Identification and Phylogeny of the Human Endogenous Retrovirus
- Slides: 1

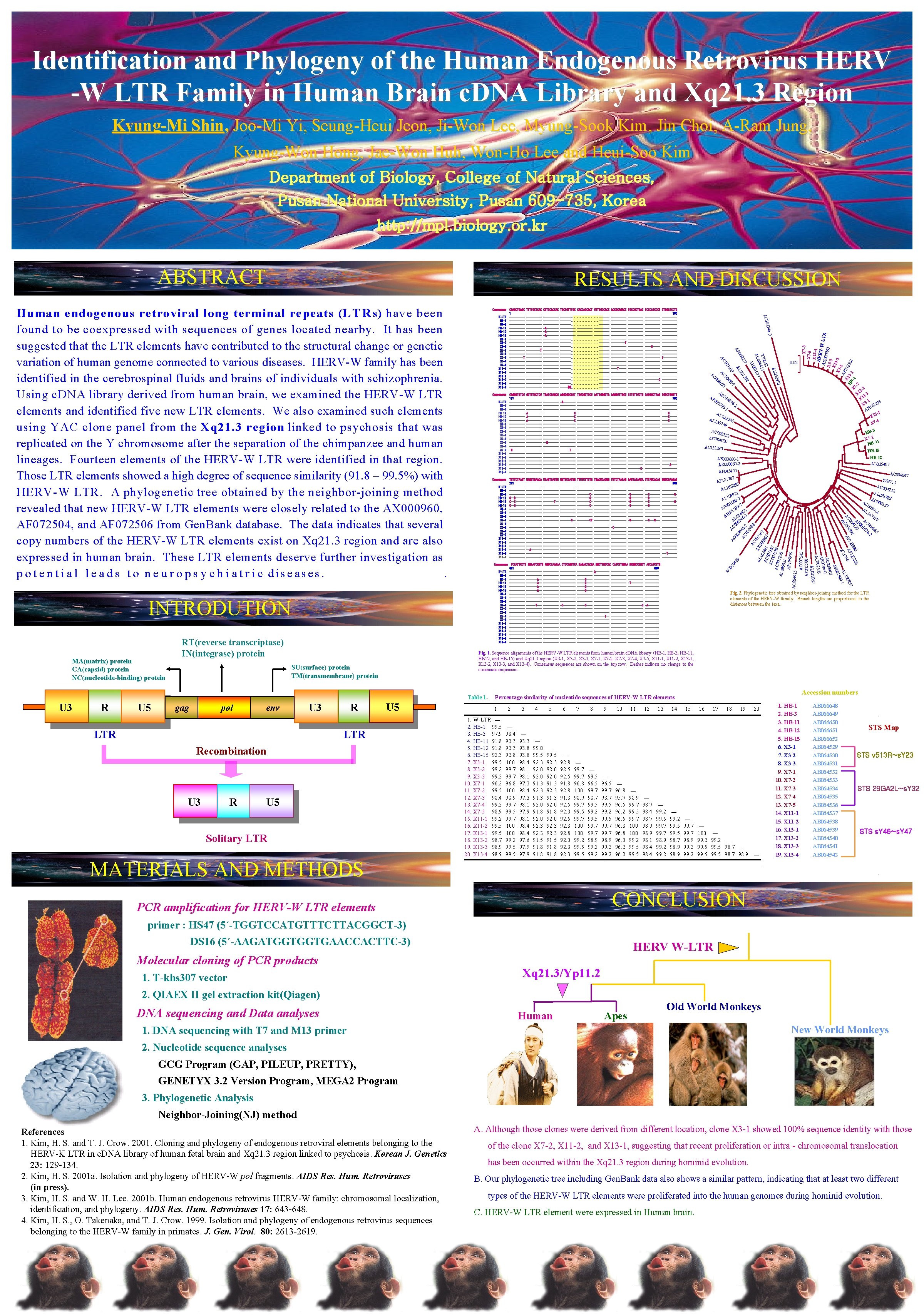
Identification and Phylogeny of the Human Endogenous Retrovirus HERV -W LTR Family in Human Brain c. DNA Library and Xq 21. 3 Region Kyung-Mi Shin, Joo-Mi Yi, Seung-Heui Jeon, Ji-Won Lee, Myung-Sook Kim, Jin Choi, A-Ram Jung, Kyung-Won Hong, Jae-Won Huh, Won-Ho Lee and Heui-Soo Kim Department of Biology, College of Natural Sciences, Pusan National University, Pusan 609 -735, Korea http: //mpl. biology. or. kr ABSTRACT RESULTS AND DISCUSSION INTRODUTION MA(matrix) protein CA(capsid) protein NC(nucleotide-binding) protein RT(reverse transcriptase) IN(integrase) protein R U 5 SU(surface) protein TM(transmembrane) protein gag pol env U 3 LTR R U 5 LTR Recombination U 3 R AP 02 00 05 68 HERV X 7 -3 X 7 -5 X 13 -4 2 1 X 1 3 -1 X 1 1 6 X 3 7250 0 AF 3 -2 X 1 4 X 7 - 98 00 -2 -1 AL -W LT AX 0 R 0096 0 X 33 X 13 -3 X 3 AF 2 07 X 25 HB 1104 1 X 7 - 1 2 AB 022 AL 1 357 49 396 3 HBX 7 -1 1 HB-1 AC 0 0035 2 AC 004 020 AL 031390 HB-15 HB-12 AL 035407 AE 000660 -1 AE 000660 -2 AF 045450 AC 004067 1782 AF 12 0 6328 1 L A 22 099 1 L A -2 00 16 -2 0 0 99 AP 15 402 0 0 4 AP L 03 6 -1 A 994 00 AC Z 69715 AC 00 AL 0 99 AC 4600 2 24 88 AC 0 AX 07 AL 56 0 16 28 0096 7 91 AC 7 0 AC 05187 007 53 AC 0 0725 8 AL 09 6821 0 AF 19697 0 AC 004915 AC 007543 AF 205588 AC 00 94 89 TCCATTCCTT GGAATCCGTG AGGCCAAGAA CTCCAGGTCA GAGAATACGA GGCTTGCCAC CATCTTGGAA GCGGCCTGCT ACCATCTTG 301 389 ---------- ----- A---------- ---------- ---------- ---------A-- ----------C-----------------A-- ----------C---------- -T------------------A-- ---------- -----C-------------------------------- ---------- ---------- ---------- ---------- ---------- ---------T--- ----- -C-----C----------C -A---------------- ---------- ---------- ---------- ---------- ----------T-- ---------- ---------- ------------------ ---------- ---------- ---------- ---------- ---------- ---------- ------------------ ---------- ---------- ---------- ---------- ---------- ----- AC 4242 3198 006 137 AC 00 GAGCTGAACA CTAGTCACTG GGTTCCATGG TTCTCTTCTG TGACCCACGG CTTCTAATAG AACTATAACA CTTACCACAT GGCCCAAGAT 300 ---------- ---------- ---------- ---------- ------------------- ----C- ---------- -G---- --C-------G---------- ----C- ---------- -G---- --C---------- -------T-- --------C- ---------- -G---- --C-------G---------- ----C- ---------- -G---- --C------------------- ---------- ---------- ---------- ---------- ---------- -------------------C-- ----CT ---------- -G---- --C---G--- ---------- ---------- ------------------- ----- -C---------- -------------------- ---------- ---------- ---------- ------------------- ---------- ---------- -------------------------------A---------- ---------- ----------- ---------- ----- ---T---------- ---------- ---------- 3 00 AL 301 16 4 32 19 AC 00 48 65 Fig. 2. Phylogenetic tree obtained by neighbor-joining method for the LTR elements of the HERV-W family. Branch lengths are proportional to the distances between the taxa. Fig. 1. Sequence alignments of the HERV-W LTR elements from human brain c. DNA library (HB-1, HB-3, HB-11, HB 12, and HB-15) and Xq 21. 3 region (X 3 -1, X 3 -2, X 3 -3, X 7 -1, X 7 -2, X 7 -3, X 7 -4, X 7 -5, X 11 -1, X 11 -2, X 13 -1, X 13 -2, X 13 -3, and X 13 -4). Consensus sequences are shown on the top row. Dashes indicate no change to the consensus sequences. Table 1. U 3 GCTGTGCTCC TGATCCAGCG AGGCGCCCAT TGCCGCTCCC AATTGGGCTA AAGGCTTGCC ATTGTTCCTG CACGGCTAAG TGCCTGGGTT 200 ---------- -A---------- ---------- ---------- ------------------- ---------- ----------A---------- ---------C ---------- ---------- ----------C----A---------C ---------- -----C-------------- ---------- ---------- ---------- ------------------- ---------- ---------- ---------- ------------ ---------- --------------- --G---------- ---------- ---------- -------------------- ------T--- ---------- ---------- ---------- ---------- ---------- ------------------- ---------- ---------- ---------- -----A---------- ---------- ----C- ------------------- ---------- ----- 23 AC W-LTR HB-1 HB-3 HB-11 HB-12 HB-15 X 3 -1 X 3 -2 X 3 -3 X 7 -1 X 7 -2 X 7 -3 X 7 -4 X 7 -5 X 11 -1 X 11 -2 X 13 -1 X 13 -2 X 13 -3 X 13 -4 60 2 467 01 P 0 A 29 0 26 72 88 41 6 00 499 F 123 AF 12 857 A AC 00 46 132 7 AL -1 75 AC Z 99 015 AP 0 8407 AC 00 9441 AB 01 8 55 AC 002 AL 022067 Consensus 00 310 W-LTR HB-1 HB-3 HB-11 HB-12 HB-15 X 3 -1 X 3 -2 X 3 -3 X 7 -1 X 7 -2 X 7 -3 X 7 -4 X 7 -5 X 11 -1 X 11 -2 X 13 -1 X 13 -2 X 13 -3 X 13 -4 TGTTCTAATT 201 --------------C--C-----C--C------------------------------------------------------------------ AC 0. 02 50 AL 0 Consensus -2 W-LTR HB-1 HB-3 HB-11 HB-12 HB-15 X 3 -1 X 3 -2 X 3 -3 X 7 -1 X 7 -2 X 7 -3 X 7 -4 X 7 -5 X 11 -1 X 11 -2 X 13 -1 X 13 -2 X 13 -3 X 13 -4 CAGGGTGTCC 101 -----------------------------------------------------------G------------------------------- 7244 Consensus TTTTGCTCAC CGTCCACCAC TGCTGTTTGC CACCACCACT GTTTGCCACC ACCGCAGACC TGCCGCTGAC TCCCATCCCT CTGGATCCTG 100 ----------. . . . ---------- ---------- ------------------- -------. . . . ---------- ---------- ------------------- -A---------- ---------- ----------- -A---------- ---------- ---------- -------. . . . ---------- ----------- -------. . . . -------T ---------- -------. . . . ---------- ----G---------T ----------. . . . ---------- ---------- ------------------- -------. . . . ---------- ----- T------------------ -------. . . . ---------- ---------- ---------- -------. . . . -------T----------------------------- -------. . . . ---------- ---------- -------. . . . ---------- -----C ---------- -------. . . . ---------- ----------- -----GG. . . . ---------- ---------- AC 00 W-LTR HB-1 HB-3 HB-11 HB-12 HB-15 X 3 -1 X 3 -2 X 3 -3 X 7 -1 X 7 -2 X 7 -3 X 7 -4 X 7 -5 X 11 -1 X 11 -2 X 13 -1 X 13 -2 X 13 -3 X 13 -4 CGAGCTGAGC 1 ------------------------------------------------------T ------------------------------------- 042 Z 70 455 004 677 AC 001 AP 8 36 27 21 00 00 L 0 A AP 97 09 48 25 00 00 AC AC Human endogenous retroviral long terminal repeats (L T R s) have been found to be coexpressed with sequences of genes located nearby. It has been suggested that the LTR elements have contributed to the structural change or genetic variation of human genome connected to various diseases. HERV-W family has been identified in the cerebrospinal fluids and brains of individuals with schizophrenia. Using c. DNA library derived from human brain, we examined the HERV-W LTR elements and identified five new LTR elements. We also examined such elements using YAC clone panel from the Xq 21. 3 region linked to psychosis that was replicated on the Y chromosome after the separation of the chimpanzee and human lineages. Fourteen elements of the HERV-W LTR were identified in that region. Those LTR elements showed a high degree of sequence similarity (91. 8 – 99. 5%) with HERV-W LTR. A phylogenetic tree obtained by the neighbor-joining method revealed that new HERV-W LTR elements were closely related to the AX 000960, AF 072504, and AF 072506 from Gen. Bank database. The data indicates that several copy numbers of the HERV-W LTR elements exist on Xq 21. 3 region and are also expressed in human brain. These LTR elements deserve further investigation as p o t e n t i a l l e a d s t o n e u r o p s y c h i a t r i c diseases. . Consensus U 5 Solitary LTR Accession numbers Percentage similarity of nucleotide sequences of HERV-W LTR elements 1 1. W-LTR ㅡ 2. HB-1 99. 5 3. HB-3 97. 9 4. HB-11 91. 8 5. HB-12 91. 8 6. HB-15 92. 3 7. X 3 -1 99. 5 8. X 3 -2 99. 2 9. X 3 -3 99. 2 10. X 7 -1 96. 2 11. X 7 -2 99. 5 12. X 7 -3 98. 4 13. X 7 -4 99. 2 14. X 7 -5 98. 9 15. X 11 -1 99. 2 16. X 11 -2 99. 5 17. X 13 -1 99. 5 18. X 13 -2 98. 7 19. X 13 -3 98. 9 20. X 13 -4 98. 9 2 ㅡ 98. 4 92. 3 92. 8 100 99. 7 96. 8 100 98. 9 99. 7 99. 5 99. 7 100 99. 2 99. 5 3 ㅡ 93. 3 93. 8 98. 4 98. 1 97. 3 98. 4 97. 3 98. 1 97. 9 98. 1 98. 4 97. 6 97. 9 4 ㅡ 99. 0 99. 5 92. 3 92. 0 91. 3 92. 3 91. 3 92. 0 91. 8 92. 0 92. 3 91. 5 91. 8 5 ㅡ 99. 5 92. 3 92. 0 91. 3 92. 3 91. 3 92. 0 91. 8 92. 0 92. 3 91. 5 91. 8 6 ㅡ 92. 8 92. 5 91. 8 92. 8 91. 8 92. 5 92. 3 92. 5 92. 8 92. 0 92. 3 7 ㅡ 99. 7 96. 8 100 98. 9 99. 7 99. 5 99. 7 100 99. 2 99. 5 8 ㅡ 99. 5 96. 5 99. 7 98. 7 99. 5 99. 2 99. 5 99. 7 98. 9 99. 2 9 ㅡ 96. 5 99. 7 98. 7 99. 5 99. 2 99. 5 99. 7 98. 9 99. 2 10 ㅡ 96. 8 95. 7 96. 5 96. 2 96. 5 96. 8 96. 0 96. 2 11 ㅡ 98. 9 99. 7 99. 5 99. 7 100 99. 2 99. 5 12 ㅡ 98. 7 98. 4 98. 7 98. 9 98. 1 98. 4 13 ㅡ 99. 2 99. 5 99. 7 98. 9 99. 2 14 ㅡ 99. 2 99. 5 98. 7 98. 9 15 ㅡ 99. 7 98. 9 99. 2 16 ㅡ 100 99. 2 99. 5 17 18 19 20 ㅡ 99. 2 ㅡ 99. 5 98. 7 98. 9 ㅡ MATERIALS AND METHODS 1. HB-1 2. HB-3 3. HB-11 4. HB-12 5. HB-15 6. X 3 -1 7. X 3 -2 8. X 3 -3 9. X 7 -1 10. X 7 -2 11. X 7 -3 12. X 7 -4 13. X 7 -5 14. X 11 -1 15. X 11 -2 16. X 13 -1 17. X 13 -2 18. X 13 -3 19. X 13 -4 AB 066648 AB 066649 AB 066650 AB 066651 AB 066652 AB 064529 AB 064530 AB 064531 AB 064532 AB 064533 AB 064534 AB 064535 AB 064536 AB 064537 AB 064538 AB 064539 AB 064540 AB 064541 AB 064542 STS Map STS v 513 R~s. Y 23 STS 29 GA 2 L~s. Y 32 STS s. Y 46~s. Y 47 CONCLUSION PCR amplification for HERV-W LTR elements primer : HS 47 (5´-TGGTCCATGTTTCTTACGGCT-3) DS 16 (5´-AAGATGGTGGTGAACCACTTC-3) Molecular cloning of PCR products 1. T-khs 307 vector HERV W-LTR Xq 21. 3/Yp 11. 2 2. QIAEX II gel extraction kit(Qiagen) DNA sequencing and Data analyses Human Apes Old World Monkeys New World Monkeys 1. DNA sequencing with T 7 and M 13 primer 2. Nucleotide sequence analyses GCG Program (GAP, PILEUP, PRETTY), GENETYX 3. 2 Version Program, MEGA 2 Program 3. Phylogenetic Analysis Neighbor-Joining(NJ) method References 1. Kim, H. S. and T. J. Crow. 2001. Cloning and phylogeny of endogenous retroviral elements belonging to the HERV-K LTR in c. DNA library of human fetal brain and Xq 21. 3 region linked to psychosis. Korean J. Genetics 23: 129 -134. 2. Kim, H. S. 2001 a. Isolation and phylogeny of HERV-W pol fragments. AIDS Res. Hum. Retroviruses (in press). 3. Kim, H. S. and W. H. Lee. 2001 b. Human endogenous retrovirus HERV-W family: chromosomal localization, identification, and phylogeny. AIDS Res. Hum. Retroviruses 17: 643 -648. 4. Kim, H. S. , O. Takenaka, and T. J. Crow. 1999. Isolation and phylogeny of endogenous retrovirus sequences belonging to the HERV-W family in primates. J. Gen. Virol. 80: 2613 -2619. A. Although those clones were derived from different location, clone X 3 -1 showed 100% sequence identity with those of the clone X 7 -2, X 11 -2, and X 13 -1, suggesting that recent proliferation or intra - chromosomal translocation has been occurred within the Xq 21. 3 region during hominid evolution. B. Our phylogenetic tree including Gen. Bank data also shows a similar pattern, indicating that at least two different types of the HERV-W LTR elements were proliferated into the human genomes during hominid evolution. C. HERV-W LTR element were expressed in Human brain.