I 519 Introduction to Bioinformatics 2011 Microarray clustering

I 519 Introduction to Bioinformatics, 2011 Microarray & clustering algorithms Yuzhen Ye (yye@indiana. edu) School of Informatics & Computing, IUB

Outline § Microarrays § Clustering algorithms – Hierarchical Clustering – K-Means Clustering

Microarrays and expression analysis § Microarrays allow biologists to infer gene function even when sequence similarity alone is insufficient to infer function. § Microarrays measure the activity (expression level) of the genes under varying conditions/time points § Expression level is estimated by measuring the amount of m. RNA for that particular gene – A gene is active if it is being transcribed – More m. RNA usually indicates more gene activity

Steps in microarray experiment § Experimental Design § Signal Extraction – Image Analysis – Normalization: remove the artifacts across arrays § Data Analysis – Selection of Genes differentially expressed – Clustering and classification

Microarray experiments § Produce c. DNA from m. RNA (DNA is more stable) § Attach phosphor to c. DNA to see when a particular gene is expressed § Different color phosphors are available to compare many samples at once § Hybridize c. DNA over the microarray § Scan the microarray with a phosphor-illuminating laser § Illumination reveals transcribed genes § Scan microarray multiple times for the different color phosphor’s
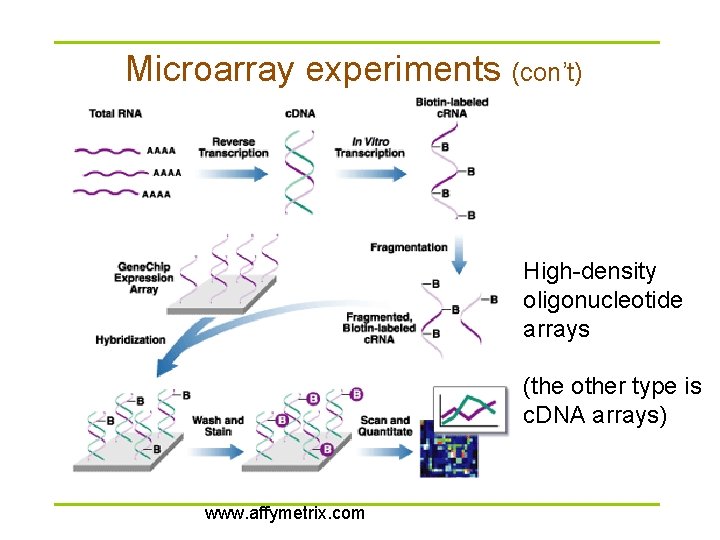
Microarray experiments (con’t) High-density oligonucleotide arrays (the other type is c. DNA arrays) www. affymetrix. com

Microarray experiment design § Type I: (n = 2) – How is this gene expressed in target 1 as compared to target 2? (e. g. , treated versus untreated; disease versus normal) – Which genes show up/down regulation between the two targets? § Type II: (n > 2) – How does the expression of gene A vary over time, tissues, or treatments? – Do any of the expression profiles exhibit similar patterns of expression?
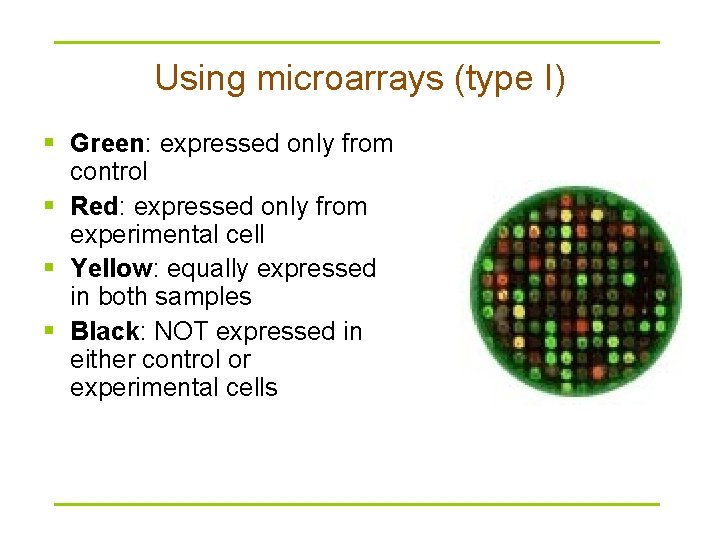
Using microarrays (type I) § Green: expressed only from control § Red: expressed only from experimental cell § Yellow: equally expressed in both samples § Black: NOT expressed in either control or experimental cells
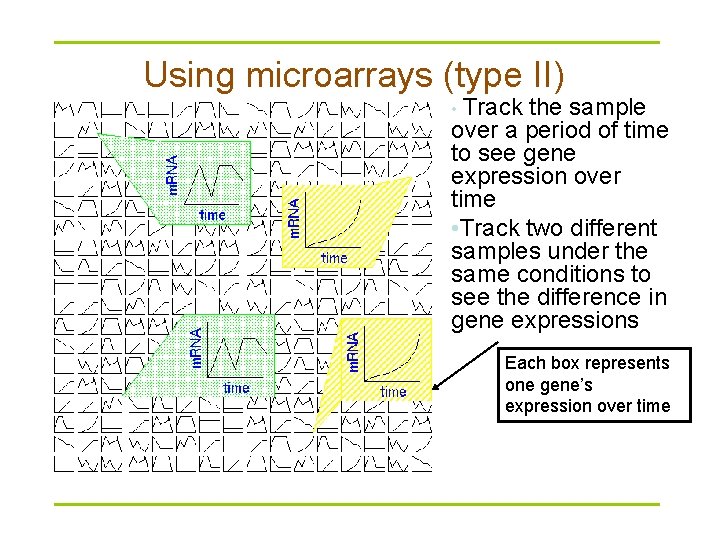
Using microarrays (type II) • Track the sample over a period of time to see gene expression over time • Track two different samples under the same conditions to see the difference in gene expressions Each box represents one gene’s expression over time

Computational & statistical problems § Image and data processing – Normalization – Noise reduction § Data analysis – Clustering – Classification – Network inference
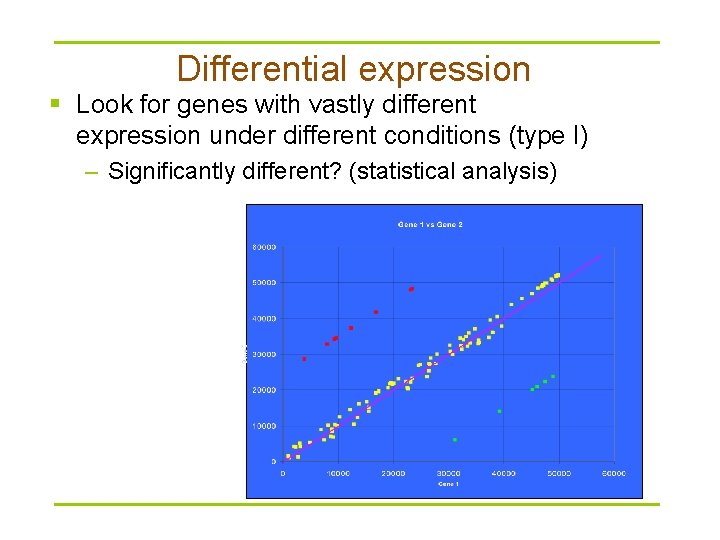
Differential expression § Look for genes with vastly different expression under different conditions (type I) – Significantly different? (statistical analysis)

Statistical considerations are essential for analysis of microarray data § Data quality & background substraction – Poorer-quality or uninformative data should be removed § Data normalization – To remove uneven variation between two labels (e. g. . cy 5 vs cy 3) in one slide or between slides – An observed intensity is the sum of contributions from variables, such as slide-to-slide variation, dye variation, etc. – Lowess normalization (data within a small window of expression values are fitted to a straight line by linear regression) § Differential expression – Analysis of variance (ANOVA) – T-test
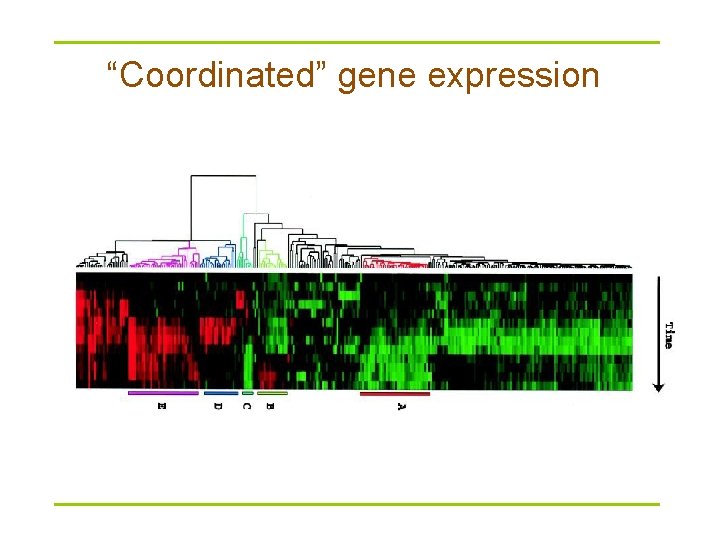
“Coordinated” gene expression

Time series microarray data § Microarray data are usually transformed into an intensity matrix (below) § The intensity matrix allows biologists to make correlations between different genes (even if they are dissimilar) and to understand how genes functions might be related § Clustering comes into play Intensity (expression level) of gene at measured time Time: Time X Time Y Time Z Gene 1 10 8 10 Gene 2 10 0 9 Gene 3 4 8. 6 3 Gene 4 7 8 3 Gene 5 1 2 3

Applications of clustering § Viewing and analyzing vast amounts of biological data as a whole set can be perplexing § It is easier to interpret the data if they are partitioned into clusters combining similar data points.

Clustering of microarray data § Plot each datum as a point in N-dimensional space § Make a distance matrix for the distance between every two gene points in the N-dimensional space § Genes with a small distance share the same expression characteristics and might be functionally related or similar. § Clustering reveal groups of functionally related genes
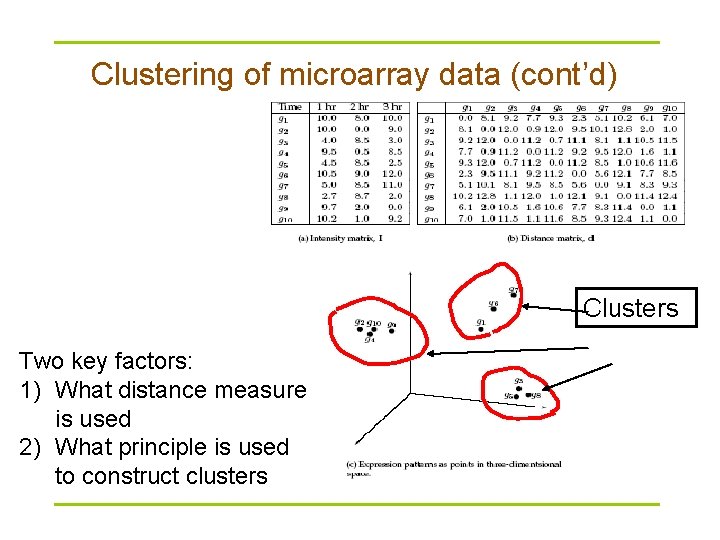
Clustering of microarray data (cont’d) Clusters Two key factors: 1) What distance measure is used 2) What principle is used to construct clusters

Distance measure § Correlation coefficient (between two variables X and Y) – Pearson correlation coefficient (sensitive to outliers) (with values range from -1 to 1; value of 1 implies that a linear equation describes the relationship between X and Y perfectly, with all data points lying on a line for which Y increases as X increases) covariance – Spearman correlation coefficient (nonparametric version; use the ranks) § Absolute value of correlation coefficient, |r| § Euclidean distance

Homogeneity and separation principles § § § Homogeneity: Elements within a cluster are close to each other Separation: Elements in different clusters are further apart from each other …clustering is not an easy task! Given these points, a clustering algorithm might make two distinct clusters as follows
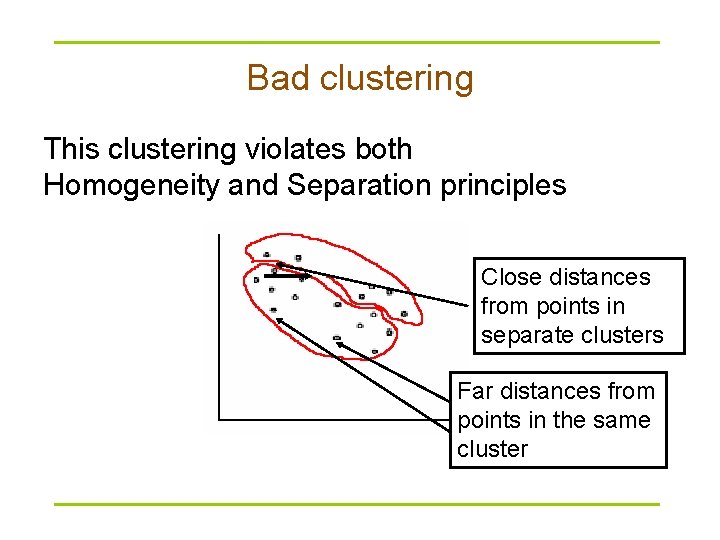
Bad clustering This clustering violates both Homogeneity and Separation principles Close distances from points in separate clusters Far distances from points in the same cluster
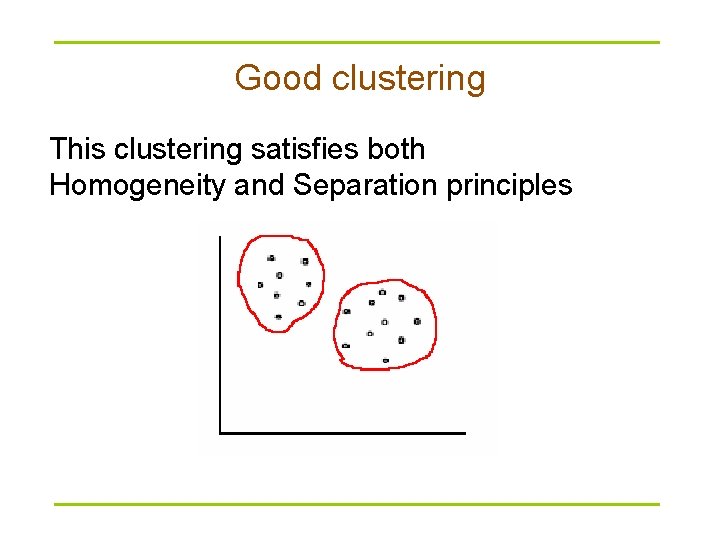
Good clustering This clustering satisfies both Homogeneity and Separation principles

Clustering techniques § Agglomerative: Start with every element in its own cluster, and iteratively join clusters together § Divisive: Start with one cluster and iteratively divide it into smaller clusters § Hierarchical: Organize elements into a tree, leaves represent genes and the length of the pathes between leaves represents the distances between genes. Similar genes lie within the same subtrees
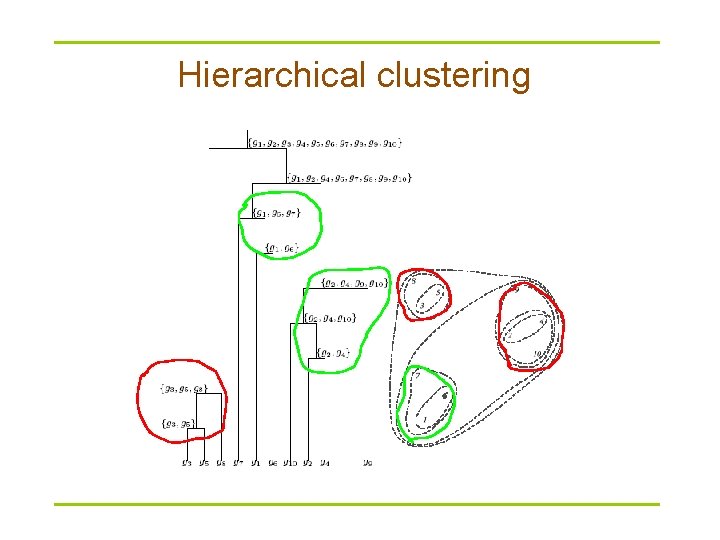
Hierarchical clustering
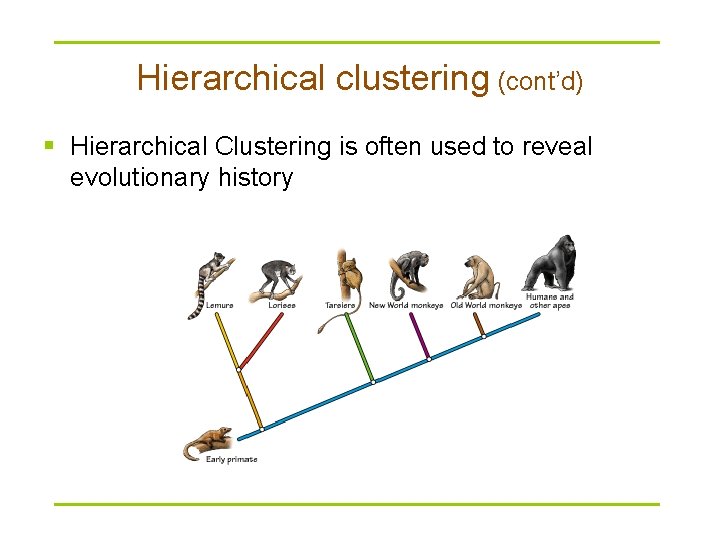
Hierarchical clustering (cont’d) § Hierarchical Clustering is often used to reveal evolutionary history

Hierarchical clustering algorithm 1. Hierarchical Clustering (d , n) 2. Form n clusters each with one element 3. Construct a graph T by assigning one vertex to each cluster 4. while there is more than one cluster 5. Find the two closest clusters C 1 and C 2 6. Merge C 1 and C 2 into new cluster C with |C 1| +|C 2| elements 7. Compute distance from C to all other clusters 8. Add a new vertex C to T and connect to vertices C 1 and C 2 9. Remove rows and columns of d corresponding to C 1 and C 2 10. Add a row and column to d corrsponding to the new cluster C 11. return T The algorithm takes a nxn distance matrix d of pairwise as an input. Different ways to define distances between clusters may lead to different clusterings

Hierarchical clustering: Recomputing distances § dmin(C, C*) = min d(x, y) for all elements x in C and y in C* – Distance between two clusters is the smallest distance between any pair of their elements § davg(C, C*) = (1 / |C*||C|) ∑ d(x, y) for all elements x in C and y in C* – Distance between two clusters is the average distance between all pairs of their elements

K-Means clustering problem § Input: A set, V, consisting of n points and a parameter k § Output: A set X consisting of k points (cluster centers) that minimizes the squared error distortion d(V, X) over all possible choices of X Given a data point v and a set of points X, define the distance from v to X d(v, X) as the (Euclidian) distance from v to the closest point from X. Given a set of n data points V={v 1…vn} and a set of k points X, define the Squared Error Distortion d(V, X) = ∑d(vi, X)2 / n 1<i<n

1 -Means clustering problem § Input: A set, V, consisting of n points § Output: A single point x (cluster center) that minimizes the squared error distortion d(V, x) over all possible choices of x 1 -Means Clustering problem is easy. However, it becomes very difficult (NP-complete) for more than one center. An efficient heuristic method for K-Means clustering is the Lloyd algorithm

K-Means clustering: Lloyd algorithm 1. Lloyd Algorithm 2. Arbitrarily assign the k cluster centers 3. while the cluster centers keep changing 4. Assign each data point to the cluster Ci corresponding to the closest cluster representative (center) (1 ≤ i ≤ k) 5. After the assignment of all data points, compute new cluster representatives according to the center of gravity of each cluster, that is, the new cluster representative is ∑v |C| for all v in C for every cluster C *This may lead to merely a locally optimal clustering.
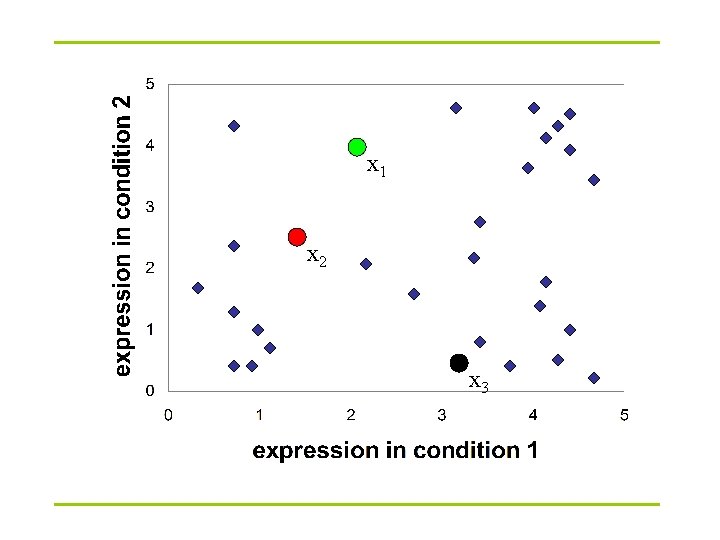
x 1 x 2 x 3
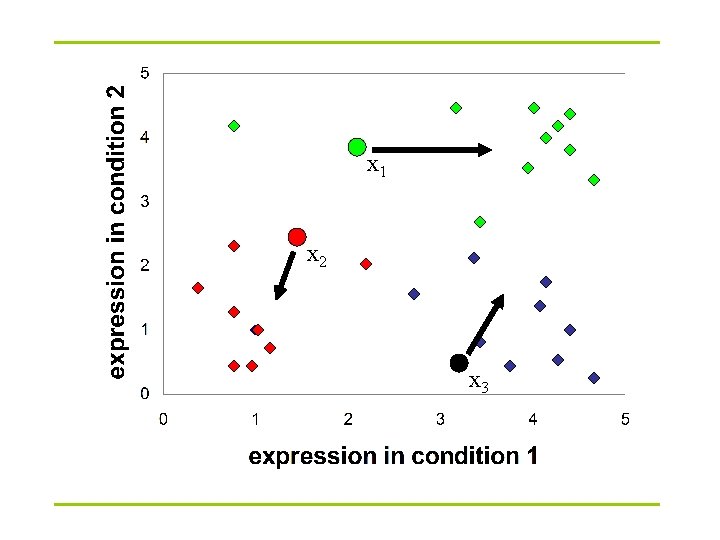
x 1 x 2 x 3
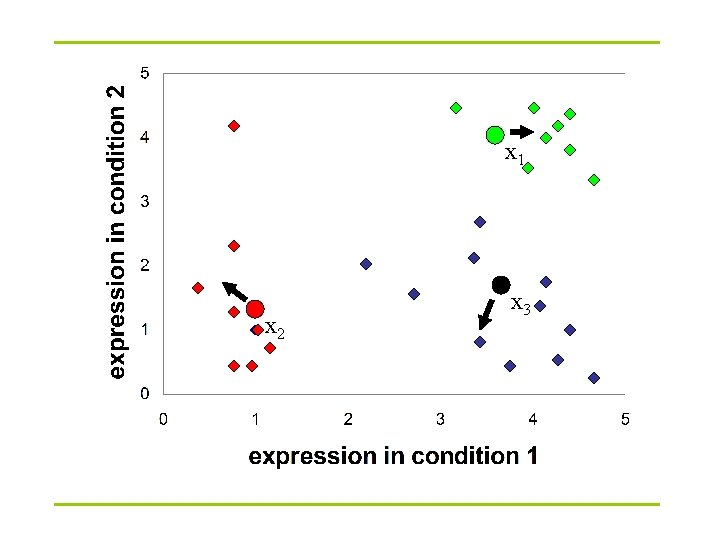
x 1 x 2 x 3
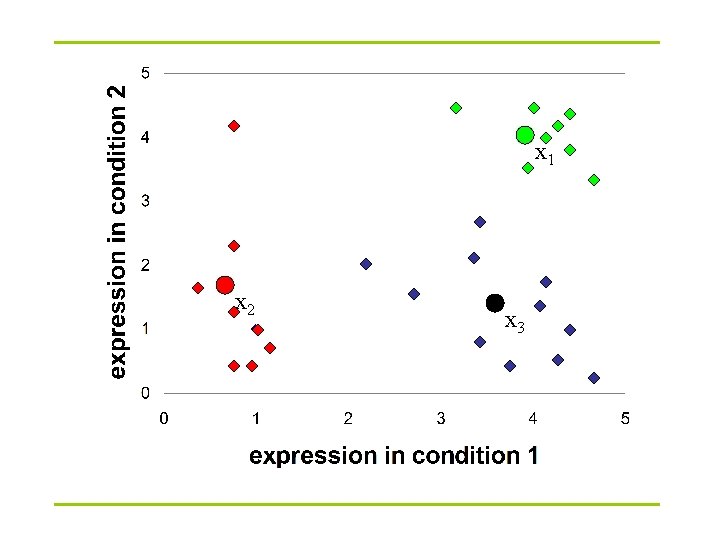
x 1 x 2 x 3

Biclustering § If two genes are related (have similar functions or are co-regulated), their expression profiles should be similar (e. g. low Euclidean distance or high correlation). § However, they can have similar expression patterns only under some conditions (e. g. they have similar response to a certain external stimulus, but each of them has some distinct functions at other time). § Similarly, for two related conditions, some genes may exhibit different expression patterns (e. g. two tumor samples of different sub-types).
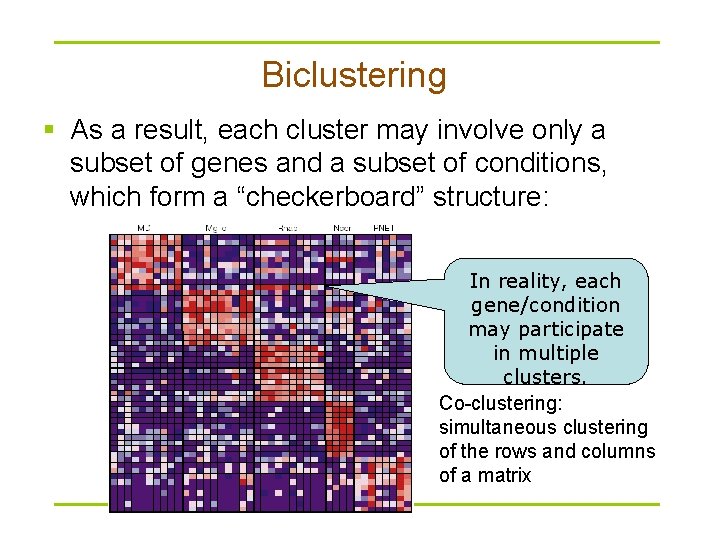
Biclustering § As a result, each cluster may involve only a subset of genes and a subset of conditions, which form a “checkerboard” structure: In reality, each gene/condition may participate in multiple clusters. Co-clustering: simultaneous clustering of the rows and columns of a matrix

Biclustering § To discover such data patterns, some “biclustering” methods have been proposed to cluster both genes and conditions simultaneously. § Differences with projected clustering (by observation, not be definition): – Projected clustering has a primary clustering target, biclustering usually treats rows and columns equally. – Most projected clustering methods define attribute relevance based on value distances, most biclustering methods define biclusters based on other measures. – Some biclustering methods do not have the concept of irrelevant attributes.

Sample classification § Classify samples based on gene expression pattern AML: acute myeloid leukemia ALL: acute lymphoblastic leukemia c: idealized expression pattern c*: random idealized expression pattern The prediction of a new sample is based on "weighted votes" of a set of informative genes (each gene votes for AML or ALL depending its expression level) Ref: Molecular Classification of Cancer: Class Discovery and Class Prediction by Gene Expression Monitoring (Science, 1999)

Packages for microarray data analysis § Packages in R – LIMMA, a library for the analysis of gene expression microarray data, especially the use of linear models for analysing designed experiments and the assessment of differential expression; part of Bioconductor – Bioconductor is open source software for bioinformatics, and it consists of 352 packages (Bio. C 2. 5)
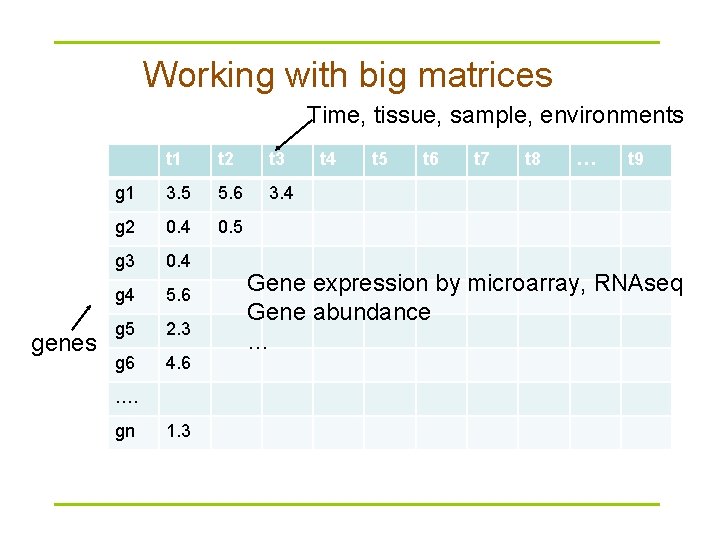
Working with big matrices Time, tissue, sample, environments genes t 1 t 2 t 3 g 1 3. 5 5. 6 3. 4 g 2 0. 4 0. 5 g 3 0. 4 g 4 5. 6 g 5 2. 3 g 6 4. 6 …. gn 1. 3 t 4 t 5 t 6 t 7 t 8 … t 9 Gene expression by microarray, RNAseq Gene abundance …
- Slides: 39