i 2 b 2 Clinical Research Chart and

i 2 b 2 Clinical Research Chart and Hive Architecture Henry Chueh Shawn Murphy Isaac Kohane, PI i 2 b 2 National Center for Biomedical Computing

Summary • • Background Intro to the Clinical Research Chart (CRC) Hive / Cell Software Architecture More details on establishing and using the CRC i 2 b 2 National Center for Biomedical Computing

Background • Clinical documentation is…clinical • Lack of systematic approach for organizing clinical data for research • Ownership issues are unique • Consent issues are a challenge i 2 b 2 National Center for Biomedical Computing

Driving Biological Projects • • Asthma Hypertension Huntington’s Disease Diabetes i 2 b 2 National Center for Biomedical Computing

Clinical Research Chart (CRC) • Organize and transform clinical data to maximize its utility for research • Develop an Application and Database framework to serve this goal • Establish an architecture that allows data from different studies done on this platform to be integrated i 2 b 2 National Center for Biomedical Computing
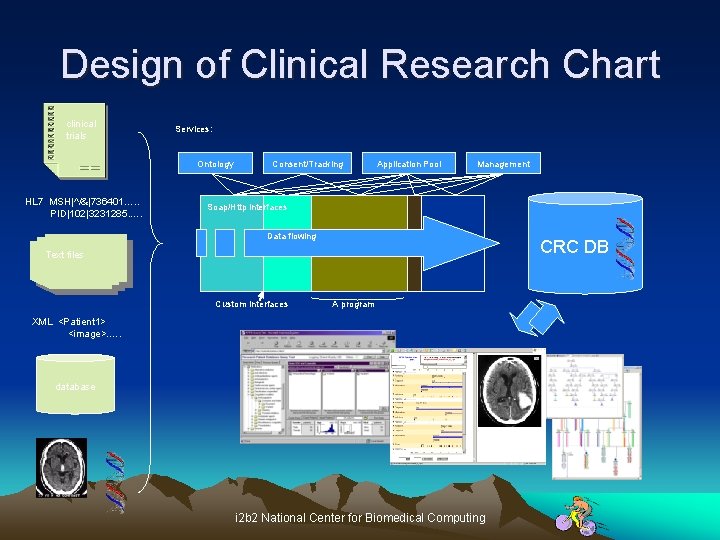
Design of Clinical Research Chart clinical trials Services: Ontology HL 7 MSH|^/&|736401…. . PID|102|3231285. …. Consent/Tracking Application Pool Management Soap/Http interfaces Data flowing CRC DB Text files Custom Interfaces A program XML <Patient 1> <image>. …. database i 2 b 2 National Center for Biomedical Computing
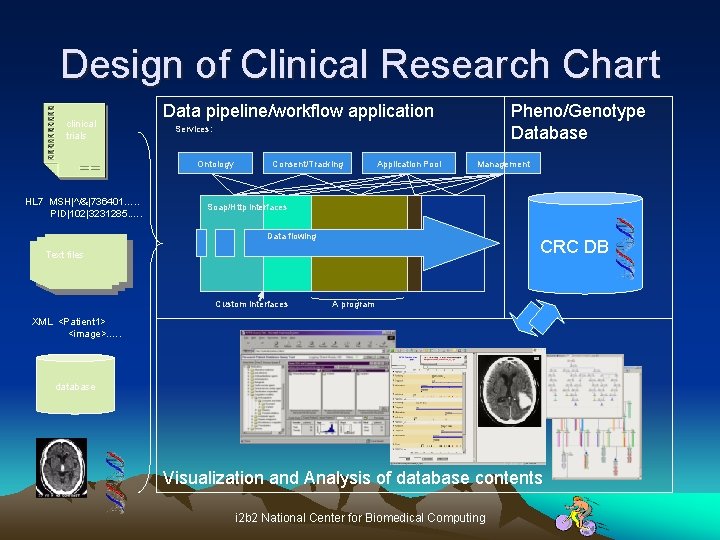
Design of Clinical Research Chart clinical trials Data pipeline/workflow application Services: Ontology HL 7 MSH|^/&|736401…. . PID|102|3231285. …. Pheno/Genotype Database Consent/Tracking Application Pool Management Soap/Http interfaces Data flowing CRC DB Text files Custom Interfaces A program XML <Patient 1> <image>. …. database Visualization and Analysis of database contents i 2 b 2 National Center for Biomedical Computing
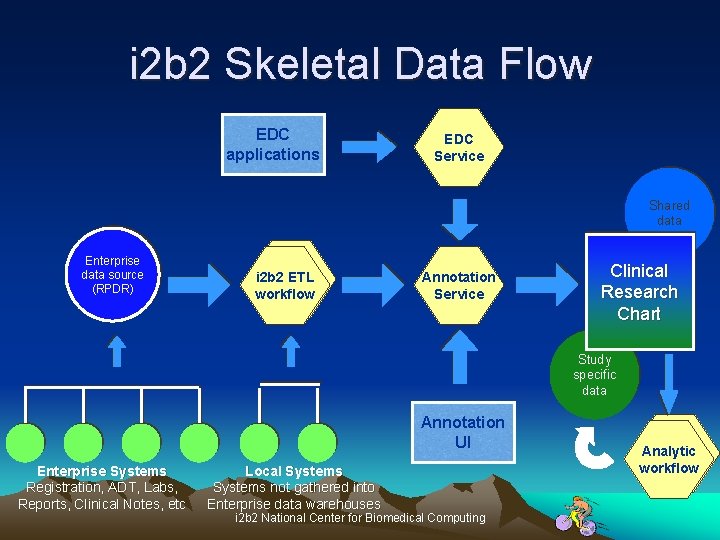
i 2 b 2 Skeletal Data Flow EDC applications EDC Service Shared data Enterprise data source (RPDR) i 2 b 2 ETL workflow Annotation Service Clinical Research Chart Study specific data Annotation UI Enterprise Systems Registration, ADT, Labs, Reports, Clinical Notes, etc Local Systems not gathered into Enterprise data warehouses i 2 b 2 National Center for Biomedical Computing Analytic workflow

Overall Themes • Framework to allow development of application services in a maximally decoupled fashion. • Linux and Windows OS support • Java and C++ programming languages • Use Cases for construction of CRC come from Driving Biology Projects and experience with clients of Partners Research Patient Data Registry i 2 b 2 National Center for Biomedical Computing

Focus on Workflow • Necessary for both pre-CRC and post. CRC processes • Needed for scientific flexibility • Implies a consistent environment for data pipelining and flow control i 2 b 2 National Center for Biomedical Computing

i 2 b 2 Hive • Formed as a collection of interoperable Cells, or services • Loosely coupled • Makes no assumptions about proximity • Connected by Web services • Activated by a workflow engine that forms basis of choreography among Cells for complex interactions i 2 b 2 National Center for Biomedical Computing

Complex choreography i 2 b 2 National Center for Biomedical Computing

i 2 b 2 Cell • Behaves as a functional service • Separates interactions conceptually into transactions and semantics • Focuses on facilitating transactions with simple semantics (e. g. , datatype) • Leaves deep semantics to be defined by the services provided by a Cell • Does not restrict language implementation i 2 b 2 National Center for Biomedical Computing

Target layer for i 2 b 2 Semantic Objects I 2 b 2 platform Web Services TCP/IP i 2 b 2 National Center for Biomedical Computing

Cell examples • Concept extraction from clinical narratives • Simple transformations; e. g. , basic text format conversion • Complex encoding; e. g. , encoding MIAME in MAGE • Microarray data normalization • … i 2 b 2 National Center for Biomedical Computing

Exposing Cells • Protocols layered on top of SOAP • At the WSDL level for integrators; ie, bioinformaticians & software engineers • At a functional level for investigators • i 2 b 2 toolkits to allow integrators to expose controlled functionality to investigators (Automator) i 2 b 2 National Center for Biomedical Computing
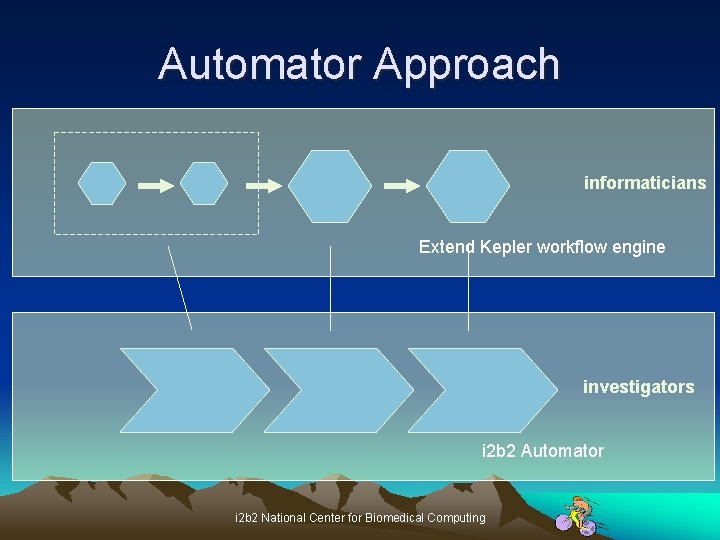
Automator Approach informaticians Extend Kepler workflow engine investigators i 2 b 2 Automator i 2 b 2 National Center for Biomedical Computing
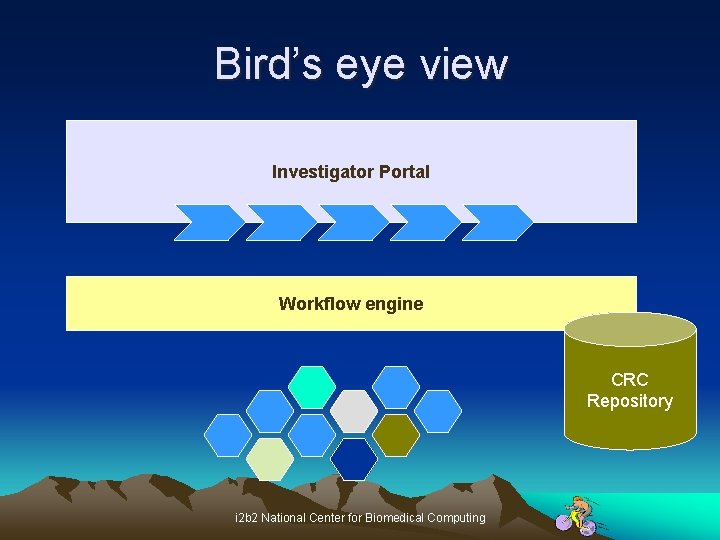
Bird’s eye view Investigator Portal Workflow engine CRC Repository i 2 b 2 National Center for Biomedical Computing

Current Implementation • Extending Kepler workflow engine for i 2 b 2 • Data model for CRC repository • Defining protocols necessary for interaction (in addition to SOAP) • Created Cell for concept extraction from narratives • Early designs for Automator toolkit i 2 b 2 National Center for Biomedical Computing

i 2 b 2 Architecture Key Points • Leverage existing workflow standards and software • Use Web services as basic form of interaction • Assume unlimited choreography, but… • Provide tools to distill complexity into basic automation for clinical investigators i 2 b 2 National Center for Biomedical Computing

SW Licensing and Distribution • Commit to Open Source software • Use GNU Lesser General Public License • Establish local i 2 b 2 repository exposed through i 2 b 2 website • Contribute to a more global NCBC Source. Forge style repository if it emerges ? NIH Forge • Keep i 2 b 2 protocols fully open i 2 b 2 National Center for Biomedical Computing

Interoperability across NCBC • Strongly consider Web services as basic protocol for generic shared interactions • Consider sharing datasets • Promote diversity of approach and use of shared software (don’t impose uniformity) • Facilitate/promote NCBC Open Source project teams i 2 b 2 National Center for Biomedical Computing

Pre-CRC Data Pipeline/Workflow Populating the Clinical Research Chart (CRC) i 2 b 2 National Center for Biomedical Computing

Pre-CRC Data Pipeline/Workflow • Use workflow framework to choreograph applications services in specific sequences • Used to extract, transform, conform, and load data and metadata into the CRC i 2 b 2 National Center for Biomedical Computing
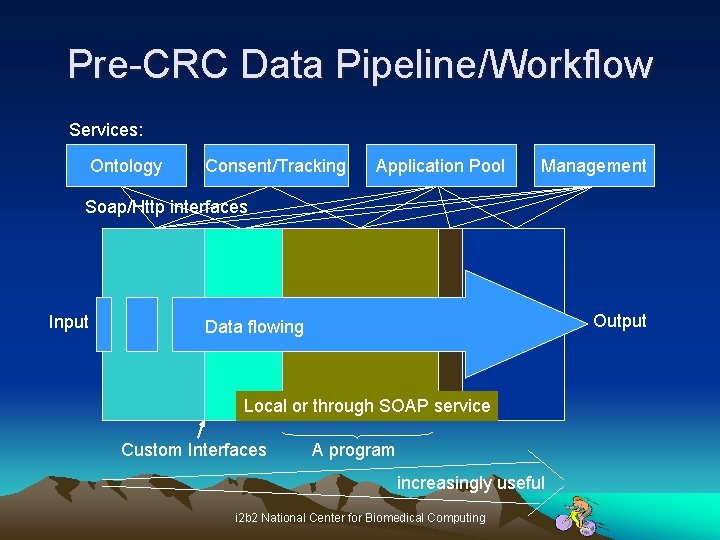
Pre-CRC Data Pipeline/Workflow Services: Ontology Consent/Tracking Application Pool Management Soap/Http interfaces Input Output Data flowing Local or through SOAP service Custom Interfaces A program increasingly useful i 2 b 2 National Center for Biomedical Computing

Ontology Service Ontology Consent/Tracking Application Pool Management • Manages mappings of terms to common vocabularies • Provides lists of acceptable (enumerated) values for various attribute and value slots. • Allows for management of hierarchies, groupings, and relationships between terms i 2 b 2 National Center for Biomedical Computing

Person Consent/Tracking Service Ontology Consent/Tracking Application Pool Management • Provides mappings between patient/subject identifiers • Tracks patient/subject consent information • Allows identification of the patient/subject based upon fuzzy demographic matches i 2 b 2 National Center for Biomedical Computing

Application Pool (CVS) Service Ontology • • Consent/Tracking Application Pool Management Stores programs/scripts used in pipeline Provides applications to be downloaded when needed Manages versioning of software Provides documentation i 2 b 2 National Center for Biomedical Computing

Management Service Ontology • • • Consent/Tracking Application Pool Management Stores workflow execution plan Starts and controls workflow execution Schedules workflow execution Monitors workflow execution and data locations Controls permissions associated with workflow execution i 2 b 2 National Center for Biomedical Computing
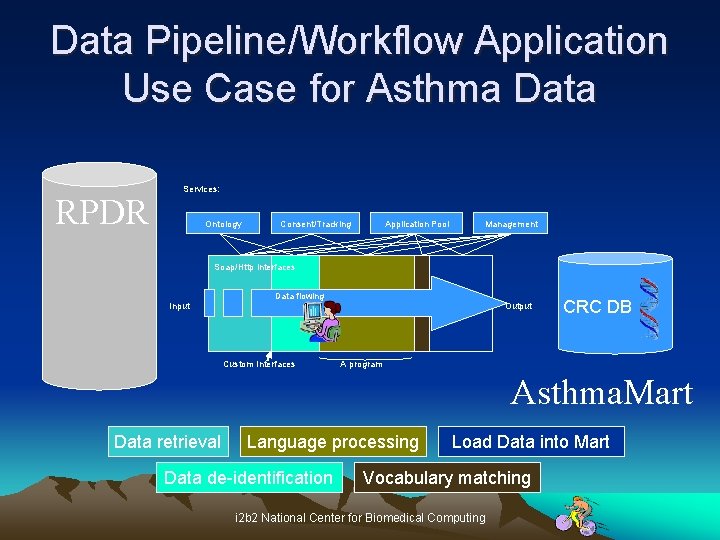
Data Pipeline/Workflow Application Use Case for Asthma Data RPDR Services: Ontology Consent/Tracking Application Pool Management Soap/Http interfaces Input Data flowing Custom Interfaces Output CRC DB A program Asthma. Mart Data retrieval Language processing Data de-identification Load Data into Mart Vocabulary matching i 2 b 2 National Center for Biomedical Computing

Data Pipeline/Workflow Implementation • Define standard XML representation for workflow - Mo. ML • Define standards for SOAP services and resource discovery • Adopt and extend open source workflow package (Kepler) • Prototypes by July timeframe • BIRN -> NAMIC and LONI collaboration • Can follow construction details at http: //diagon/i 2 b 2 National Center for Biomedical Computing

Phenotype/Genotype Database i 2 b 2 National Center for Biomedical Computing

Phenotype/Genotype Database Principles • Analytical database schema that does not need to change with new data types and concepts • Defined fundamental unit of data (atomic fact) = observation • Defined metadata strategy • Various levels of de-identification (reviewed and approved by IRB) i 2 b 2 National Center for Biomedical Computing

Phenotype/Genotype Database Architecture (see preprint) i 2 b 2 National Center for Biomedical Computing

Phenotype/Genotype Database Use Case • Smoking observations represented in database Provider_id Provider_path Name_char M 0022303 MGHNeurologyM 0022303 Concept_cd Concept_path Name_char CT-A-SMK Asth. V 1DRpt. NLPTobacco UseSmoker Smoking IC 9 -3051 V 2DiagnosisMental Disorders (290 -319)Nonpsychotic disorders (300 -316)(305) Nondependent abuse of drugs(305 -1) Tobacco use disorder(30511) Tobacco use disorder, co~ Tobacco Use Disorder, continuous use CT-A-NSK Asth. V 1DRpt. NLPTobacco UseNon smoker Never smoked Patient_id_e Concept_cd Start_date Provider_id Confidence_num Z 234 CT-A-SMK 1/1/1997 M 0022303 3 Z 234 CT-A-SMK 1/1/1998 M 0034125 9 Z 234 IC 9 -3051 1/1/2001 M 0022303 3 Z 234 CT-A-NSK 1/1/2002 M 0034125 9 Patient_id_e Birth_date Sex_cd Race_cd Z 234 3/4/1924 Female Black i 2 b 2 National Center for Biomedical Computing Death_date 4/5/2003

Phenotype/Genotype Database Implementation • Asthma CRC DB “primed” with data from 90, 000 patients from Research Patient Data Registry • Serves as fundamental data structure for i 2 b 2 supported data Querying and Visualization Application Suite • CRC DB’s able to fuse seamlessly together • Various levels of de-identification to be supported for data sharing and publication i 2 b 2 National Center for Biomedical Computing

Visualization and Analysis of CRC database Post-CRC workflow i 2 b 2 National Center for Biomedical Computing

Visualization and Analysis Principles • Supported application suite to query and view CRC database contents • Outside applications for analysis and viewing able to plug in to application suite • Pipeline/Workflow framework may be used for analysis and re-entry of derived data into CRC database i 2 b 2 National Center for Biomedical Computing

Visualization and Analysis Architecture • Supported Applications, Querying and Visualization – Standard querying – Data exploration i 2 b 2 National Center for Biomedical Computing

Visualization and Analysis Architecture • Supported Applications, ontology management – Ontology Management • Integrate (outside? ) population analysis applications i 2 b 2 National Center for Biomedical Computing

Visualization and Analysis Architecture • Supported applications have plug-in architecture for outside analytic tools: – Standard web-link support with GET and POST oriented data transfer – Support transfer of specifically transformed data to outside applications – Complex analysis supported with workflow application i 2 b 2 National Center for Biomedical Computing
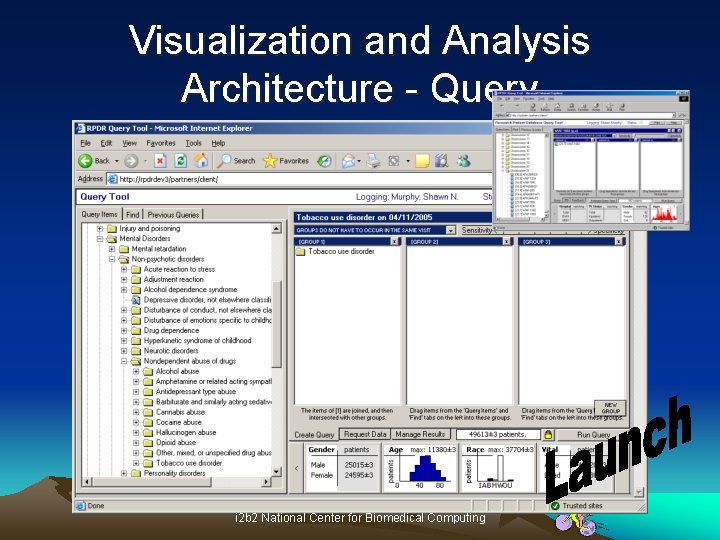
Visualization and Analysis Architecture - Query i 2 b 2 National Center for Biomedical Computing
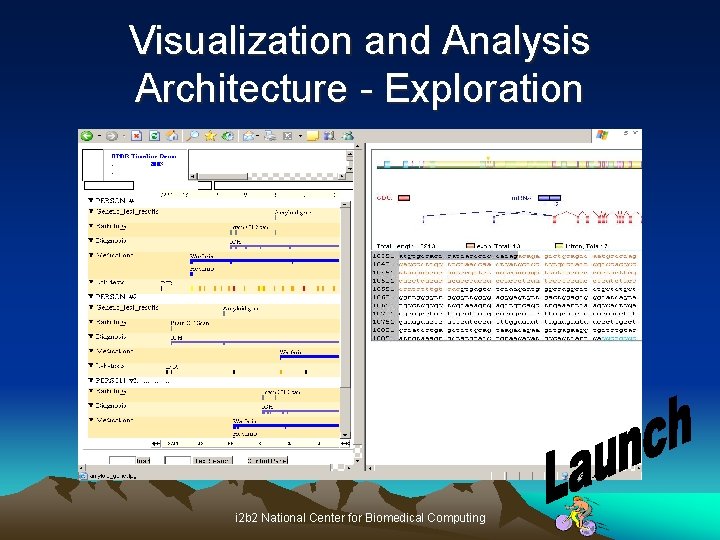
Visualization and Analysis Architecture - Exploration i 2 b 2 National Center for Biomedical Computing
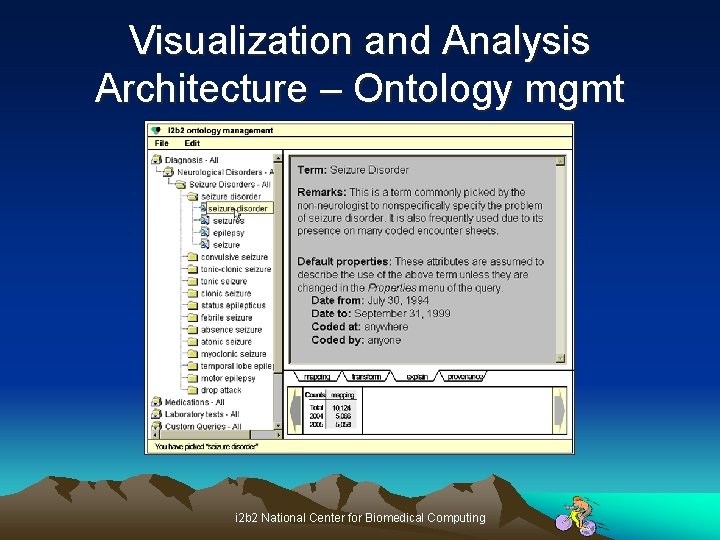
Visualization and Analysis Architecture – Ontology mgmt i 2 b 2 National Center for Biomedical Computing
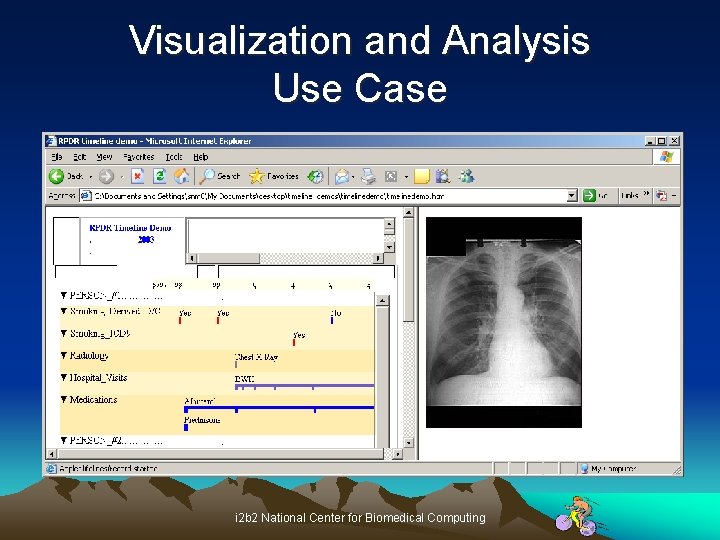
Visualization and Analysis Use Case i 2 b 2 National Center for Biomedical Computing
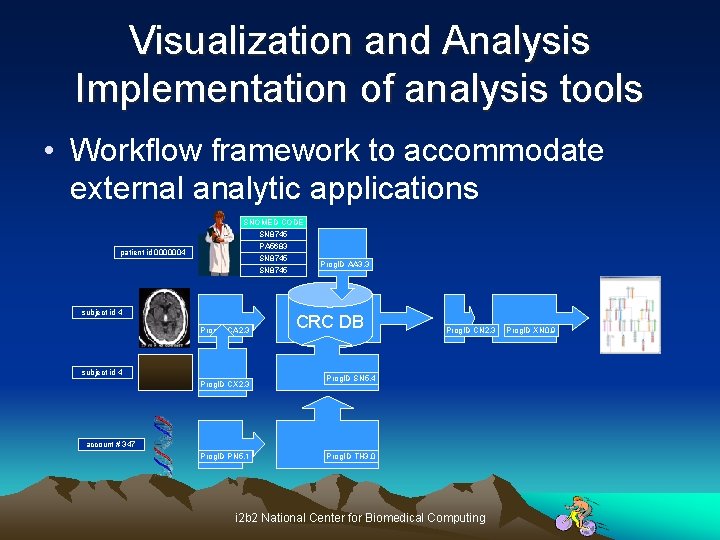
Visualization and Analysis Implementation of analysis tools • Workflow framework to accommodate external analytic applications SNOMED CODE SN 8745 PA 5683 SN 8745 patient id 0000004 subject id 4 Prog. ID CA 2. 3 subject id 4 Prog. ID CX 2. 3 Prog. ID AA 3. 3 CRC DB Prog. ID CN 2. 3 Prog. ID SN 5. 4 account # 347 Prog. ID PN 5. 1 Prog. ID TH 3. 0 i 2 b 2 National Center for Biomedical Computing Prog. ID XN 0. 9
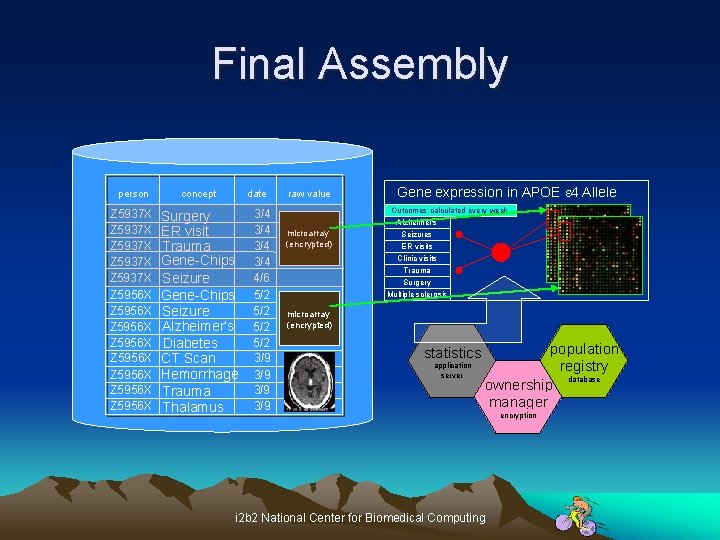
Final Assembly person concept Z 5937 X Z 5937 X Z 5956 X Z 5956 X Surgery ER visit Trauma Gene-Chips Seizure Alzheimer’s Diabetes CT Scan Hemorrhage Trauma Thalamus date 3/4 3/4 4/6 5/2 5/2 3/9 3/9 raw value microarray (encrypted) Gene expression in APOE e 4 Allele Outcomes calculated every week Alzheimer's Seizures ER visits Clinic visits Trauma Surgery Multiple sclerosis microarray (encrypted) statistics application server population registry database ownership manager i 2 b 2 National Center for Biomedical Computing encryption

i 2 b 2 National Center for Biomedical Computing
- Slides: 48