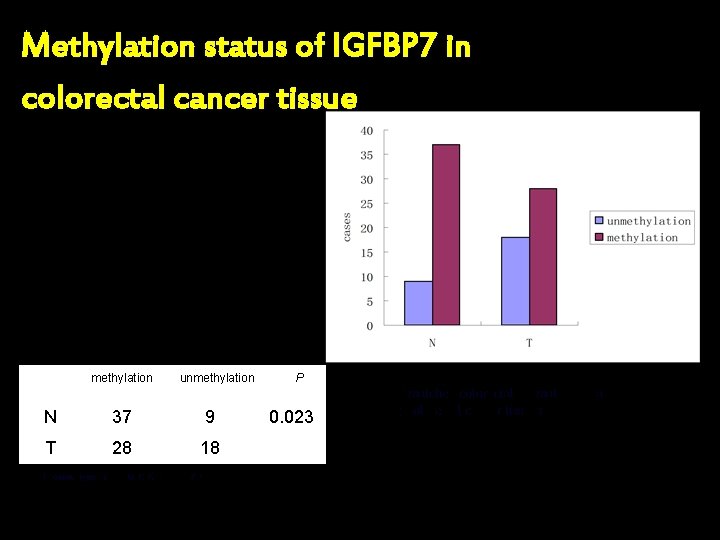
Methylation status of IGFBP 7 in colorectal cancer tissue methylation unmethylation N 37 9 T 28 18 Wilcoxon signed ranks test: Z=-2. 268, P=0. 023 P 0. 023 N: matched colorectal normal mucosa; T: colorectal cancer tissues

Relationship between IGFBP 7 expression and the exon 1 methylation status m. RNA Expression Methylation status (%) P N 0. 21 80. 4 0. 044 T 0. 61 60. 9 r=-0. 210, P=0. 044 IHC unmethylation P Low (0, 1) 15 51 0. 004 High (2, 3) 12 14 r=-0. 299, P=0. 004

III: The biological behaviour of IGFBP 7 in CRC cells生物学行为
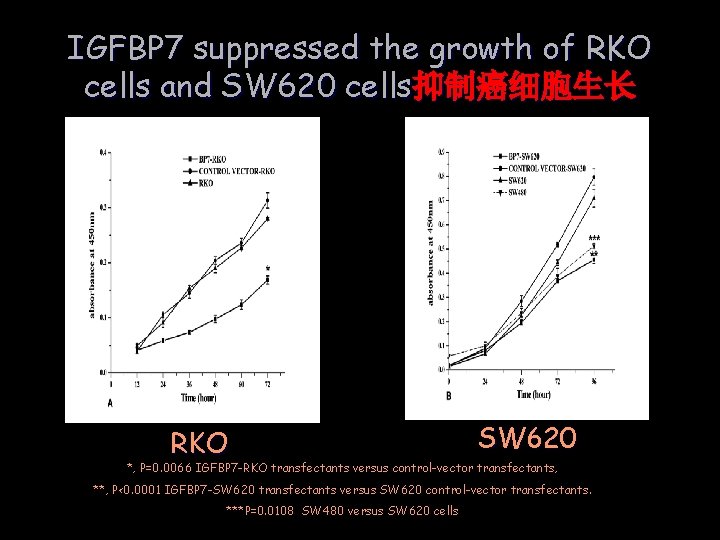
IGFBP 7 suppressed the growth of RKO cells and SW 620 cells抑制癌细胞生长 RKO SW 620 *, P=0. 0066 IGFBP 7 -RKO transfectants versus control-vector transfectants, **, P<0. 0001 IGFBP 7 -SW 620 transfectants versus SW 620 control-vector transfectants. ***P=0. 0108 SW 480 versus SW 620 cells
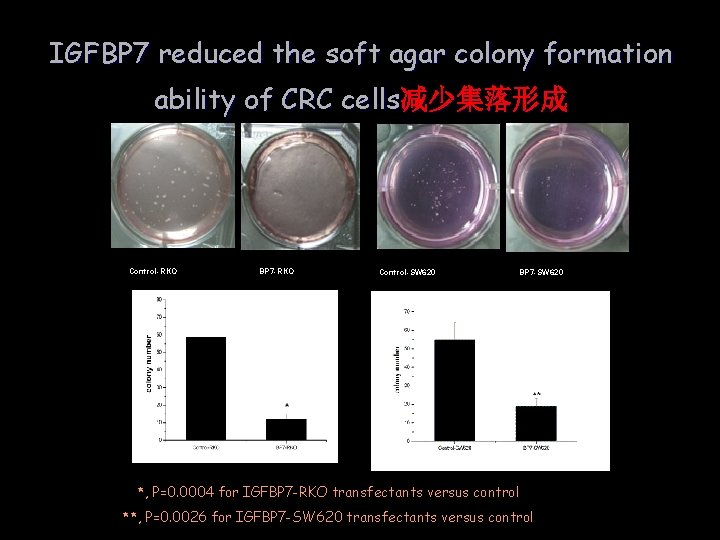
IGFBP 7 reduced the soft agar colony formation ability of CRC cells减少集落形成 Control-RKO BP 7 -RKO Control-SW 620 BP 7 -SW 620 *, P=0. 0004 for IGFBP 7 -RKO transfectants versus control **, P=0. 0026 for IGFBP 7 -SW 620 transfectants versus control
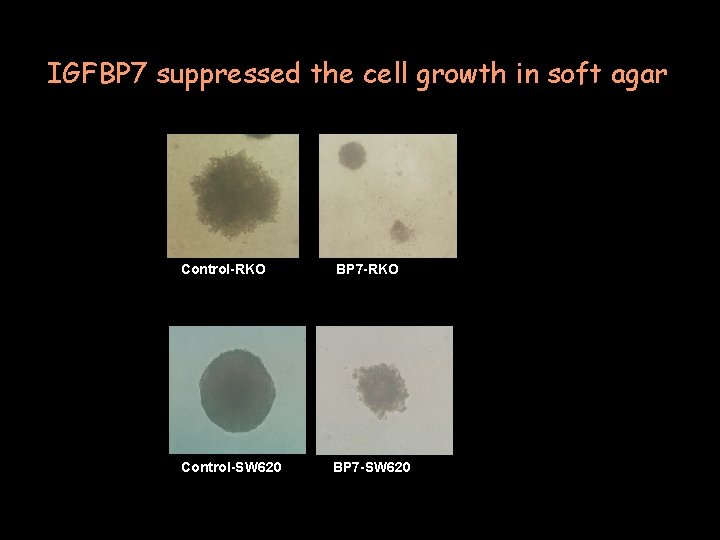
IGFBP 7 suppressed the cell growth in soft agar Control-RKO BP 7 -RKO Control-SW 620 BP 7 -SW 620
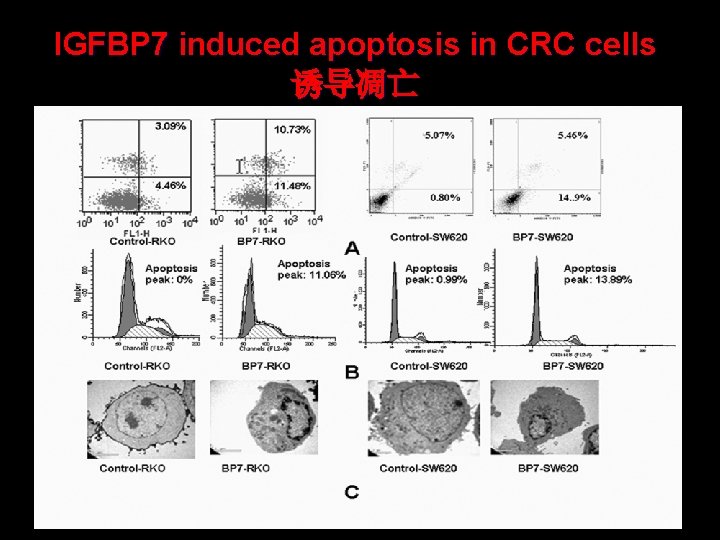
IGFBP 7 induced apoptosis in CRC cells 诱导凋亡
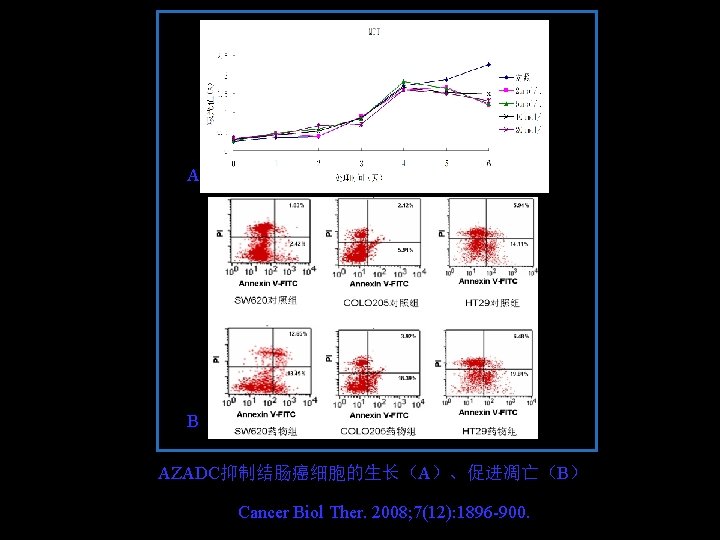
A B AZADC抑制结肠癌细胞的生长(A)、促进凋亡(B) Cancer Biol Ther. 2008; 7(12): 1896 -900.
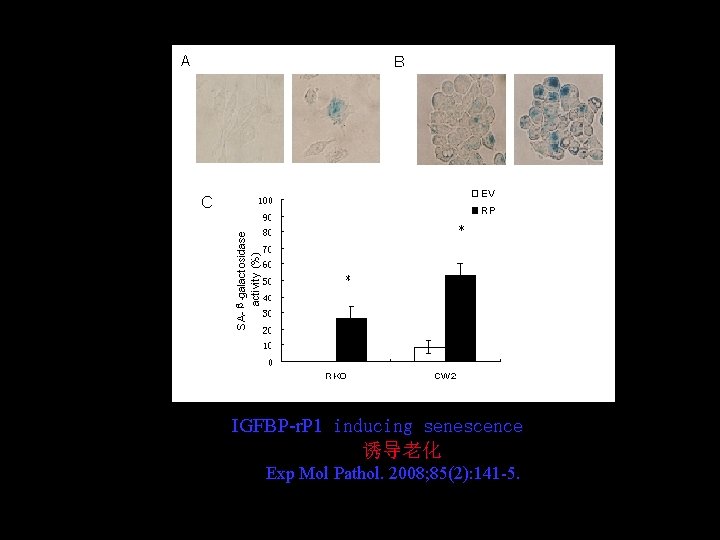
IGFBP-r. P 1 inducing senescence 诱导老化 Exp Mol Pathol. 2008; 85(2): 141 -5.

l IGFBP 7 was a potential tumor suppressor gene in colon cancer 是一个抑癌基因 Cancer Biol Ther. 2007; 6(3): 354 -9 Exp Mol Pathol. 2008; 85(2): 141 -5
l Explore the molecular mechanism…… IGFBP 7 transfectants ? Control What’s different inside
Identify the differentially expressed genes in IGFBP 7 -RKO transfectants versus control RP 5 vs EV 5 RP 7 RP 6 vs vs EV 7 EV 6
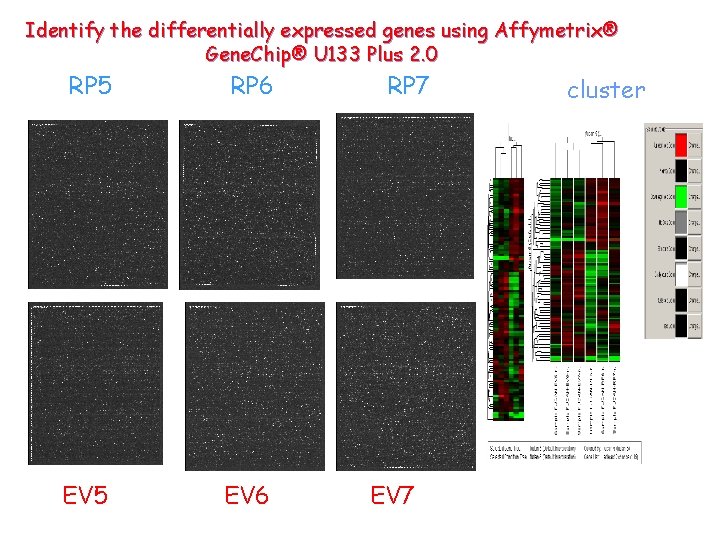
Identify the differentially expressed genes using Affymetrix® Gene. Chip® U 133 Plus 2. 0 RP 5 RP 6 EV 5 EV 6 RP 7 EV 7 cluster

Interesting information resides in the chip results: Reproducible in at least 2 clones: 91 genes KEGG pathway analysis: MAPK pathway: GADD 45 B,FOS,FLNB,NR 4 A 1, RASA 1 TGF- pathway: SMAD 3, CDKN 2 B, ID 1, ID 3 IGFBP 7 could activate MAPK pathway and inhibit TGF- pathway.
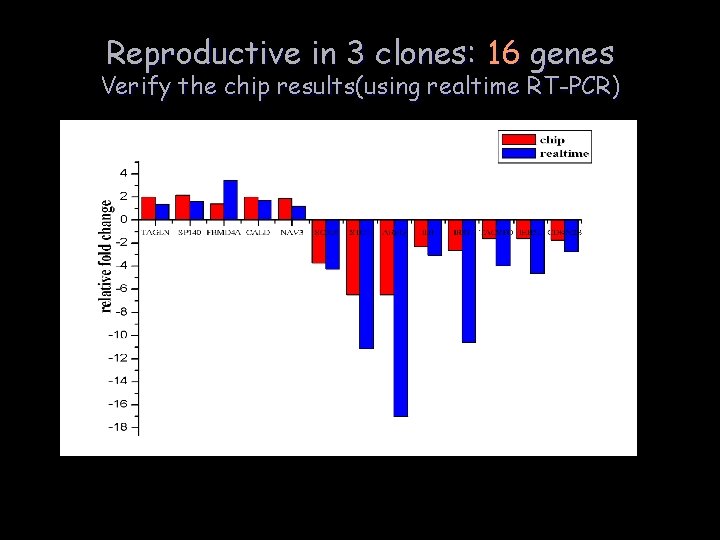
Reproductive in 3 clones: 16 genes Verify the chip results(using realtime RT-PCR)
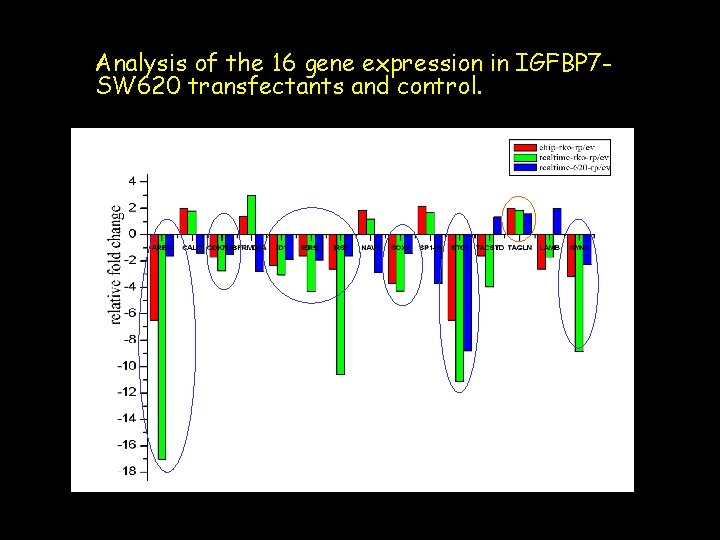
Analysis of the 16 gene expression in IGFBP 7 SW 620 transfectants and control.
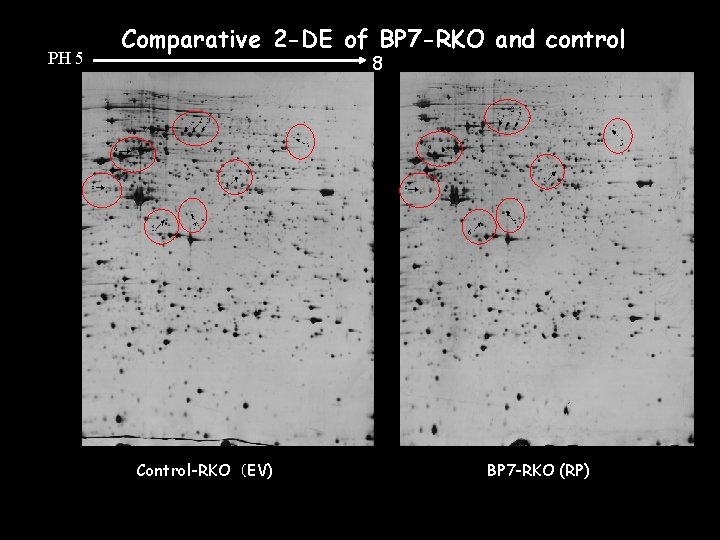
PH 5 Comparative 2 -DE of BP 7 -RKO and control 8 Control-RKO(EV) BP 7 -RKO (RP)

EV RP EV EV Spot RP EV RP Protein description RP EV RP Sequence coverage(%)* Swissprot ID Theoretical Mr /Pi** 1 Serum albumin 5. 74% P 02768 69367/6. 42 2 Serum albumin 7. 97% P 02768 69367/6. 42 3 Serum albumin 6. 86% P 02768 69367/6. 42 4 pyruvate kinase, muscle 22. 45% Q 9 UK 31 6002/7. 58 5 Phenylalanyl-t. RNA synthetase beta chain 12. 56% Q 9 NSD 9 66130/6. 39 6 Actin, cytoplasmic 1 or 2 33. 33% P 63261 41793/5. 31 7 Actin, cytoplasmic 1 or 2 23. 20% P 63261 41793/5. 31 8 60 k. Da heat shock protein, mitochondrial precursor 2. 96% P 10809 61055/5. 7 9 60 k. Da heat shock protein, mitochondrial precursor 28. 52% P 10809 61055/5. 7 *Sequence coverage was calculated by amino acid count **Calculated from the database entry without any processing Unpublished data
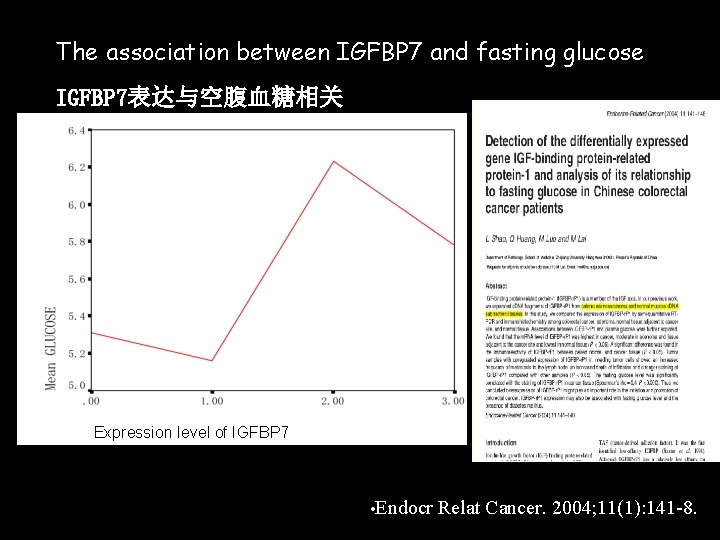
The association between IGFBP 7 and fasting glucose IGFBP 7表达与空腹血糖相关 Expression level of IGFBP 7 • Endocr Relat Cancer. 2004; 11(1): 141 -8.
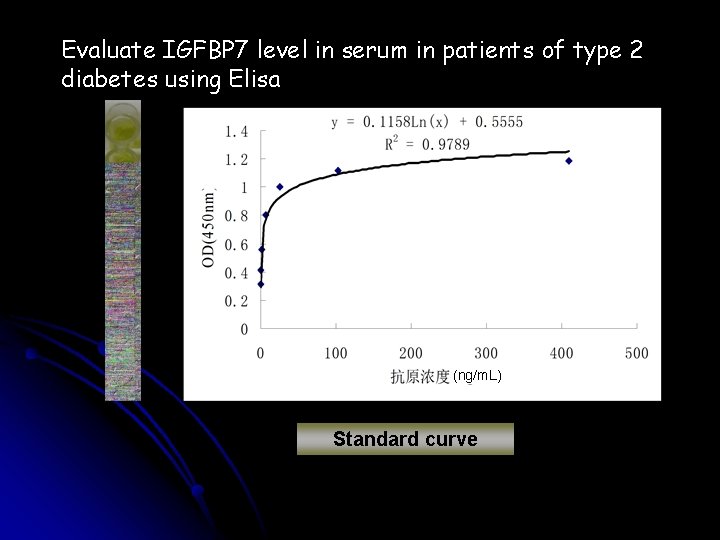
Evaluate IGFBP 7 level in serum in patients of type 2 diabetes using Elisa (ng/m. L) Standard curve
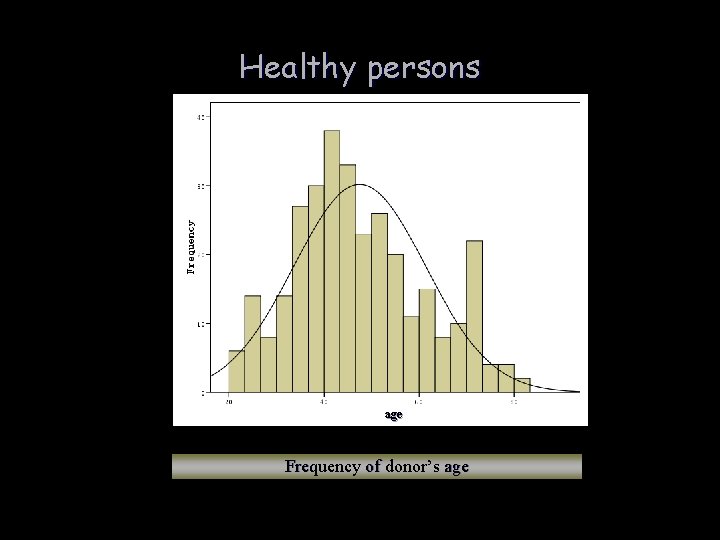
Healthy persons age Frequency of donor’s age
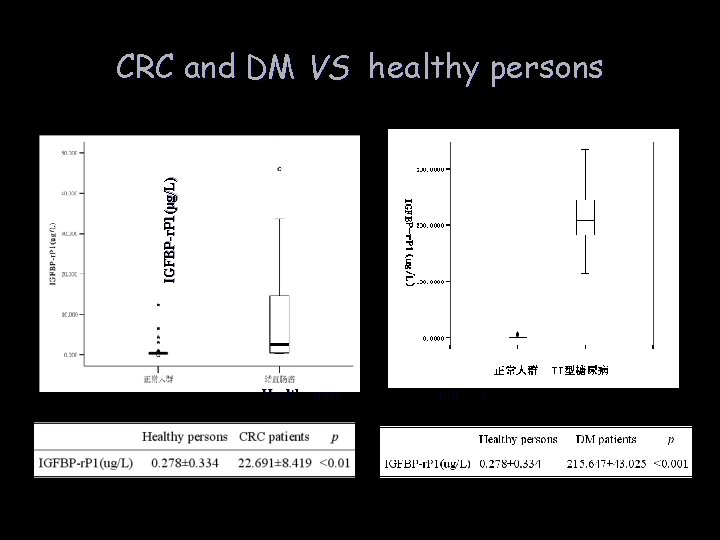
IGFBP-r. P 1(µg/L) CRC and DM VS healthy persons Healthy persons CRC patients
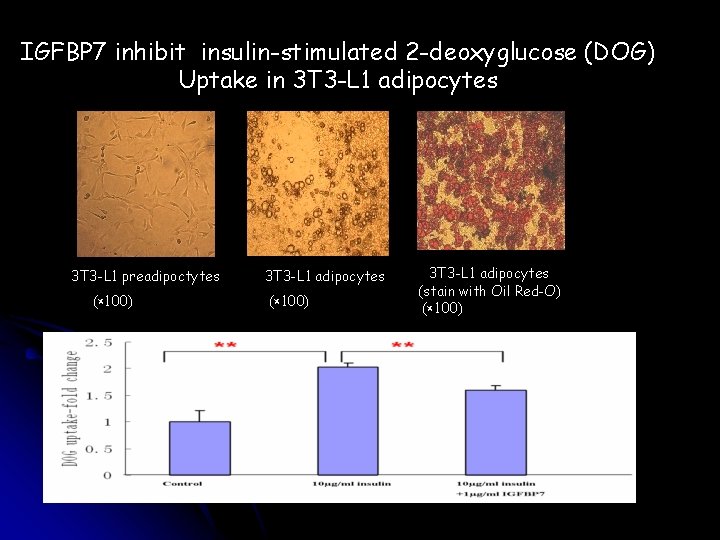
IGFBP 7 inhibit insulin-stimulated 2 -deoxyglucose (DOG) Uptake in 3 T 3 -L 1 adipocytes 3 T 3 -L 1 preadipoctytes (× 100) 3 T 3 -L 1 adipocytes (stain with Oil Red-O) (× 100)

Metabolic syndrome is reversible 代谢综合征是二十一世纪流行病,可逆
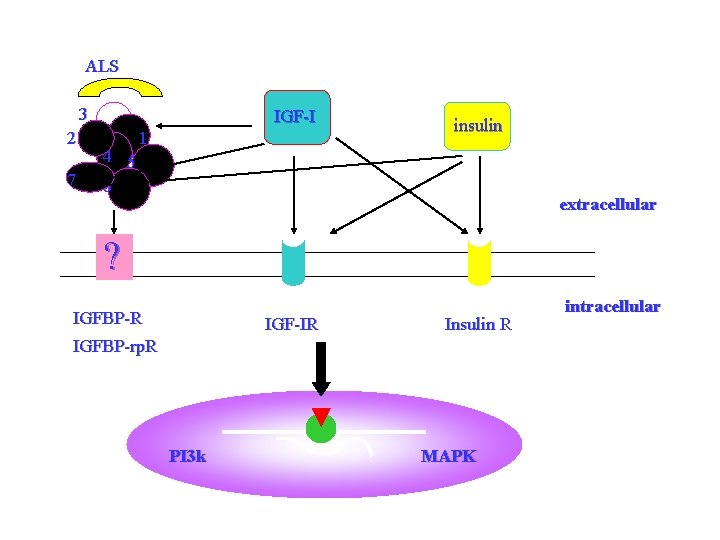
ALS 3 2 7 IGF-I 4 5 6 1 insulin extracellular ? IGFBP-R IGF-IR Insulin R IGFBP-rp. R PI 3 k MAPK intracellular
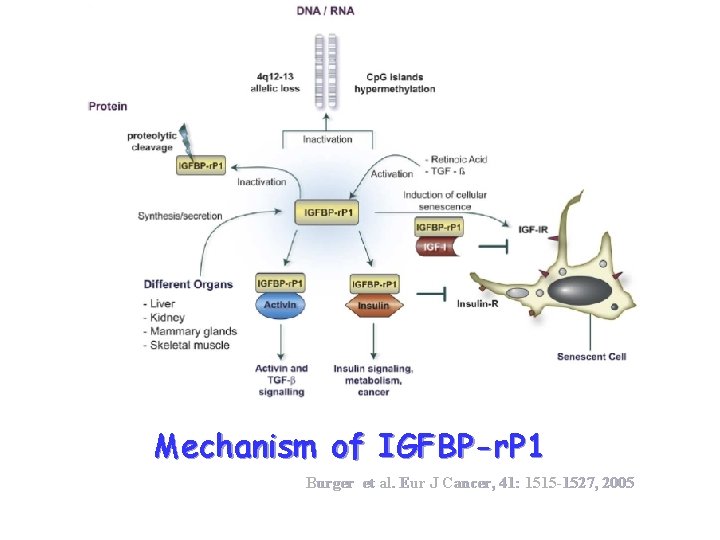
Mechanism of IGFBP-r. P 1 Burger et al. Eur J Cancer, 41: 1515 -1527, 2005

IGFBP 7 may be a molecule contribute to insulin resistance, which may play important roles in the initiation and development of type 2 diabetes. 可能参与2型糖尿病的发生和发展 • Unpublished data

Character of IGFBP 7 l IGFBP 7 binds insulin with high affinity: 500 fold higher than IGFBP 1 -6 does. J Biol Chem. 1997; 272(49): 30729 -34. What’s the binding structure of IGFBP 7 to insulin?
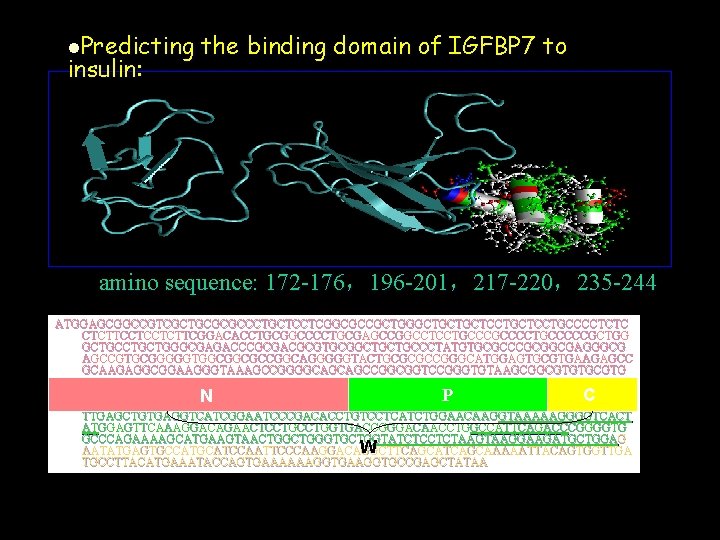
l. Predicting insulin: the binding domain of IGFBP 7 to amino sequence: 172 -176,196 -201,217 -220,235 -244 ATGGAGCGGCCGTCGCTGCGCGCCCTGCTCCTCGGCGCCGCTGGGCTGCTGCTCCTGCCCCTCTC CTCTTCCTCCTCTTCGGACACCTGCGGCCCCTGCGAGCCGGCCTCCTGCCCCTGCCCCCGCTGG GCTGCTGGGCGAGACCCGCGACGCGTGCGGCTGCTGCCCTATGTGCGCCCGCGGCGAGGGCG AGCCGTGCGGGGGTGGCGGCGCCGGCAGGGGGTACTGCGCGCCGGGCATGGAGTGCGTGAAGAGCC GCAAGAGGCGGAAGGGTAAAGCCGGGGCAGCAGCCGGCGGTCCGGGTGTAAGCGGCGTGTGCGTG TGCAAGAGCCGCTACCCGGTGTGCGGCAGCGACGGCACCACCTACCCGAGCGGCTGCCAGCTGCGC GCCGCCAGAGGGCCGAGAGCCGCGGGGAGAAGGCCATCACCCAGGTCAGCAAGGGCACCTG P C N CGAGCAAGGTCCTTCCATAGTGACGCCCCCCAAGGACATCTGGAATGTCACTGGTGCCCAG GTGTAC CGAGCAAGGTCCTTCCATAGTGACGCCCCCCAAGGACATCTGGAATGTCACTGGTGCCCAGGTGTAC TTGAGCTGTGAGGTCATCGGAATCCCGACACCTGTCCTCATCTGGAACAAGGTAAAAAGGGGTCACT ATGGAGTTCAAAGGACAGAACTCCTGGTGACCGGGACAACCTG GCCATTCAGACCCGGGGTG ATGGAGTTCAAAGGACAGAACTCCTGGTGACCGGGACAACCTGGCCATTCAGACC GCCCAGAAAAGCATGAAGTAACTGGGTGCTGGTATCTCCTCTAAGGAAGATGCTGGA G GCCCAGAAAAGCATGAAGTAACTGGGTGCTGGTATCTCCTCTAAGGAAGATGCTGGAG W AATATGAGTGCCATGCATCCAATTCCCAAGGACAGGCTTCAGCAAAAATTACAGTGGTTGA TGCCTTACATGAAATACCAGTGAAAAAAGGTGCCGAGCTATAA
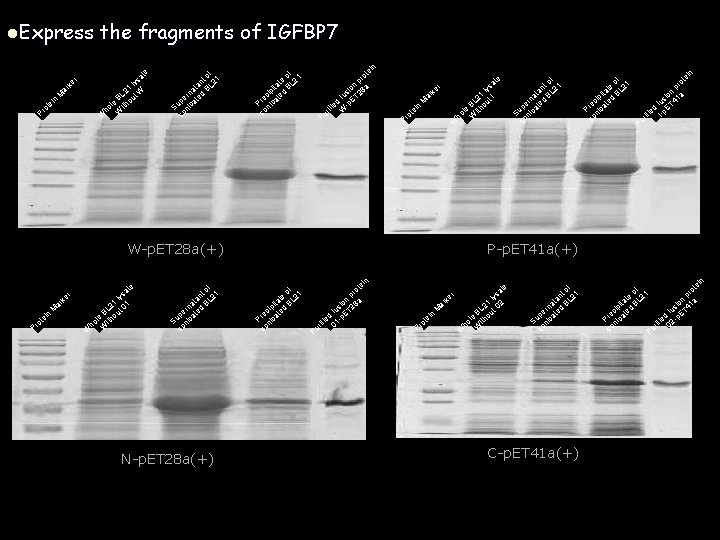
l. Express k er ar Pr ot n ei M the fragments of IGFBP 7 te sa ly 21 W BL out le th ho Wi W of nt L 21 a t na B er ated p Su nic so n ei ot pr on a si 8 fu ET 2 ed -p ifi ur W of 1 te L 2 a it B ip d ec ate r P nic so P e r ke ar M in te o Pr W W-p. ET 28 a(+) r ke ar ot Pr n ei M te sa ly 21 01 BL out le th ho i W W f to 1 an L 2 t na B er ated p Su nic so N-p. ET 28 a(+) f to 1 an L 2 t na B er ated p Su nic so t sa ly 21 t I BL hou le it ho W of 1 te 2 ti a BL ip d ec te Pr nica so in e ot pr n o si 1 a fu T 4 d E e ifi I-p r Pu P-p. ET 41 a(+) of e L 21 t ita B ip d ec ate r P nic so n P ei ot pr on a si 28 fu ET ed p ifi 1 ur 0 a er rk Pr n ei ot M e t sa ly 21 02 BL out le th ho i W W f to 1 an L 2 t na B er ated p Su nic so C-p. ET 41 a(+) i te ro of 1 p e 2 t L n io 1 a ita B ip ed us T 4 c f t e E d Pr nica ie -p rif 02 u so P n

Analyze the binding affinity of the IGFBP 7 fragments to insulin SDS-PAGE cut the gel at 6 KD transmembrane Insulin Incubated with fusion protein wash with His-tag Incubated with His-tag Ab Incubated with secondary Ab For odssey W wash P N C 正在对关键氨基酸进行鉴定

Conclusions 1. The expression of IGFBP 7 was upregulated in CRC tissue. Overexpression of IGFBP 7 correlates with favourable prognosis in CRC patients. 2. Methylation of exon 1 was the key regulating mechanism of IGFBP 7. 3. IGFBP 7 played potential tumor suppressor role in colorectal carcinogenesis. 4. The patients with insulin resistance are associated with higher incidence of some cancers. The serum IGFBP 7 level was higher in type 2 diabetes and colorectal cancer patients. IGFBP 7 may be a molecule contributing to insulin resistance.

Grants l China National “ 973” program (2007 CB 914304) l the National Scientific Foundation of China (30200333, 30570840, 30770989, 30900236) l Zhejiang Natural Science Foundation of China ( D 2080011)

Persons contributed to the research Department of Pathology, Zhejiang University Dr. Maode Lai Dr. Minjie Luo Dr. Lina Shao Dr. Jie Lin Dr. Wenjing Ruan Dr. Yu Ma Dr. Fangying Xu Lipei Wang Haibing Wang Youzhao …… Li Department of Chemistry, Zhejiang University Dr. Tao Wu Dr. Xin Chen
- Slides: 60