IDEA Image Display Enhancement and Analysis LINKS Learningbased

IDEA Image Display, Enhancement, and Analysis LINKS: Learning-based multi-source Integratio. N framewor. K for Segmentation of infant brain images Li Wang, Yaozong Gao, Feng Shi, Gang Li, Dinggang Shen Presented by Li Wang 09 -18 -2014 Department of Radiology and BRIC, UNC-Chapel Hill

Content Motivation n Proposed method n Experimental results n Conclusion n Department of Radiology and BRIC, UNC-Chapel Hill
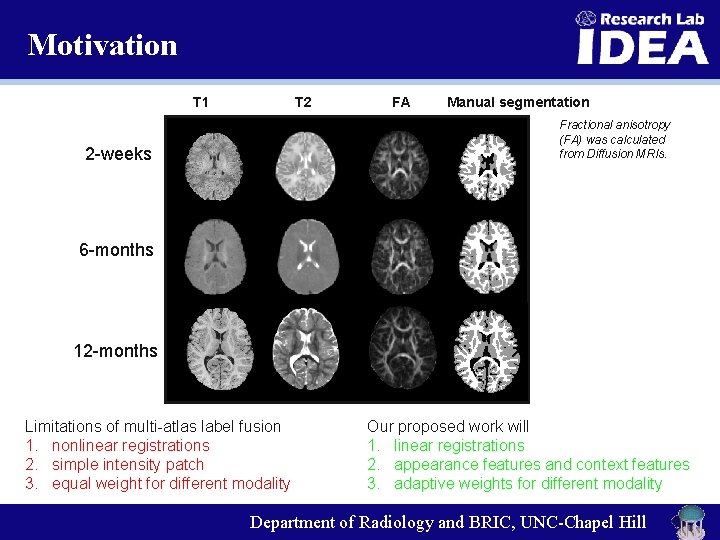
Motivation T 2 T 1 FA Manual segmentation Fractional anisotropy (FA) was calculated from Diffusion MRIs. 2 -weeks 6 -months 12 -months Limitations of multi-atlas label fusion 1. nonlinear registrations 2. simple intensity patch 3. equal weight for different modality Our proposed work will 1. linear registrations 2. appearance features and context features 3. adaptive weights for different modality Department of Radiology and BRIC, UNC-Chapel Hill
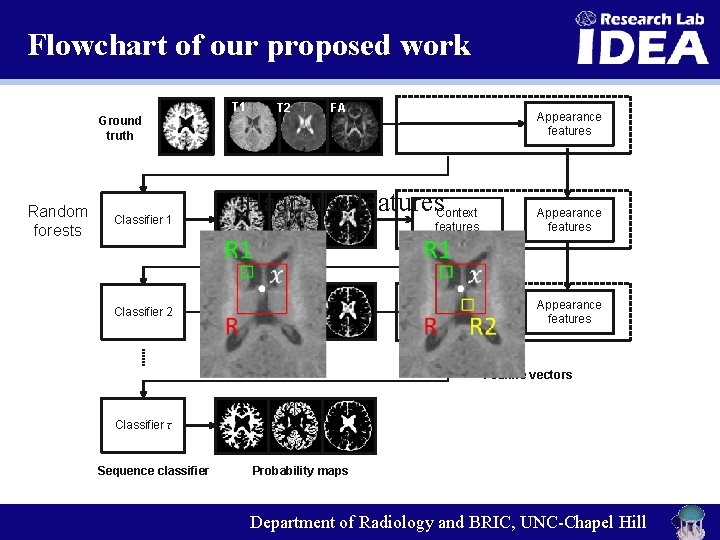
Flowchart of our proposed work T 1 Ground truth Random forests Classifier 1 T 2 FA Appearance features Haar-like features. Context Classifier 2 features Appearance features Context features Appearance features Feature vectors Classifier τ Sequence classifier Probability maps Department of Radiology and BRIC, UNC-Chapel Hill
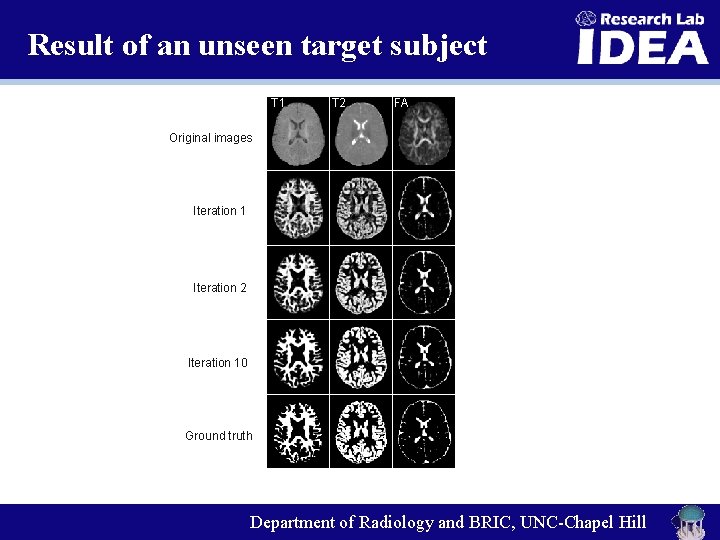
Result of an unseen target subject T 1 T 2 FA Original images Iteration 1 Iteration 2 Iteration 10 Ground truth Department of Radiology and BRIC, UNC-Chapel Hill

Post-processing: Anatomical constraint To deal with the possible artifacts due to independent voxel-wise classification, we use patch-based sparse representation to impose an anatomical constraint [1] into the segmentation. Ground truth of training images Probabilities of training image by the random forest Probabilities of target image by the random forest Without anatomical With anatomical Ground truth 1. Wang, L. , Shi, F. , Gao, Y. , Li, G. , Gilmore, J. H. , Lin, W. , Shen, D. , 2014. Integration of sparse multi-modality representation and anatomical constraint for isointense infant brain MR image segmentation. Neuro. Image 89, 152 -164. Department of Radiology and BRIC, UNC-Chapel Hill

Dataset 1: UNC 119 infants consisting of 26, 22, 23, and 26 subjects at 0 -, 3 -, 6 -, 9 - and 12 -months of age, respectively. § Dataset 2: Neo. Brain. S 12 MICCAI 2012 Challenge. § Dataset 3: SATA MICCAI 2013 Challenge. § Department of Radiology and BRIC, UNC-Chapel Hill
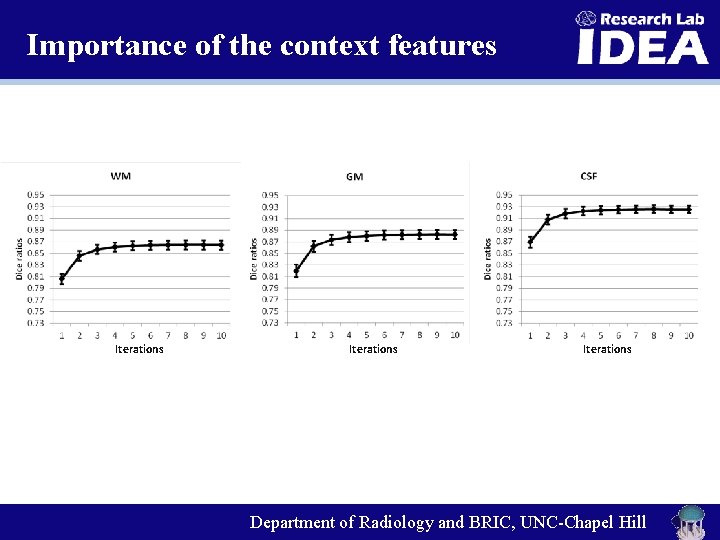
Importance of the context features Iterations Department of Radiology and BRIC, UNC-Chapel Hill
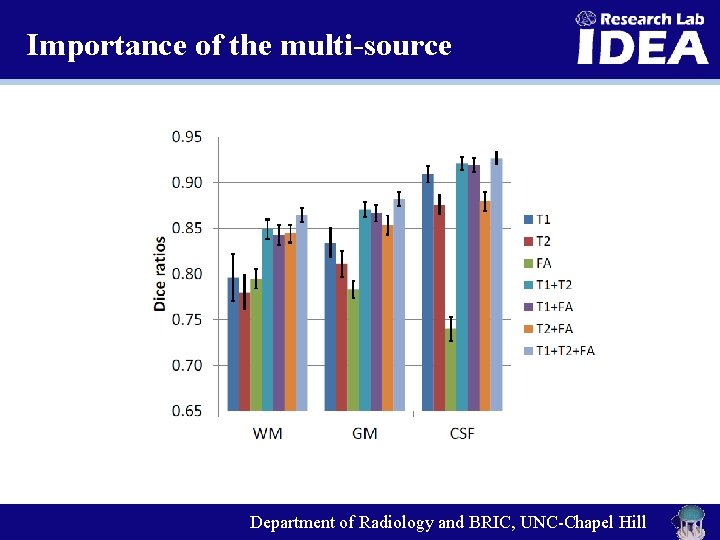
Importance of the multi-source Department of Radiology and BRIC, UNC-Chapel Hill

Dataset 1: UNC 119 infants (a) (b) (c) (d) (e) (f) Majority voting (MV) Nonlocal label fusion [1] Atlas forest [2] Patch-based sparse labeling [3] Proposed 1 (Random forest) Proposed 2 (Random forest + Anatomical constraint) 1. Coupé, P. , Manjón, J. , Fonov, V. , Pruessner, J. , Robles, M. , Collins, D. L. , 2011. Patch-based segmentation using expert priors: Application to hippocampus and ventricle segmentation. Neuro. Image 54, 940 -954. 2. Zikic, D. , Glocker, B. , Criminisi, A. , 2013. Atlas Encoding by Randomized Forests for Efficient Label Propagation. MICCAI 2013, pp. 66 -73. 3. Wang, L. , Shi, F. , Gao, Y. , Li, G. , Gilmore, J. H. , Lin, W. , Shen, D. , 2014. Integration of sparse multi-modality representation and anatomical constraint for isointense infant brain MR image segmentation. Neuro. Image 89, 152 -164. Department of Radiology and BRIC, UNC-Chapel Hill
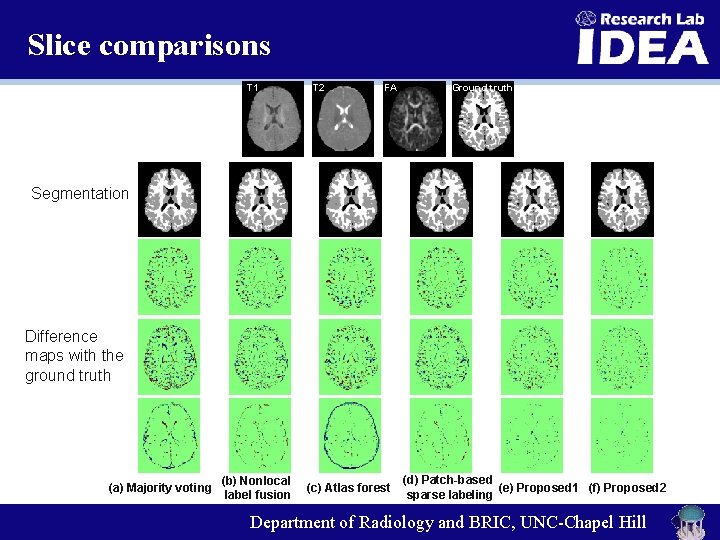
Slice comparisons T 1 T 2 FA Ground truth Segmentation Difference maps with the ground truth (a) Majority voting (b) Nonlocal label fusion (c) Atlas forest (d) Patch-based (e) Proposed 1 (f) Proposed 2 sparse labeling Department of Radiology and BRIC, UNC-Chapel Hill
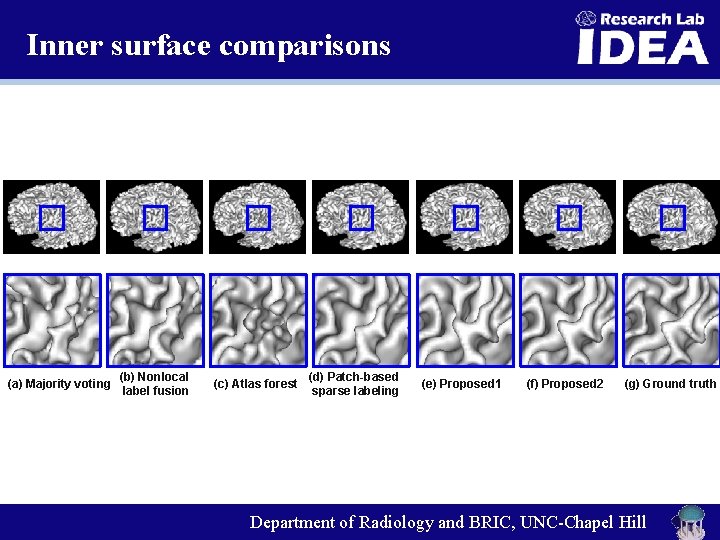
Inner surface comparisons (a) Majority voting (b) Nonlocal label fusion (c) Atlas forest (d) Patch-based sparse labeling (e) Proposed 1 (f) Proposed 2 (g) Ground truth Department of Radiology and BRIC, UNC-Chapel Hill
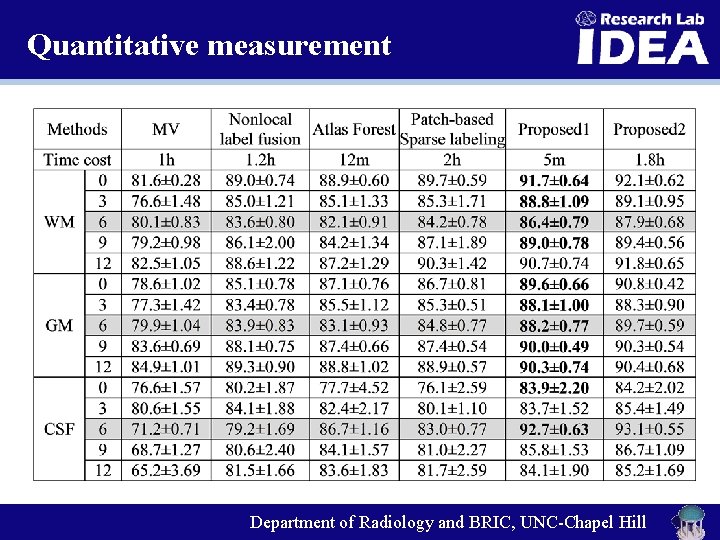
Quantitative measurement Department of Radiology and BRIC, UNC-Chapel Hill

Dataset 2: Neobrain. S 12 MICCAI Challenge Ø 2 training images with the manual segmentations. Ø 3 target images for testing. Department of Radiology and BRIC, UNC-Chapel Hill
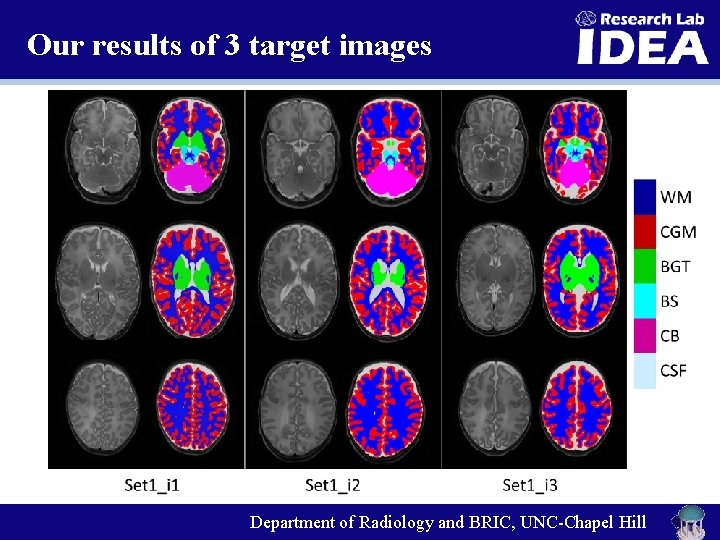
Our results of 3 target images Department of Radiology and BRIC, UNC-Chapel Hill
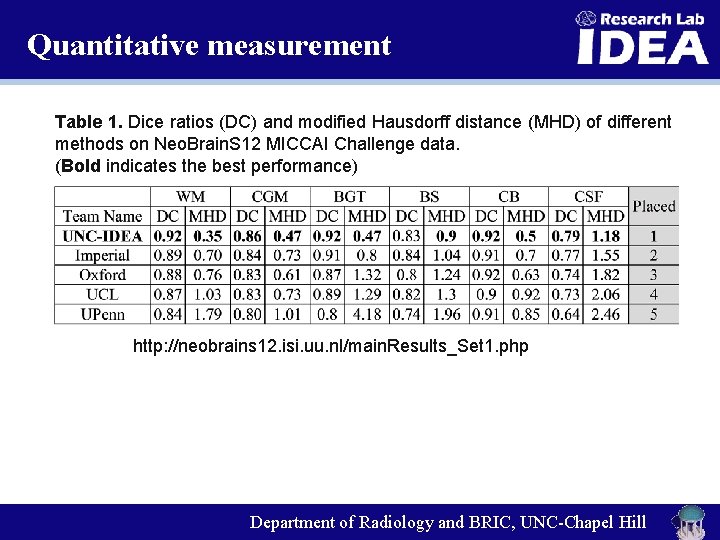
Quantitative measurement Table 1. Dice ratios (DC) and modified Hausdorff distance (MHD) of different methods on Neo. Brain. S 12 MICCAI Challenge data. (Bold indicates the best performance) http: //neobrains 12. isi. uu. nl/main. Results_Set 1. php Department of Radiology and BRIC, UNC-Chapel Hill

Dataset 3: SATA MICCAI 2013 Challenge Ø 35 training images with the 14 ROIs in subcortical regions. Ø 12 target images for testing. Department of Radiology and BRIC, UNC-Chapel Hill
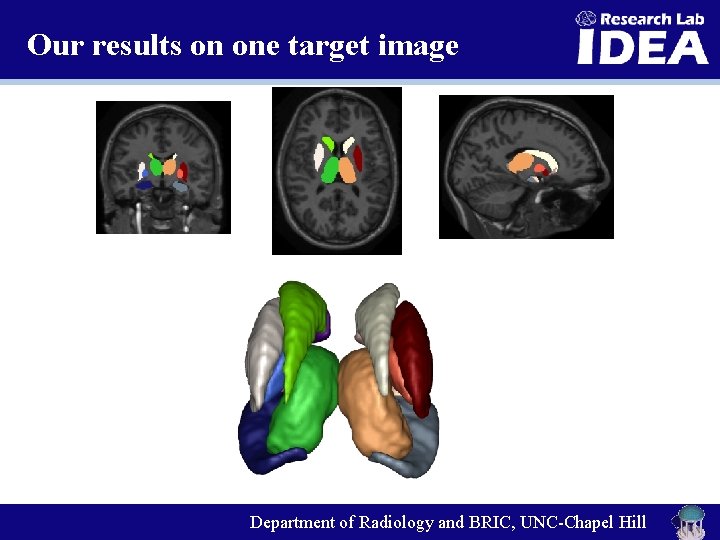
Our results on one target image Department of Radiology and BRIC, UNC-Chapel Hill
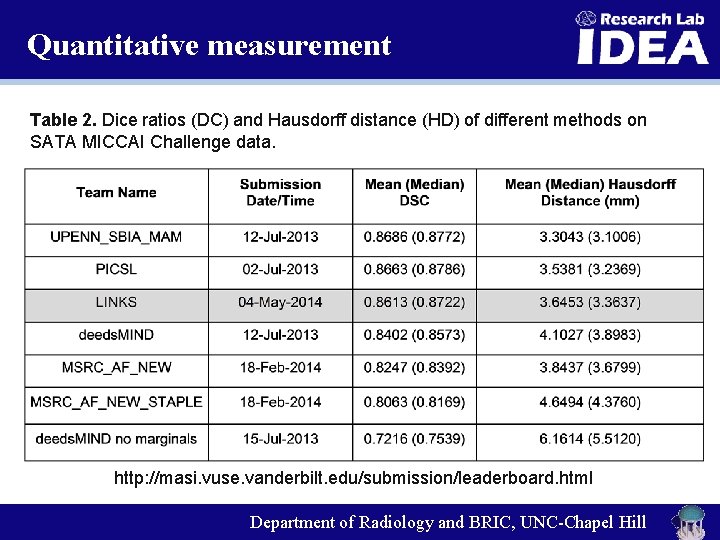
Quantitative measurement Table 2. Dice ratios (DC) and Hausdorff distance (HD) of different methods on SATA MICCAI Challenge data. http: //masi. vuse. vanderbilt. edu/submission/leaderboard. html Department of Radiology and BRIC, UNC-Chapel Hill

Conclusion We have presented a learning-based method (LINKS) to effectively integrate multi-source images and the tentatively estimated tissue probability maps for infant brain image segmentation. n Experimental results on 119 infant subjects and MICCAI grand challenge show that the proposed method achieves better performance than other state-of-the-art automated segmentation methods. n Department of Radiology and BRIC, UNC-Chapel Hill

n Source code can be found: https: //liwang. web. unc. edu n Google scholar Department of Radiology and BRIC, UNC-Chapel Hill
- Slides: 21