Identification of Ortholog Groups by Ortho MCL Protein
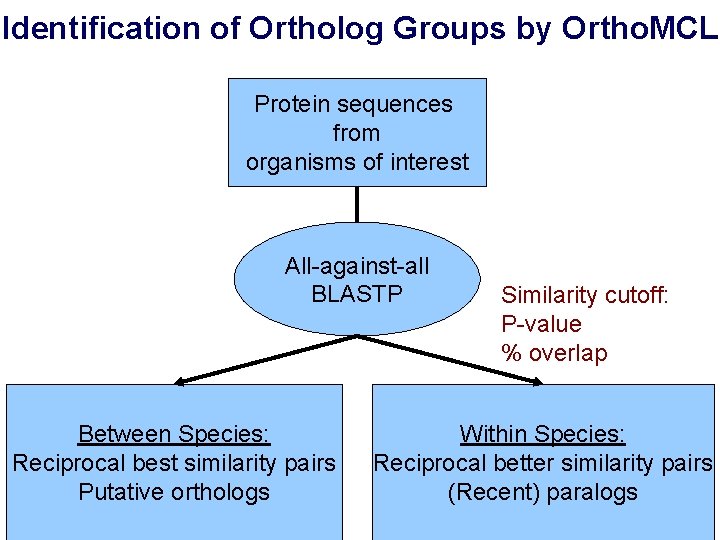
Identification of Ortholog Groups by Ortho. MCL Protein sequences from organisms of interest All-against-all BLASTP Between Species: Reciprocal best similarity pairs Putative orthologs Similarity cutoff: P-value % overlap Within Species: Reciprocal better similarity pairs (Recent) paralogs
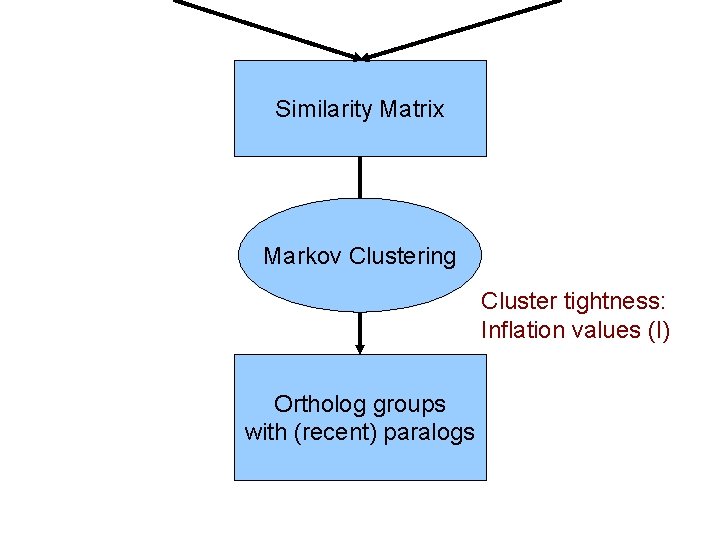
Similarity Matrix Markov Clustering Cluster tightness: Inflation values (I) Ortholog groups with (recent) paralogs
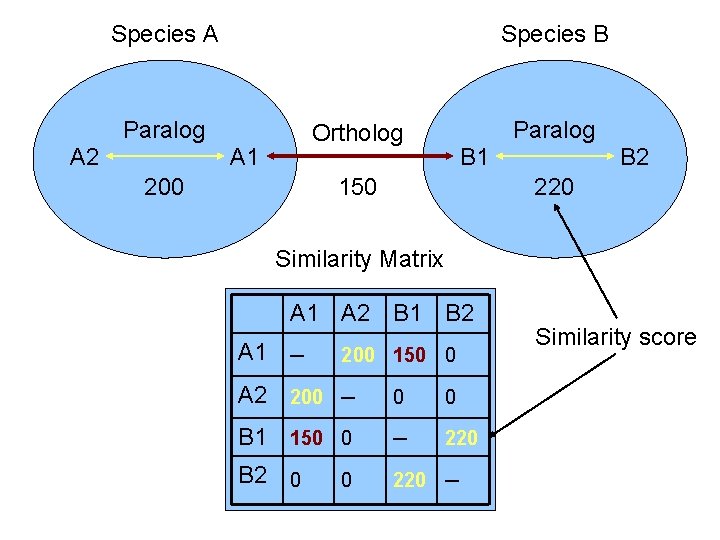
Species A Species B Paralog A 2 Paralog Ortholog A 1 200 B 1 150 B 2 220 Similarity Matrix A 1 A 2 B 1 B 2 A 1 ─ 200 150 0 A 2 200 ─ 0 0 B 1 150 0 ─ 220 B 2 0 220 ─ 0 Similarity score
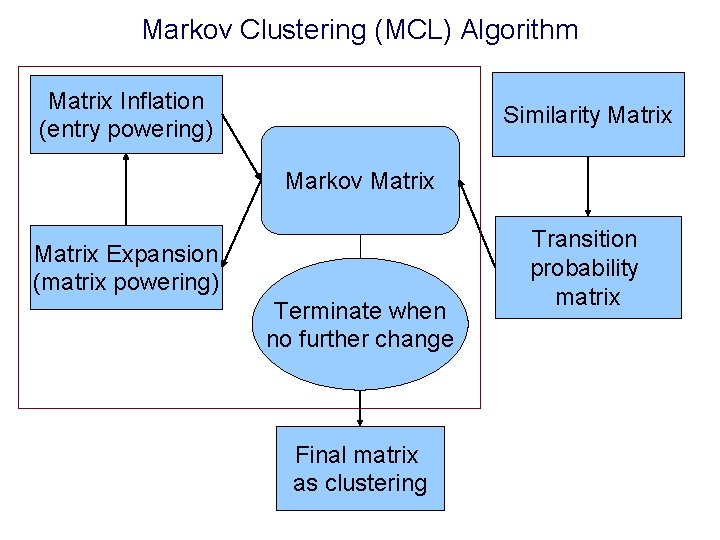
Markov Clustering (MCL) Algorithm Matrix Inflation (entry powering) Similarity Matrix Markov Matrix Expansion (matrix powering) Terminate when no further change Final matrix as clustering Transition probability matrix
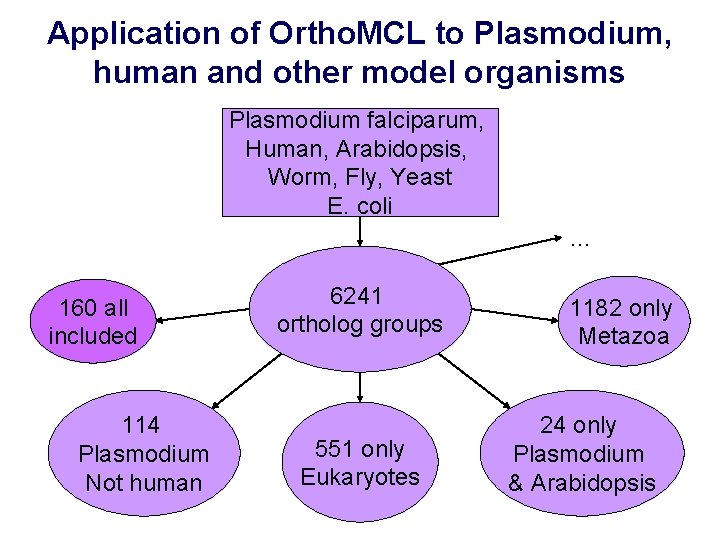
Application of Ortho. MCL to Plasmodium, human and other model organisms Plasmodium falciparum, Human, Arabidopsis, Worm, Fly, Yeast E. coli … 160 all included 114 Plasmodium Not human 6241 ortholog groups 551 only Eukaryotes 1182 only Metazoa 24 only Plasmodium & Arabidopsis
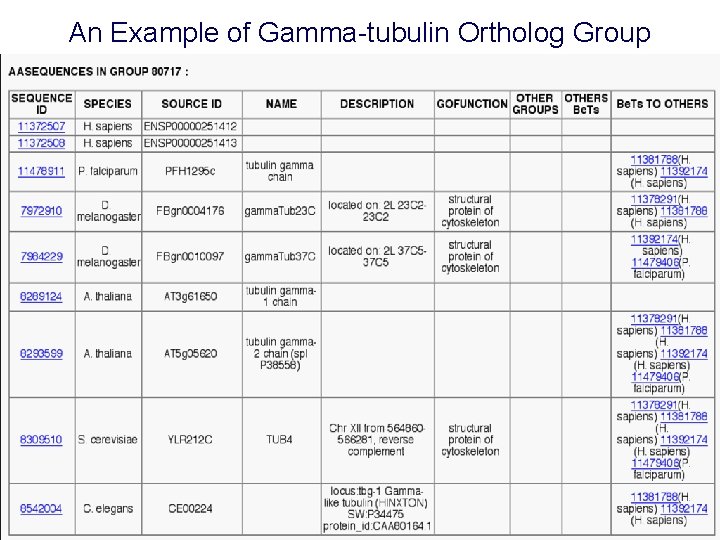
An Example of Gamma-tubulin Ortholog Group
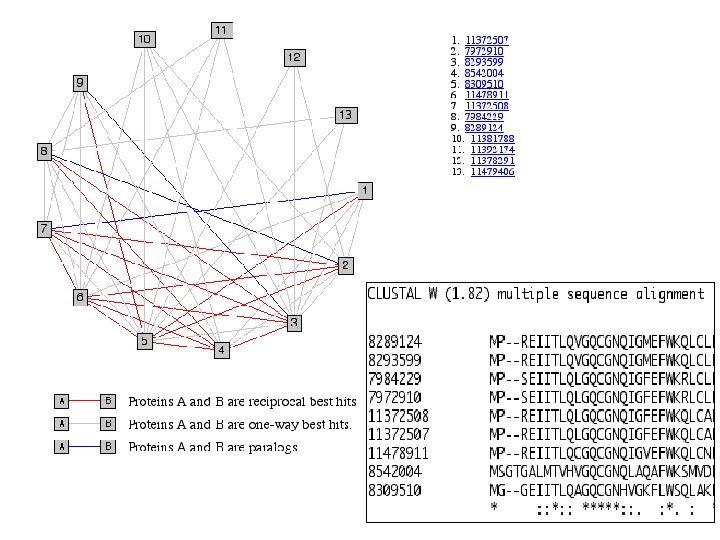

Comparing Ortho. MCL with INPARANOID ( two species) • INPARANOID clusters both orthologs and in-paralogs from two species by pairwise similarity – Find two-way best hits from pairwise similarity scores as main ortholog pair – Add additional orthologs (in-paralogs) from the same species for each main ortholog by comparing similarity scores between the main ortholog with putative in-paralogs with the score between the main ortholog pair – Resolve overlapping groups by merging, deleting, dividing them based on a set of rules • Ortho. MCL can cluster orthologs and in-paralogs from multiple species
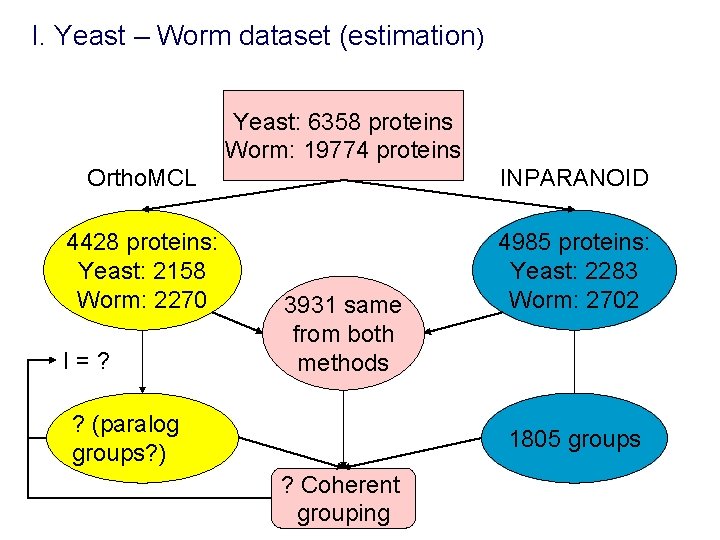
I. Yeast – Worm dataset (estimation) Yeast: 6358 proteins Worm: 19774 proteins Ortho. MCL INPARANOID 4428 proteins: Yeast: 2158 Worm: 2270 4985 proteins: Yeast: 2283 Worm: 2702 I=? 3931 same from both methods ? (paralog groups? ) 1805 groups ? Coherent grouping
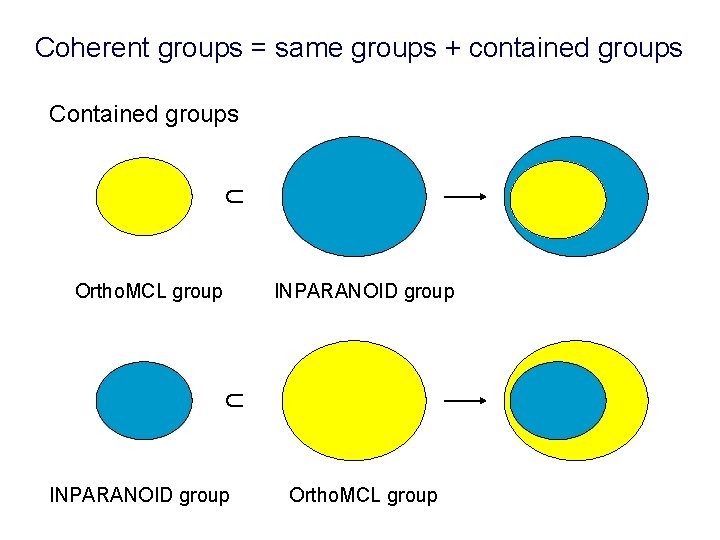
Coherent groups = same groups + contained groups ∩ Contained groups INPARANOID group ∩ Ortho. MCL group INPARANOID group Ortho. MCL group
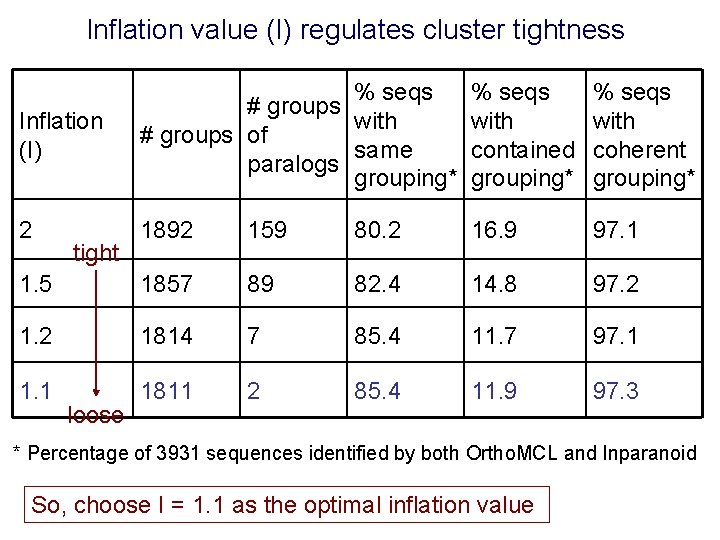
Inflation value (I) regulates cluster tightness Inflation (I) % seqs # groups with # groups of same paralogs grouping* % seqs with contained grouping* % seqs with coherent grouping* 2 1892 159 80. 2 16. 9 97. 1 1. 5 1857 89 82. 4 14. 8 97. 2 1814 7 85. 4 11. 7 97. 1 1811 2 85. 4 11. 9 97. 3 tight loose * Percentage of 3931 sequences identified by both Ortho. MCL and Inparanoid So, choose I = 1. 1 as the optimal inflation value

Possible reasons for including different sequences Ortho. MCL INPARANOID BLAST version WU-BLAST NCBI-BLAST Search All-against-all, SEG filtered, Pairwise fixed database size Similarity cutoff P<1 e-5 Score>=50 bits Overlap > 50% Reciprocal “best” hits P-value, percent identity Score Recent paralogs One-way better Bi-directional better within-species similarity from similarity orthologs
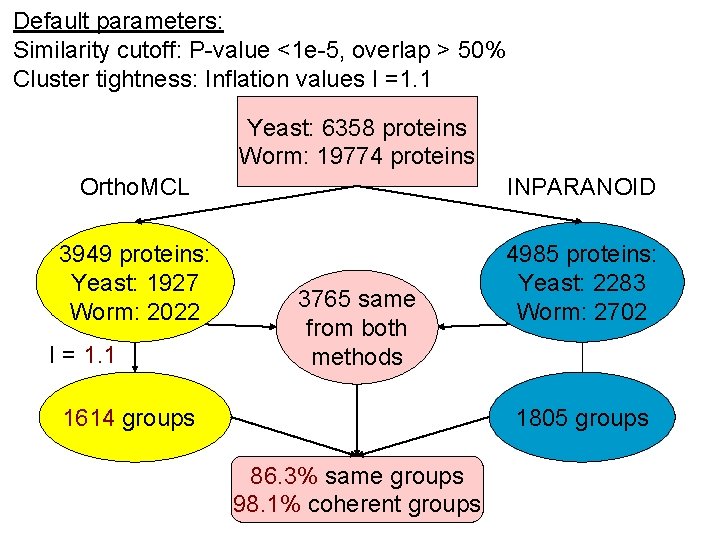
Default parameters: Similarity cutoff: P-value <1 e-5, overlap > 50% Cluster tightness: Inflation values I =1. 1 Yeast: 6358 proteins Worm: 19774 proteins Ortho. MCL INPARANOID 3949 proteins: Yeast: 1927 Worm: 2022 4985 proteins: Yeast: 2283 Worm: 2702 I = 1. 1 3765 same from both methods 1805 groups 1614 groups 86. 3% same groups 98. 1% coherent groups
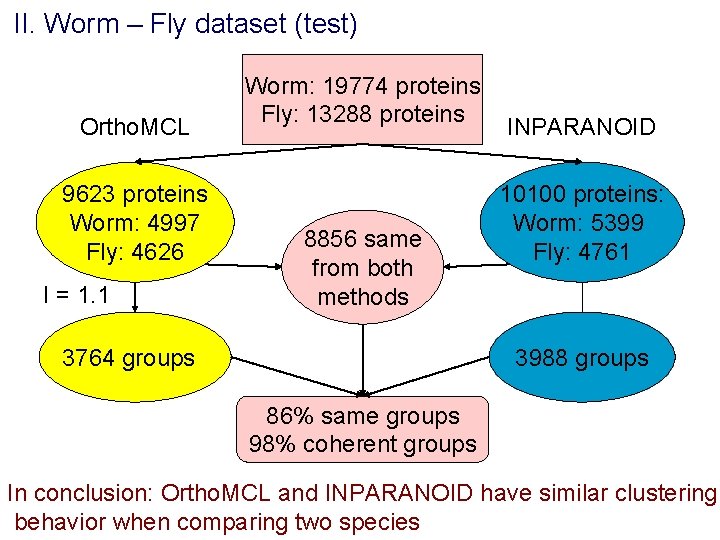
II. Worm – Fly dataset (test) Ortho. MCL 9623 proteins Worm: 4997 Fly: 4626 I = 1. 1 Worm: 19774 proteins Fly: 13288 proteins 8856 same from both methods INPARANOID 10100 proteins: Worm: 5399 Fly: 4761 3988 groups 3764 groups 86% same groups 98% coherent groups In conclusion: Ortho. MCL and INPARANOID have similar clustering behavior when comparing two species

Comparison of Ortho. MCL with EGO (multiple species) III. Yeast – Worm – Fly dataset EGO: TC/NP BLASTP 10260 seqs Protein sequences 4776 proteins Remove redundancy 4776 unique proteins formed 3125 unique groups Ortho. MCL: 12459 proteins formed 4033 groups
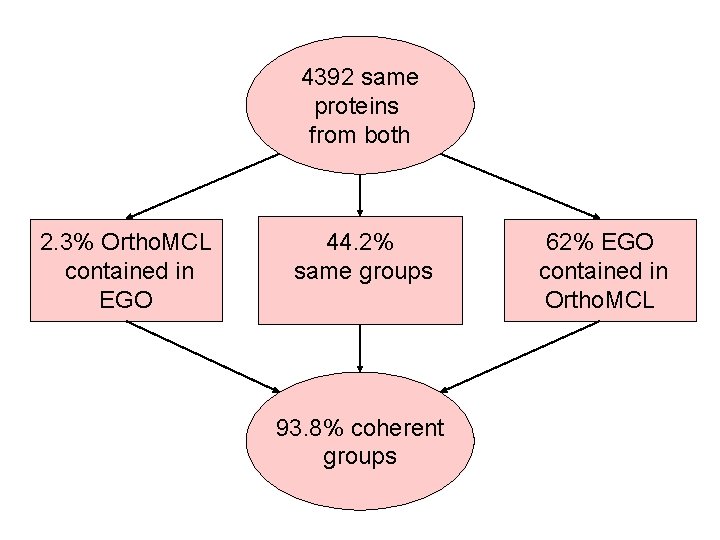
4392 same proteins from both 2. 3% Ortho. MCL contained in EGO 44. 2% same groups 93. 8% coherent groups 62% EGO contained in Ortho. MCL
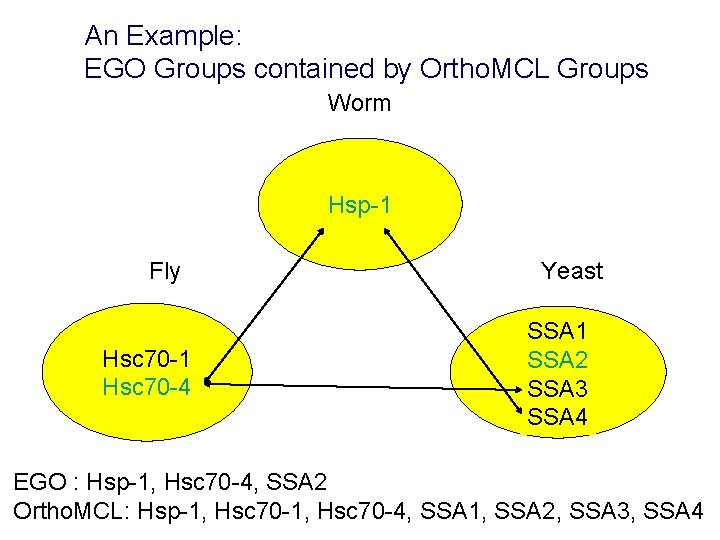
An Example: EGO Groups contained by Ortho. MCL Groups Worm Hsp-1 Fly Hsc 70 -1 Hsc 70 -4 Yeast SSA 1 SSA 2 SSA 3 SSA 4 EGO : Hsp-1, Hsc 70 -4, SSA 2 Ortho. MCL: Hsp-1, Hsc 70 -4, SSA 1, SSA 2, SSA 3, SSA 4
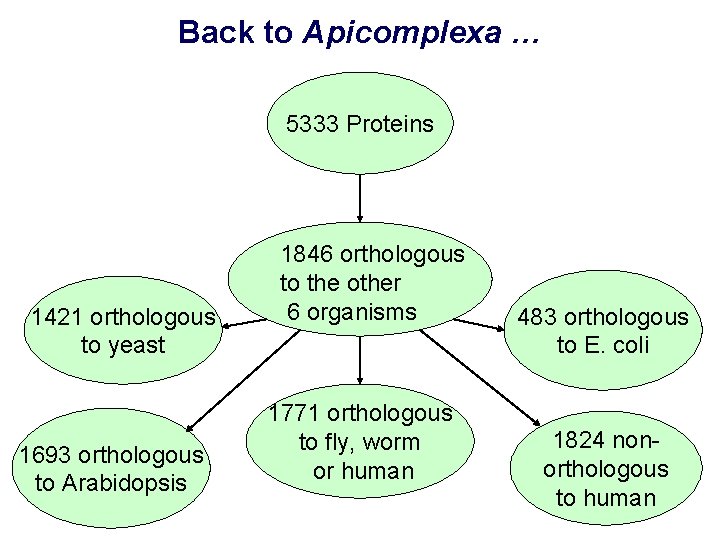
Back to Apicomplexa … 5333 Proteins 1421 orthologous to yeast 1693 orthologous to Arabidopsis 1846 orthologous to the other 6 organisms 1771 orthologous to fly, worm or human 483 orthologous to E. coli 1824 nonorthologous to human

Summary • Ortho. MCL automatically delineates the many-to-many orthologous relationship across multiple eukaryotic genomes • When applied to pairwise comparison of two species, the performance of Ortho. MCL is comparable to INPARANOID which was designed for comparing two species • When applied to multiple species and compared with EGO database, Ortho. MCL tend to identify more orthologous genes • The underlying object-based relational storage model permits integration with organismal data and queries based on user-defined species distribution provides a snapshot of shared/diversified biological processes across species

Related Posters and Reference • 114 A. Web-Based Biological Discovery using an Integrated Database. • 146 A. The Genomics Unified Schema (GUS). • 170 A. TESS-II: Describing and Finding Gene Regulatory Sequences with Grammars. • Remm et al. Automatic Clustering of Orthologs and Inparalogs from Pairwise Species Comparisons. J. MOL. Biol. (2001) 314 • Lee et al. Cross-Referencing Eukaryotic Genomes: TIGR Orthologous Gene Alignments (TOGA). Genome Res. (2002) 12 • Enright et al. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. (2002) 30
- Slides: 20