I 519 Introduction to Bioinformatics 2011 Sequencing techniques

I 519 Introduction to Bioinformatics, 2011 Sequencing techniques and genome assembly Yuzhen Ye (yye@indiana. edu) School of Informatics & Computing, IUB
Start with reads >read 1 aatgcatgcggctatgctaagctgggatccgatgacaatgcgg ctatgctaatgcggctatgcaagctgggatccgatgactatgctaagctgggatccgatga caatgcggctatgctaatggtcttgggatttaccttggaat >read 2 gctaagctgggatccgatgacaatgcggctatgctaatggtcttgggatttaccttg gaatatgctaatgcggctatgctaagctgggatccgatgacaatgcggctatgctaa tgcatgcggctatgcaagctgggatccgatgactatgctaagctgcggctatgctaatgcg gctatgctaagctgggatccgatgacaatgca >read 3 tgcggctatgctaatgcggctatgcaagctgggatcctgcggctatgctaatggtct tgggatttaccttggaatgctaagctgggatccgatgacaatgcggctatgctaatg gtcttgggatttaccttggaatatgctaatgcggctatgcta ……

What can be done § Assemble the short reads into a genome (hopefully a complete genome) – Assembly problem § Comparative analysis – Whole genome level: whole genome comparison – Individual gene level – Genome variation & SNP § Annotate the genome – What are the genes (gene structure prediction) – What are the functions of the genes

How genome sequences are generated? § Limitation on read length (new sequencers produce even shorter reads than Sanger sequencing machines) § Sequencing of long DNA sequences (a chromosome or a whole genome) relies on sequencing of short segments (carried in cloning vectors) § Two approaches to sequence large pieces of – Chromosome walking / primer walking; progresses through the entire strand, piece by piece – Shotgun sequencing; cut DNA randomly into smaller pieces; with sufficient oversampling (? ), the sequence of the target can be inferred by piecing the sequence reads together into an assembly.

Cloning vectors § Cloning Vectors: DNA vehicles in which a foreign DNA can be inserted; and stay stable § Various types – Cosmid (plasmid, containing 37 -52 kbp of DNA) – BAC (Bacterial Artificial Chromosome; takes in 100 -300 kbp of foreign DNA) – YAC (Yeast Artificial Chromosome)
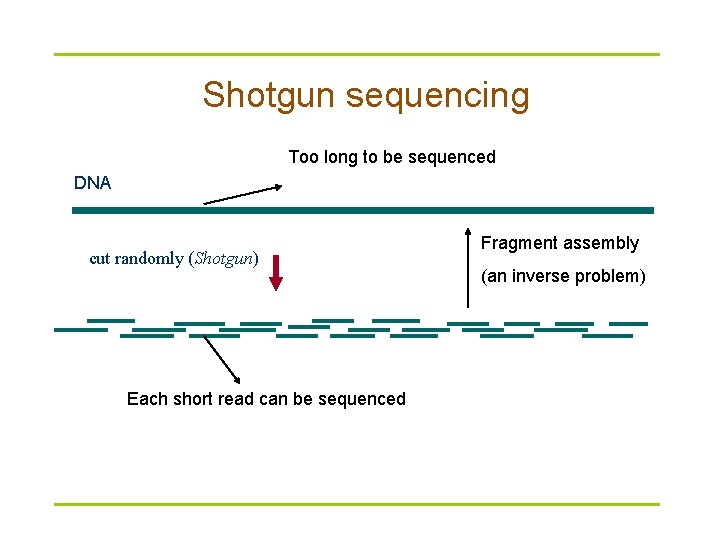
Shotgun sequencing Too long to be sequenced DNA cut randomly (Shotgun) Each short read can be sequenced Fragment assembly (an inverse problem)

Shotgun sequencing: from small viral genomes to larger genomes § Early applications of shotgun approach – small viral genomes (e. g. , lambda virus; 1982) – 30 - to 40 -kbp segments of larger genomes that could be manipulated and amplified in cosmids or other clones (physical mapping) -- hierarchical genome sequencing (divide-and-conquer sequencing) § 1994, Haemophilus influenzae -- whole-genome shotgun (WGS) sequencing – Critical to this accomplishment: use of pairs of reads, called mates, from the ends of 2 -kbp and 16 -kbp inserts randomly sampled from the genome (which used for ordering the contigs) § 2001 whole-genome shotgun sequencing of Human genome

DNA sequencing technology § Sanger sequencing – The main method for sequencing DNA for the past thirty years! § 2 nd generation sequencing techniques (next generation sequencing) – Differ from Sanger sequencing in their basic chemistry – Massively increased throughput – Smaller DNA concentration – 454 pyrosequencing, Ilumina/Solexa, SOLi. D § 3 rd generation? (single-molecule)

DNA sequencing: history Sanger method (1977): labeled dd. NTPs terminate DNA copying at random points. Gilbert method (1977): chemical method to cleave DNA at specific points (G, G+A, T+C, C). Both methods generate labeled fragments of varying lengths that are further electrophoresed (electrophoretic separation)
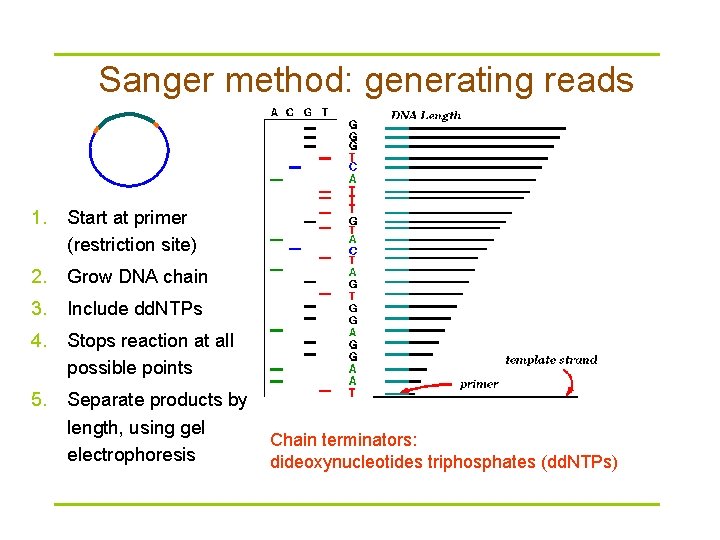
Sanger method: generating reads 1. Start at primer (restriction site) 2. Grow DNA chain 3. Include dd. NTPs 4. Stops reaction at all possible points 5. Separate products by length, using gel electrophoresis Chain terminators: dideoxynucleotides triphosphates (dd. NTPs)
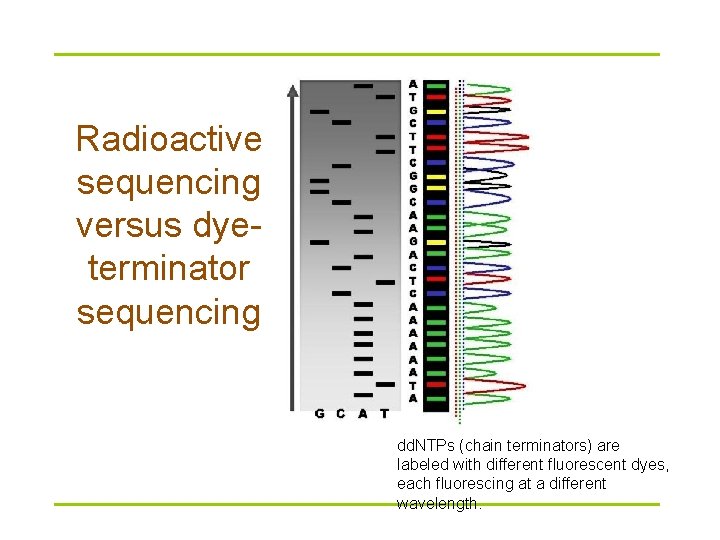
Radioactive sequencing versus dyeterminator sequencing dd. NTPs (chain terminators) are labeled with different fluorescent dyes, each fluorescing at a different wavelength.
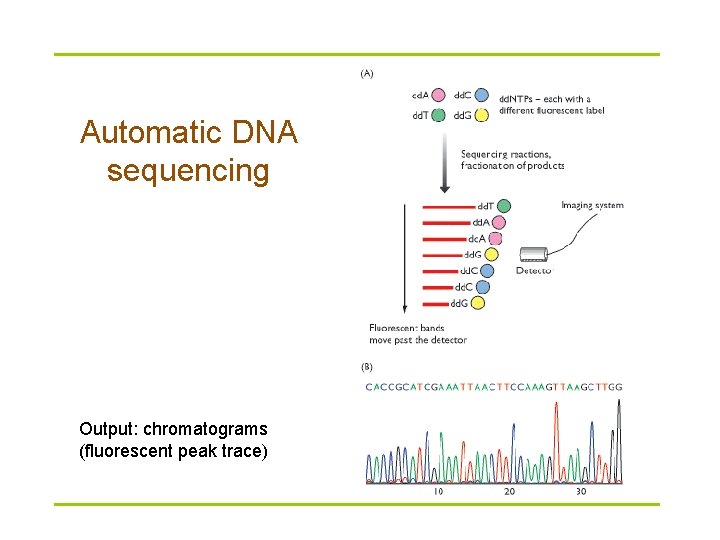
Automatic DNA sequencing Output: chromatograms (fluorescent peak trace)
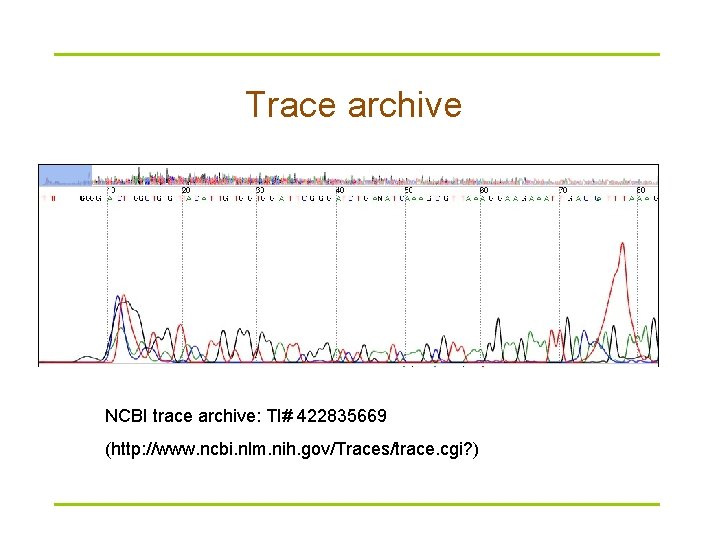
Trace archive NCBI trace archive: TI# 422835669 (http: //www. ncbi. nlm. nih. gov/Traces/trace. cgi? )

New sequencing techniques § Next Generation Sequencing (NGS) (Second Generation) – Pyrosequencing – Illumina – SOLi. D § Third generation sequencing – single-molecule sequencing technologies § NHGRI funds development of third generation DNA sequencing technologies – “More than $18 million in grants to spur the development of a third generation of DNA sequencing technologies was announced today by the National Human Genome Research Institute (NHGRI). …The cost to sequence a human genome has now dipped below $40, 000. Ultimately, NHGRI's vision is to cut the cost of whole-genome sequencing of an individual's genome to $1, 000 or less, which will enable sequencing to be a part of routine medical care. . ” – http: //www. nih. gov/news/health/sep 2010/nhgri-14. htm

Next-generation sequencing transforms today's biology 454 sequencer Ref: Nature Methods - 5, 16 - 18 (2008) Sanger sequencers

Next-generation sequencing transforms today's biology § § Genome re-sequencing Metagenomics Transcriptomics (RNA-seq) Personal genomics ($1000 for sequencing a person’s genome)
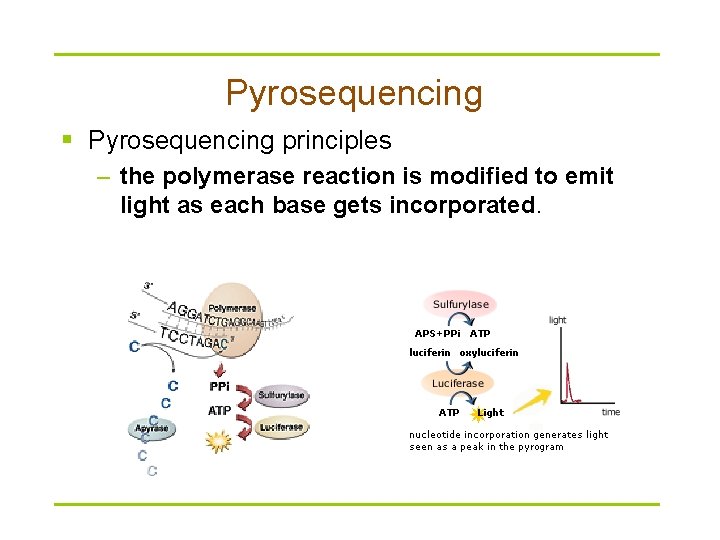
Pyrosequencing § Pyrosequencing principles – the polymerase reaction is modified to emit light as each base gets incorporated.
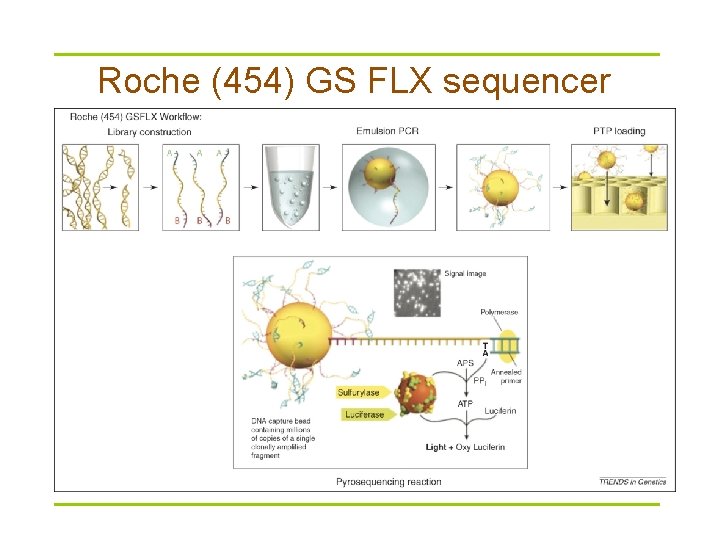
Roche (454) GS FLX sequencer

Solexa/Illumina sequencing § Ultrahigh-throughput sequencing § Keys – attachment of randomly fragmented genomic DNA to a planar, optically transparent surface – solid phase amplification to create an ultra-high density sequencing flow cell with > 10 million clusters, each containing ~1, 000 copies of template per sq. cm. § Short reads § Used for gene expression, small RNA discovery etc
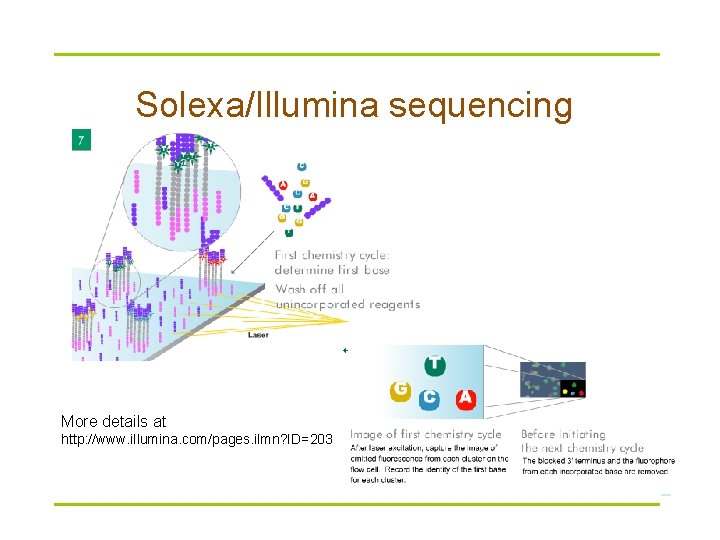
Solexa/Illumina sequencing More details at http: //www. illumina. com/pages. ilmn? ID=203

Applied Biosystems SOLi. D sequencer § Commercial release in October 2007 § Sequencing by Oligo Ligation and Detection § ~5 days to run / produces 3 -4 Gb § The chemistry is based on template-directed ligation of short, “dinucleotide-encoding”, 8 mer oligonucleotides. Dinucleotide-encoding permits discrimination of SNP’s from most chemistry and imaging errors, and subsequent in silico correction of those errors. Ref: http: //appliedbiosystems. cnpg. com/Video/flat. Files/699/index. aspx

Comparison of new sequencing techniques Applied Biosysems 3730 xl 454 GS FLX Pyrosequencer Solexa 1 G Genome Analyzer Applied Biosystems 1 G SOLi. D Analyzer 1 -2 Mbp per day/machine 100 Mbp per day/machine 800 Mbp per run/machine 1200 Mbp per run/machine 600 -900 bp 200 -300 bp 25 -40 bp 25 -30 Increased!! Mate pair No Mate pair Yes now! Libraries No No (“The new science of metagenomics” Table 4 -2) Libraries
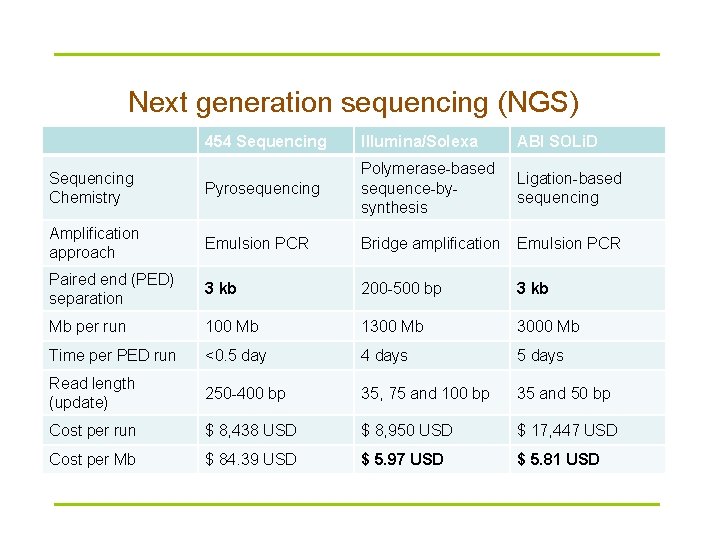
Next generation sequencing (NGS) 454 Sequencing Illumina/Solexa ABI SOLi. D techniques Pyrosequencing Polymerase-based sequence-bysynthesis Ligation-based sequencing Amplification approach Emulsion PCR Bridge amplification Emulsion PCR Paired end (PED) separation 3 kb 200 -500 bp 3 kb Mb per run 100 Mb 1300 Mb 3000 Mb Time per PED run <0. 5 day 4 days 5 days Read length (update) 250 -400 bp 35, 75 and 100 bp 35 and 50 bp Cost per run $ 8, 438 USD $ 8, 950 USD $ 17, 447 USD Cost per Mb $ 84. 39 USD $ 5. 97 USD $ 5. 81 USD Sequencing Chemistry
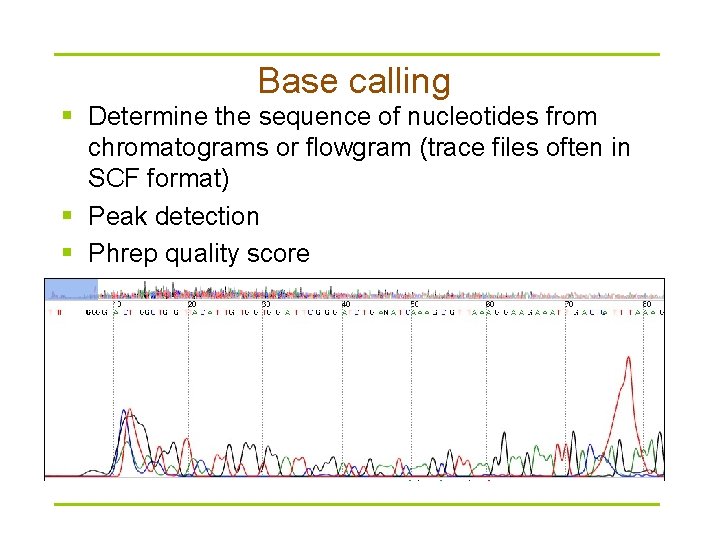
Base calling § Determine the sequence of nucleotides from chromatograms or flowgram (trace files often in SCF format) § Peak detection § Phrep quality score Q = -10 log 10(Pe)

Phrep quality score Phred Quality Score Probability of incorrect base call Base call accuracy 10 1/10 90% 20 1/100 99% (for high values the two scores are asymptotically equal)
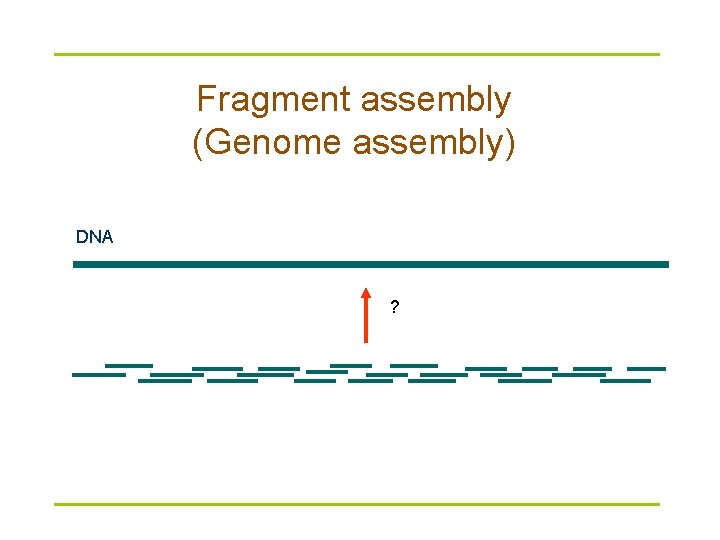
Fragment assembly (Genome assembly) DNA ?

Assembly § Comparative assembly – comparative (re-sequencing) approaches that use the sequence of a closely related organism as a guide during the assembly process. § De novo assembly – reconstructing genomes that are not similar to any organisms previously sequenced – proven to be difficult, falling within a class of problems (NP-hard) – main strategies: greedy, overlap-layoutconsensus, and Eulerian
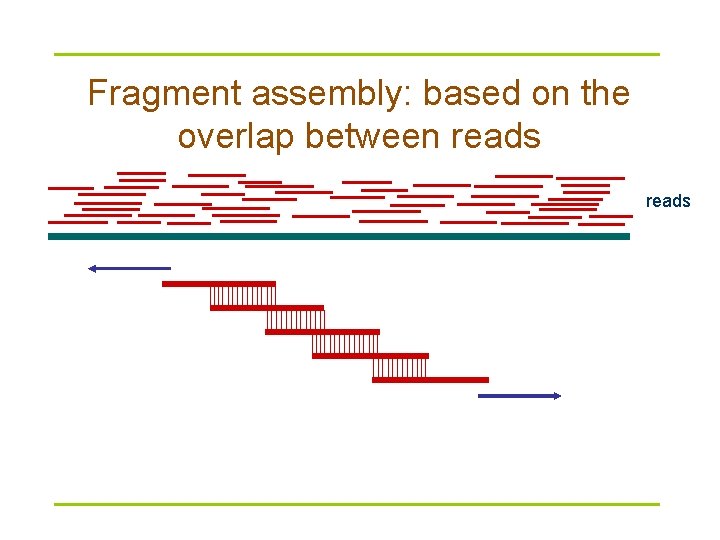
Fragment assembly: based on the overlap between reads
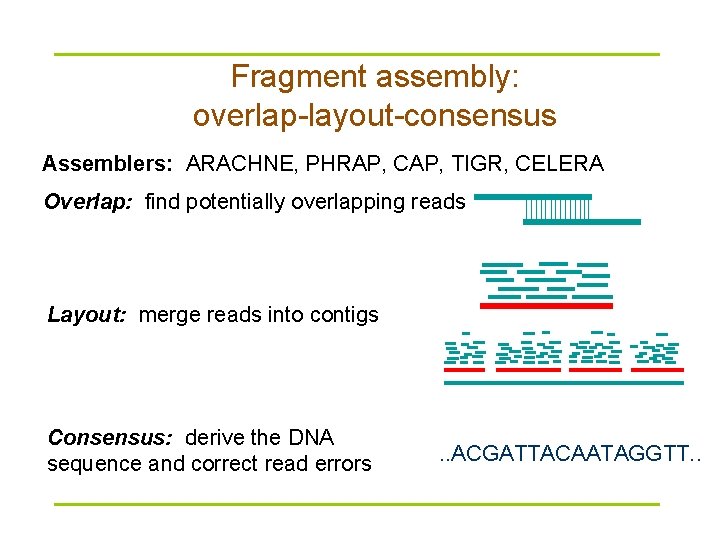
Fragment assembly: overlap-layout-consensus Assemblers: ARACHNE, PHRAP, CAP, TIGR, CELERA Overlap: find potentially overlapping reads Layout: merge reads into contigs Consensus: derive the DNA sequence and correct read errors . . ACGATTACAATAGGTT. .

Overlap § Find the best match between the suffix of one read and the prefix of another § Due to sequencing errors, need to use dynamic programming to find the optimal overlap alignment § Apply a filtration method to filter out pairs of fragments that do not share a significantly long common substring
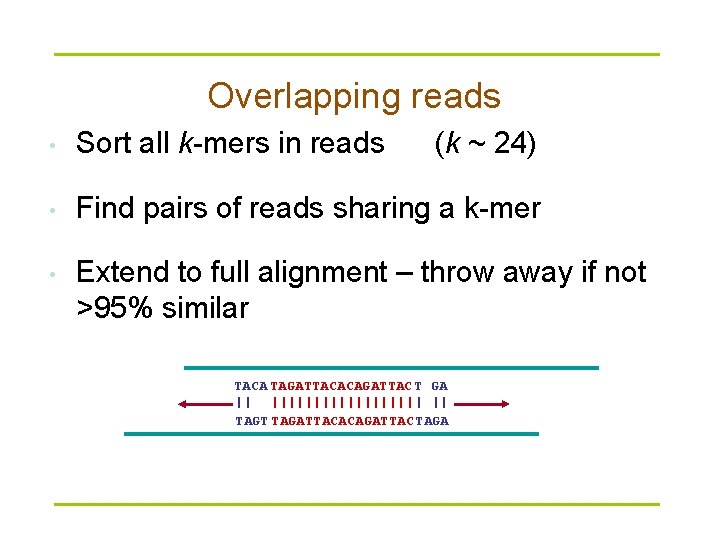
Overlapping reads • Sort all k-mers in reads (k ~ 24) • Find pairs of reads sharing a k-mer • Extend to full alignment – throw away if not >95% similar TACA TAGATTACACAGATTAC T GA || ||||||||| | || TAGT TAGATTACACAGATTAC TAGA
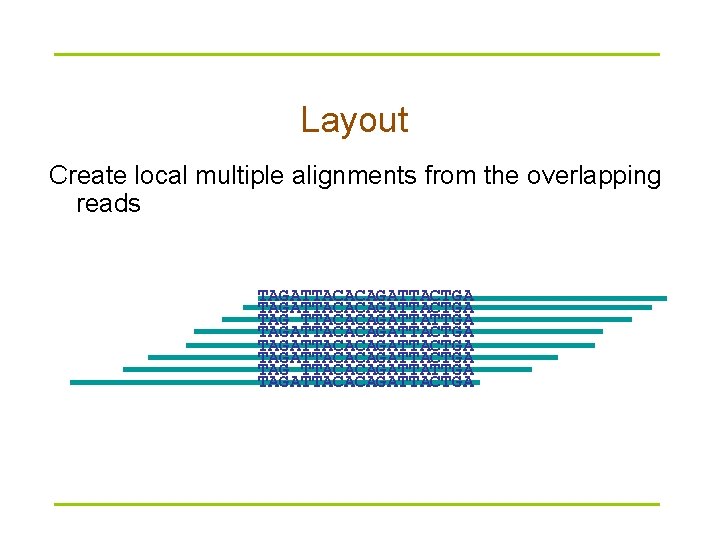
Layout Create local multiple alignments from the overlapping reads TAGATTACACAGATTACTGA TAG TTACACAGATTATTGA TAGATTACACAGATTACTGA

Derive consensus sequence TAGATTACACAGATTACTGA TTGATGGCGTAA CTA TAGATTACACAGATTACTGACTTGATGGCGTAAACTA TAG TTACACAGATTATTGACTTCATGGCGTAA CTA TAGATTACACAGATTACTGACTTGATGGGGTAA CTA TAGATTACACAGATTACTGACTTGATGGCGTAA CTA Derive multiple alignment from pairwise read alignments Derive each consensus base by weighted voting

Consensus § A consensus sequence is derived from a profile of the assembled fragments § A sufficient number of reads are required to ensure a statistically significant consensus. § Reading errors are corrected
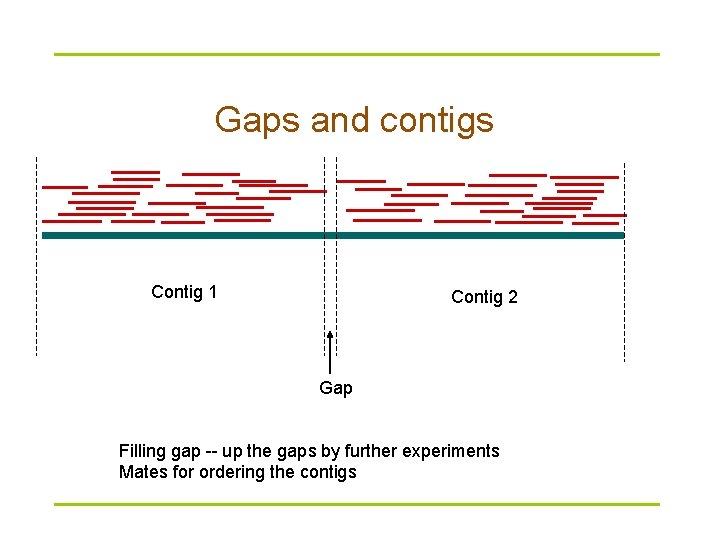
Gaps and contigs Contig 1 Contig 2 Gap Filling gap -- up the gaps by further experiments Mates for ordering the contigs

Read coverage C Assuming uniform distribution of reads: Length of genomic segment: L Number of reads: n Length of each read: l Coverage l = n l / L How much coverage is enough (or what is sufficient oversampling)? Lander-Waterman model: P(x) = (lx * e-l ) / x! P(x=0) = e -l where l is coverage
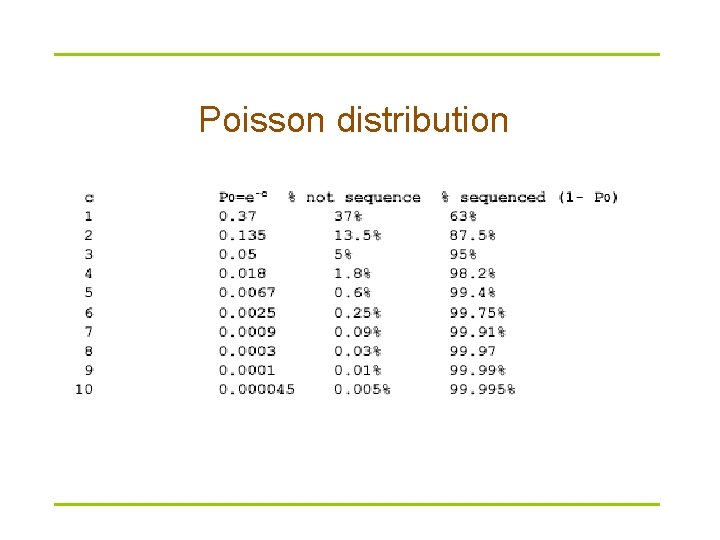
Poisson distribution
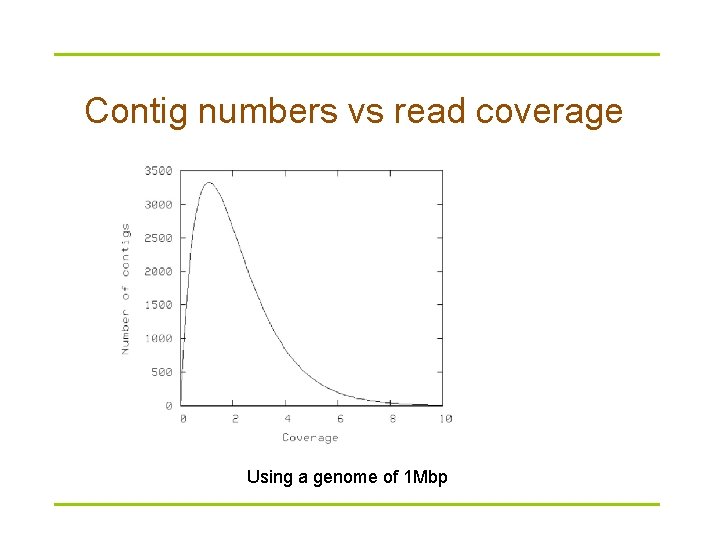
Contig numbers vs read coverage Using a genome of 1 Mbp
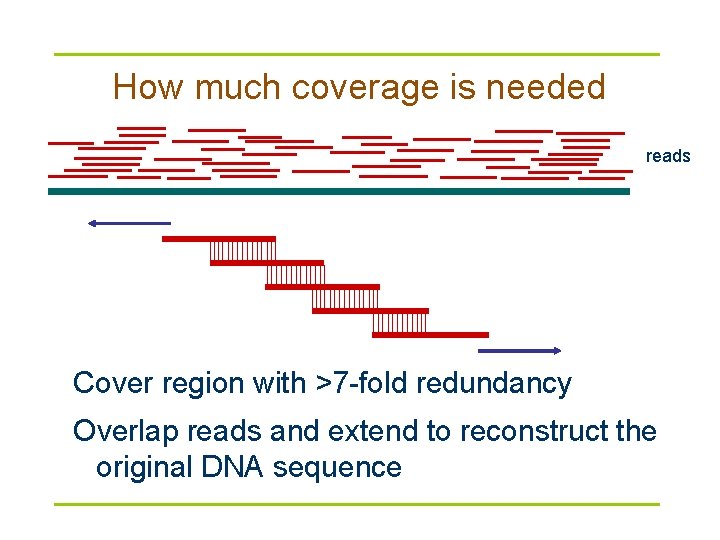
How much coverage is needed reads Cover region with >7 -fold redundancy Overlap reads and extend to reconstruct the original DNA sequence
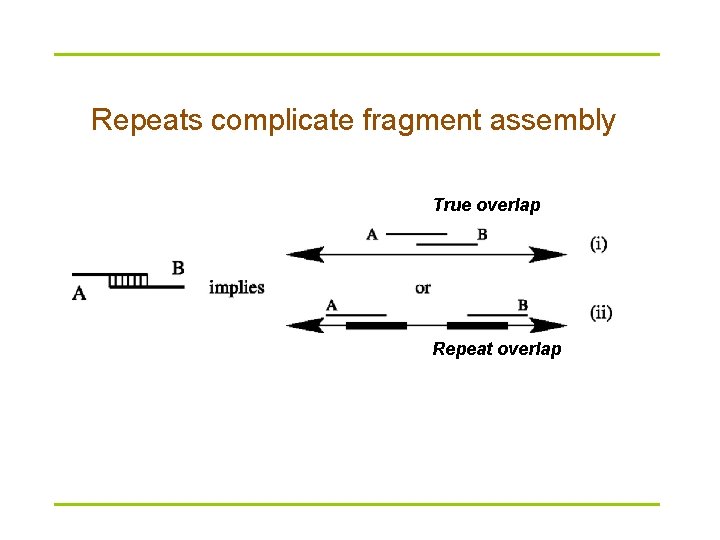
Repeats complicate fragment assembly True overlap Repeat overlap
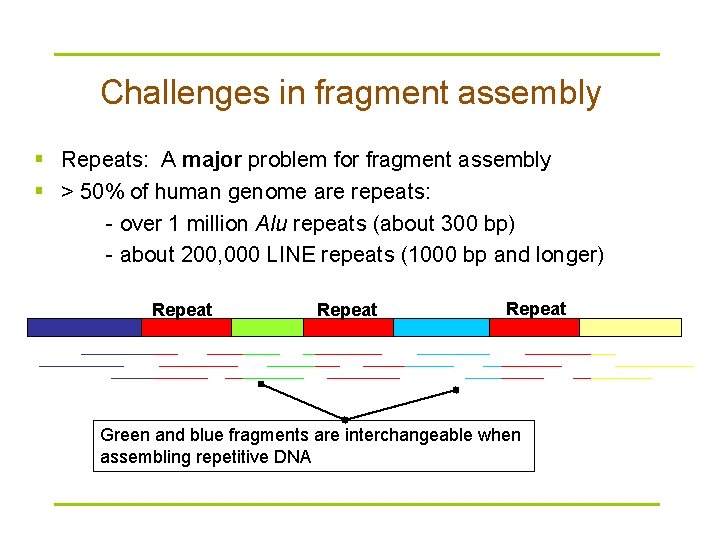
Challenges in fragment assembly § Repeats: A major problem for fragment assembly § > 50% of human genome are repeats: - over 1 million Alu repeats (about 300 bp) - about 200, 000 LINE repeats (1000 bp and longer) Repeat Green and blue fragments are interchangeable when assembling repetitive DNA

Repeat types § Low-Complexity DNA § Microsatellite repeats § (e. g. ATATACATA…) (a 1…ak)N where k ~ 3 -6 (e. g. CAGCAGTAGCAGCACCAG) Transposons/retrotransposons – SINE Short Interspersed Nuclear Elements (e. g. , Alu: ~300 bp long, 106 copies) – LINE Long Interspersed Nuclear Elements ~500 - 5, 000 bp long, 200, 000 copies – LTR retroposons Long Terminal Repeats (~700 bp) at each end genes duplicate & then diverge § Gene Families § Segmental duplications ~very long, very similar copies

Celera assembler § “The key to not being confused by repeats is the exploitation of mate pair information to circumnavigate and to fill them” § A mate pair are two reads from the same clone -we know the distance between the two reads Myers et al. 2000 “A Whole-Genome Assembly of Drosophila”. Science, 287: 2196 - 2204
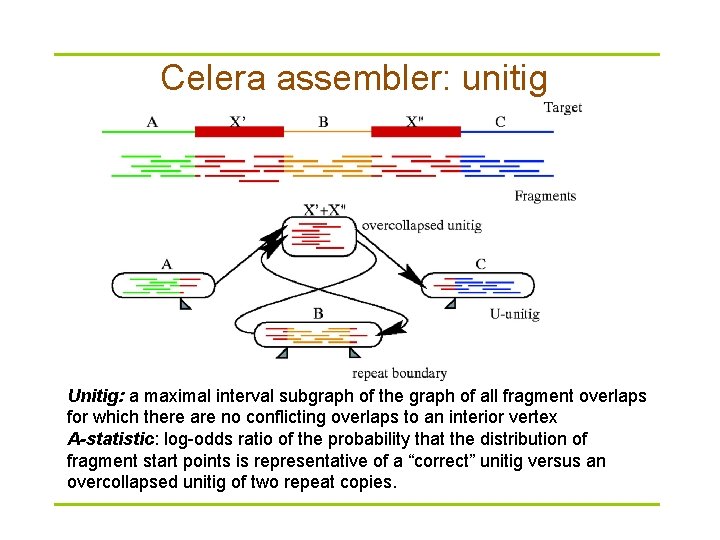
Celera assembler: unitig Unitig: a maximal interval subgraph of the graph of all fragment overlaps for which there are no conflicting overlaps to an interior vertex A-statistic: log-odds ratio of the probability that the distribution of fragment start points is representative of a “correct” unitig versus an overcollapsed unitig of two repeat copies.
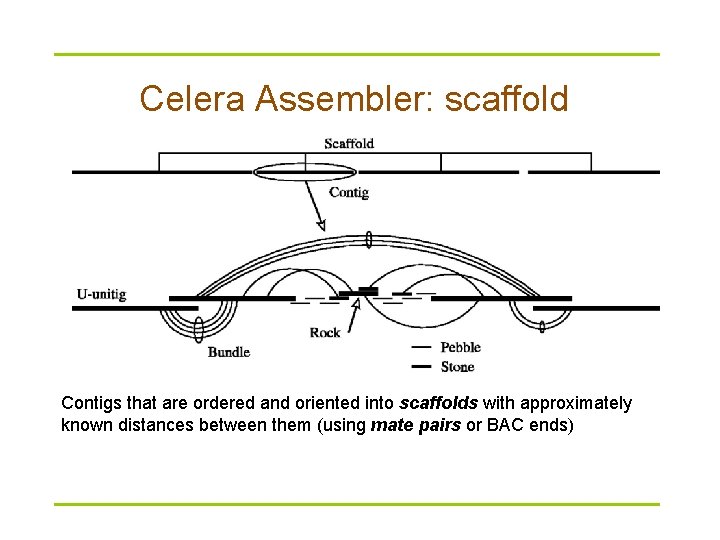
Celera Assembler: scaffold Contigs that are ordered and oriented into scaffolds with approximately known distances between them (using mate pairs or BAC ends)
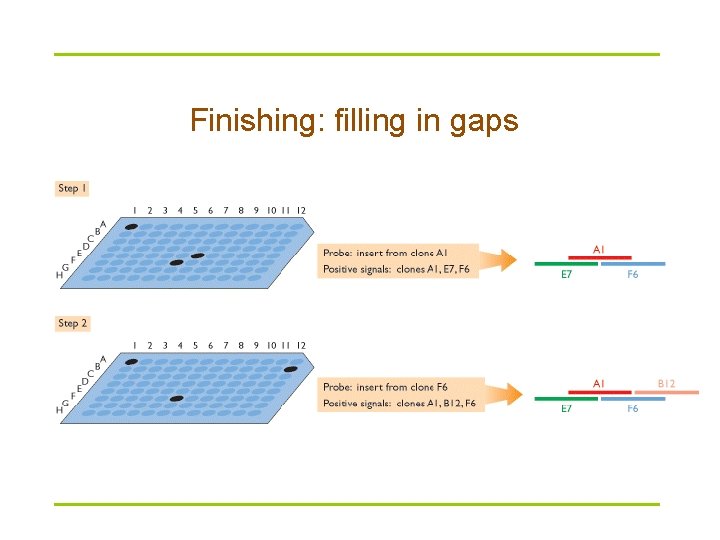
Finishing: filling in gaps

Human genome § 2001 Two assemblies of initial human genome sequences published – International Human Genome project (Hierachical sequencing; BACshotgun) – Celera Genomics: WGS approach; § Initial impact of the sequencing of the human genome (Nature 470: 187– 197, 2011)
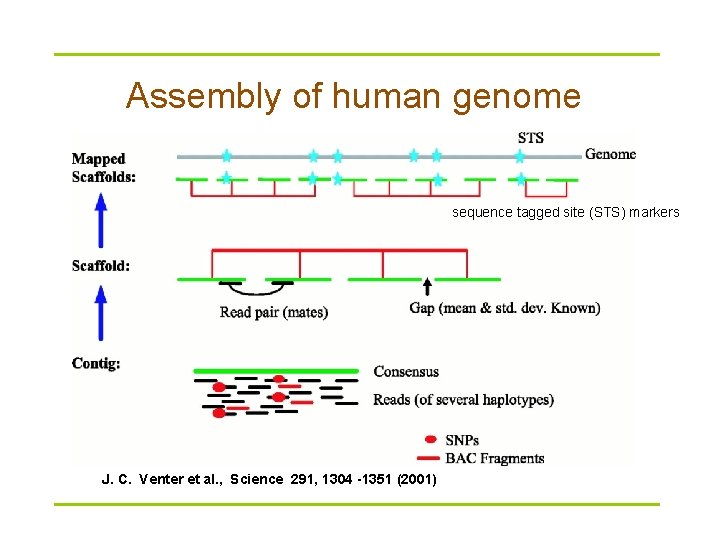
Assembly of human genome sequence tagged site (STS) markers J. C. Venter et al. , Science 291, 1304 -1351 (2001)
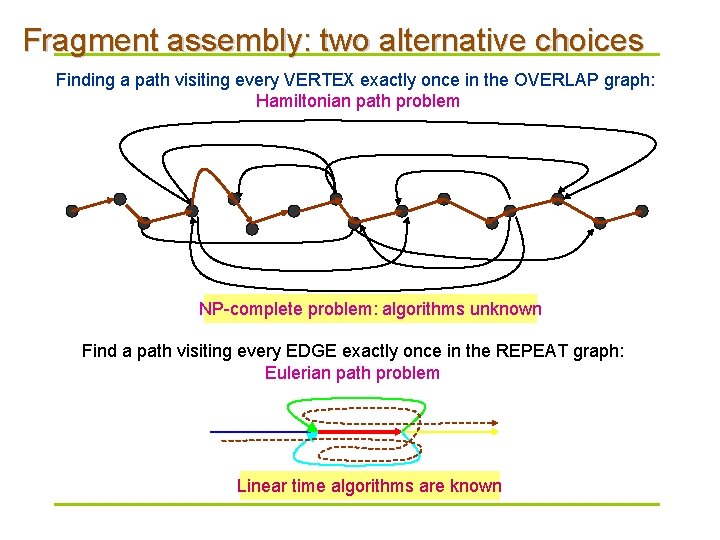
Fragment assembly: two alternative choices Finding a path visiting every VERTEX exactly once in the OVERLAP graph: Hamiltonian path problem NP-complete problem: algorithms unknown Find a path visiting every EDGE exactly once in the REPEAT graph: Eulerian path problem Linear time algorithms are known
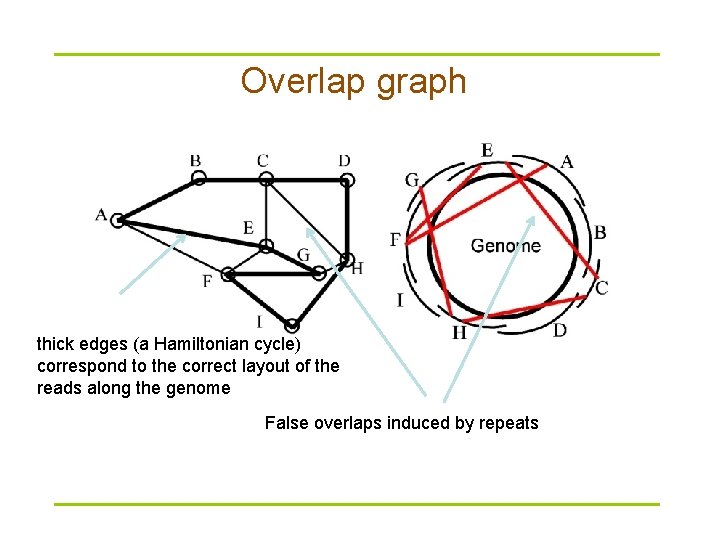
Overlap graph thick edges (a Hamiltonian cycle) correspond to the correct layout of the reads along the genome False overlaps induced by repeats
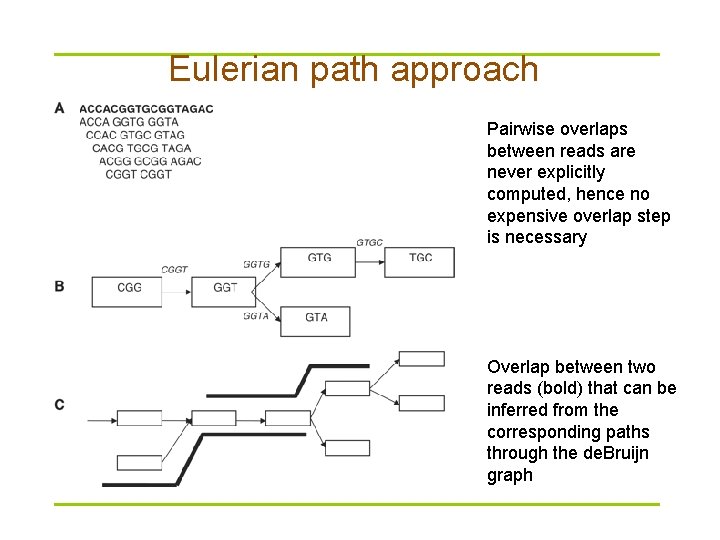
Eulerian path approach Pairwise overlaps between reads are never explicitly computed, hence no expensive overlap step is necessary Overlap between two reads (bold) that can be inferred from the corresponding paths through the de. Bruijn graph
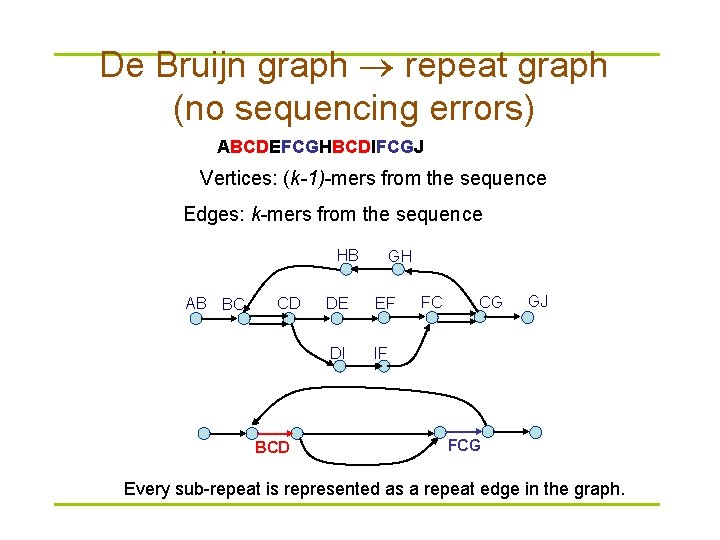
De Bruijn graph repeat graph (no sequencing errors) ABCDEFCGHBCDIFCGJ Vertices: (k-1)-mers from the sequence Edges: k-mers from the sequence HB AB BC CD BCD GH DE EF DI IF FC CG GJ FCG Every sub-repeat is represented as a repeat edge in the graph.

Repeat graph 8328 140 628 1185 2905 628 1185 381 140 628 161442 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 A-Bruijn graph repeat graph Removing bulges and whirls repeat graph Pevzner, Tang and Waterman. “A New Approach to Fragment Assembly in DNA Sequencing”. RECOMB 01
Genome assembly viewer Eagle. View

Assembly quality metrics § Number of contigs, the longest contig § N 50, defined as the contig length such that using equal or longer contigs produces half the bases of the genome (or all the contigs). – sorting all contigs from largest to smallest – contig sizes: 2 M, 1 M, 0. 5 M, 0. 3 M, 0. 2 M, … 500 bp with total bases = 4 M, then N 50 = 0. 2 M

Genome assembly reborn § Genome assembly reborn: recent computational challenges (Briefings in Bioinformatics 2009 10(4): 354 -366) § Hybrid assembler (? )

Sequencing wars § “Ion Torrent’s Fast and Cheap DNA Sequencer Catches On, Even as Biologists Tighten Belts” – semiconductor-based and almost works like a p. H meter in some respects; Personal Genome Machine in December 2010 – Jonathan Rothberg founded 454 Life Sciences, sold to Roche in 2007 – Carlsbad, CA-based Life Technologies § Sequencing Wars—The Third Generation
- Slides: 57