Humboldt University Berlin Theoretical Biophysics Max Planck Institute

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Networks in Metabolism and Signaling Edda Klipp Humboldt University Berlin Lecture 4 / WS 2007/08 Boolean Networks VL Netzwerke, WS 2007/08 Edda Klipp 1
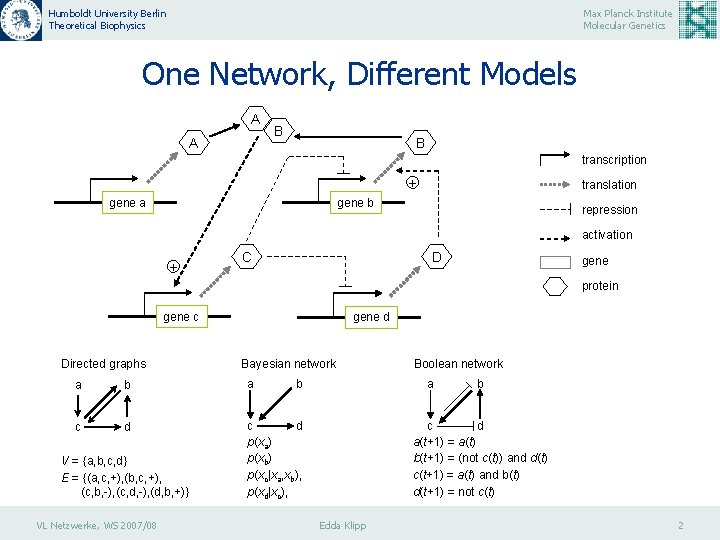
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics One Network, Different Models A A B B transcription + gene a translation gene b repression activation + C D gene protein gene c Directed graphs gene d Bayesian network a b a c d p(xa) p(xb) p(xc|xa, xb), p(xd|xc), V = {a, b, c, d} E = {(a, c, +), (b, c, +), (c, b, -), (c, d, -), (d, b, +)} VL Netzwerke, WS 2007/08 b Boolean network a b c d a(t+1) = a(t) b(t+1) = (not c(t)) and d(t) c(t+1) = a(t) and b(t) d(t+1) = not c(t) Edda Klipp 2
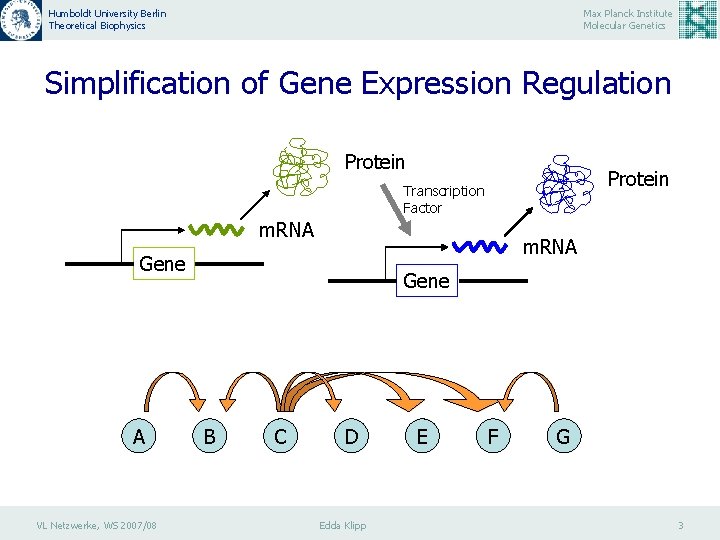
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Simplification of Gene Expression Regulation Protein Transcription Factor m. RNA Gene A VL Netzwerke, WS 2007/08 Gene B C D Edda Klipp E F G 3
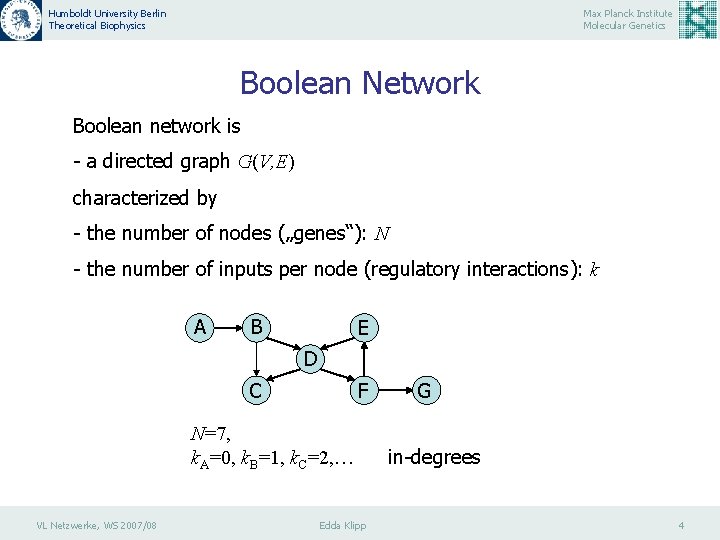
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Boolean network is - a directed graph G(V, E) characterized by - the number of nodes („genes“): N - the number of inputs per node (regulatory interactions): k A B E D C F N=7, k. A=0, k. B=1, k. C=2, … VL Netzwerke, WS 2007/08 Edda Klipp G in-degrees 4

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Logic Boolean models are discrete (in state and time) and deterministic. (George Boole, 1815 -1864) Each gene can assume one of two states: expressed („ 1“) or not expressed („ 0“) Background: Not enough information for more detailed description Increasing complexity and computational effort for more specific models Replacement of continuous functions (e. g. Hill function) by step function VL Netzwerke, WS 2007/08 Edda Klipp 5

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Boolean networks have always a finite number of possible states: 2 N and, therefore, a finite number of state transitions: A B E D C N=7, 27 states, VL Netzwerke, WS 2007/08 F G theoretically possible state transition Edda Klipp 6

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Dynamics of Boolean Network. S The dynamics are described by rules: „if input value/s at time t is/are. . , then output value at t+1 is. . “ A A(t) VL Netzwerke, WS 2007/08 B B(t+1) Edda Klipp 7

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Models: Truth functions A B A(t) B(t+1) in output 0 1 0 0 p rule VL Netzwerke, WS 2007/08 0 0 1 1 0 p not p 1 2 Edda Klipp 1 1 B(t+1) = not (A(t)) rule 2 3 8

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Dynamics of Boolean Networks with k=1 A B C D Linear chain A fixed (no input). Rules 0 and 3 not considered (since independence of input). A(t) B(t+1) C(t+2) D(t+3) The system reaches a steady state after N-1 time steps. VL Netzwerke, WS 2007/08 Edda Klipp 9

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Dynamics of Boolean Networks with k=1 A A B B C D Ring Again: Rules 0 and 3 not considered (since independence of input). A(t+1)=B(t) B(t+1)=A(t) Both rule 1 AB 00 00 00 AB 10 01 10 AB 01 10 01 AB 11 11 11 A(t+1)=not B(t) B(t+1)=A(t) Both rule 1 AB 00 10 11 01 00 AB 10 11 01 00 10 AB 01 00 10 11 01 AB 11 01 00 10 11 VL Netzwerke, WS 2007/08 Edda Klipp Fixpoint or cycle of length 2 depending on initial conditions Cycle of length 4 independent of initial conditions. 10

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Attractor The trajectory connects the successive states for increasing time. An attractor is a region of a dynamical system's state space that the system can enter but not leave, and which contains no smaller such region (a special trajectory). Fixpoint – cycle of length 1 Cycles of length L Basin of attraction: is the surrounding region in state space such that all trajectories starting in that region end up in the attractor. Bifurcation: appearance of a boarder separating two basins of attraction. VL Netzwerke, WS 2007/08 Edda Klipp 11
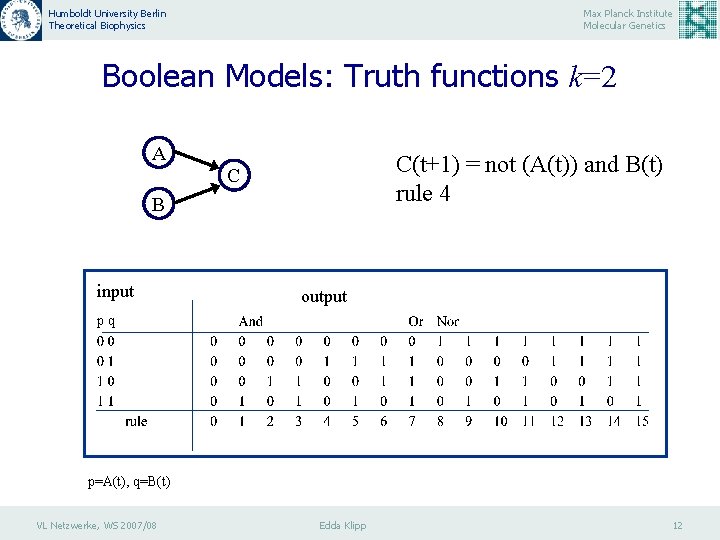
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Models: Truth functions k=2 A C(t+1) = not (A(t)) and B(t) rule 4 C B input output pq p=A(t), q=B(t) VL Netzwerke, WS 2007/08 Edda Klipp 12
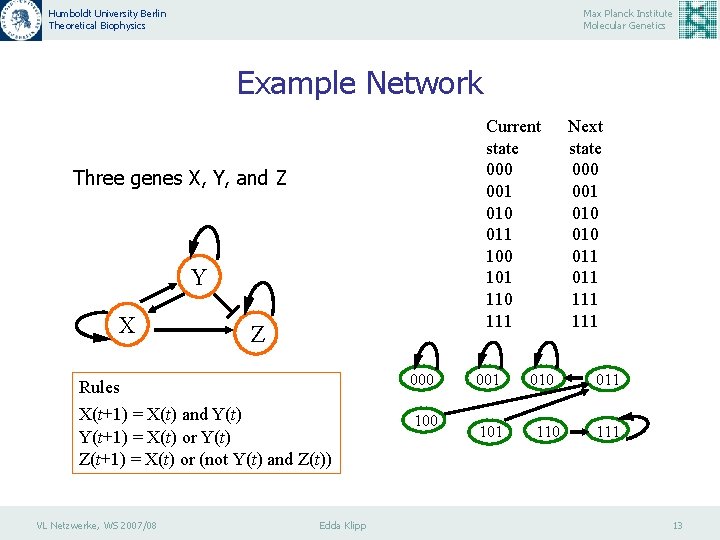
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example Network Current state 000 001 010 011 100 101 110 111 Three genes X, Y, and Z Y X Z Rules 000 X(t+1) = X(t) and Y(t) Y(t+1) = X(t) or Y(t) Z(t+1) = X(t) or (not Y(t) and Z(t)) 100 VL Netzwerke, WS 2007/08 Edda Klipp 001 101 010 110 Next state 000 001 010 011 111 011 13
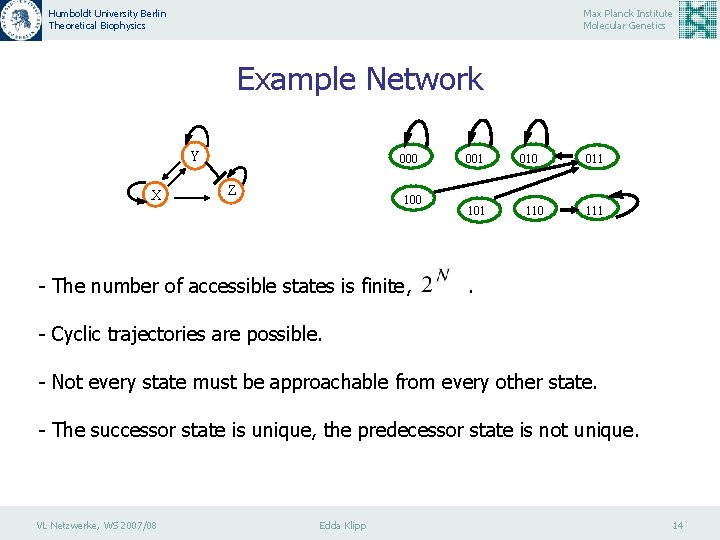
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example Network Y X 000 Z 100 - The number of accessible states is finite, 001 101 010 110 011 111 . - Cyclic trajectories are possible. - Not every state must be approachable from every other state. - The successor state is unique, the predecessor state is not unique. VL Netzwerke, WS 2007/08 Edda Klipp 14
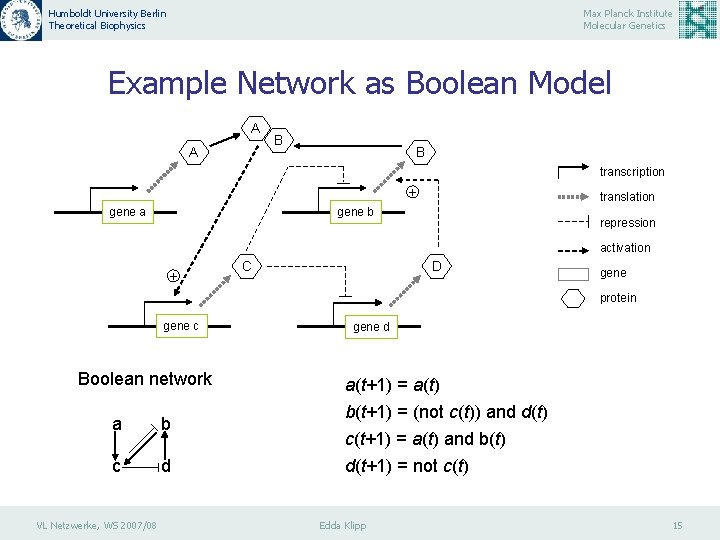
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example Network as Boolean Model A A B B transcription + gene a translation gene b repression activation + C D gene protein gene c Boolean network a b c d VL Netzwerke, WS 2007/08 gene d a(t+1) = a(t) b(t+1) = (not c(t)) and d(t) c(t+1) = a(t) and b(t) d(t+1) = not c(t) Edda Klipp 15

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example Network as Boolean Model Boolean network a b c d a(t+1) = a(t) b(t+1) = (not c(t)) and d(t) c(t+1) = a(t) and b(t) 0000 0001 0010 0011 0100 0101 0110 0111 0001 0101 0000 1000 1001 1010 1011 1100 1101 1110 1111 1001 1101 1000 1011 1111 1010 d(t+1) = not c(t) Steady state: 0101 Cycle: 1000 1001 1111 1010 1000 VL Netzwerke, WS 2007/08 Edda Klipp 16

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Naïve Reconstruction of Boolean Models If it is known -the number of vertices, N, and -the number of inputs per vertex, k, -As well as a sufficient set of successive states, one can reconstruct the network List - List for each vertex all possible input combinations - List all respective outputs Experiments: - Delete after every “experiment” all “wrong” entries of the list VL Netzwerke, WS 2007/08 Edda Klipp 17
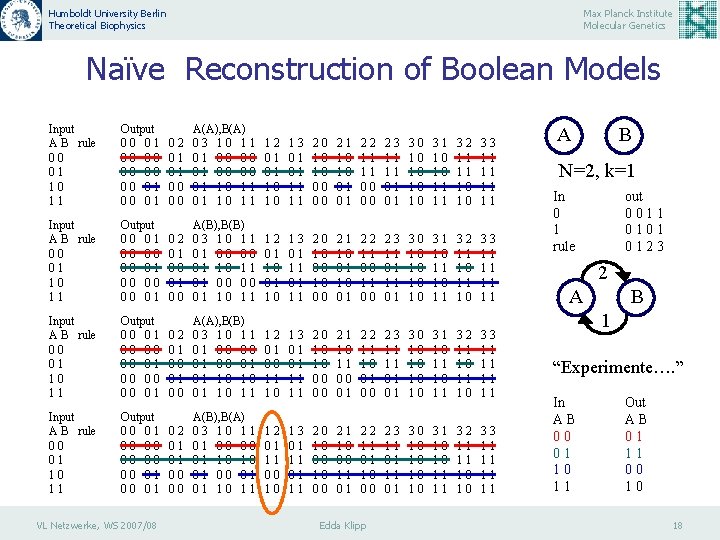
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Naïve Reconstruction of Boolean Models Input A B rule 00 01 10 11 Output 00 01 00 00 00 01 02 01 01 00 00 A(A), B(A) 03 10 11 01 00 00 01 10 11 12 01 01 10 10 13 01 01 11 11 20 10 10 00 00 21 10 10 01 01 22 11 11 00 00 23 11 11 01 01 30 10 10 31 10 10 11 11 32 11 11 10 10 33 11 11 Input A B rule 00 01 10 11 Output 00 01 00 00 00 01 02 01 00 A(B), B(B) 03 10 11 01 00 00 01 10 11 12 01 10 13 01 11 20 10 00 21 10 01 22 11 00 23 11 01 30 10 10 31 10 11 32 11 10 33 11 11 Input A B rule 00 01 10 11 Output 00 01 00 00 00 01 02 01 00 A(A), B(B) 03 10 11 01 00 00 01 01 10 10 01 10 11 12 01 00 11 10 13 01 01 11 11 20 10 10 00 00 21 10 11 00 01 22 11 10 01 00 23 11 11 01 01 30 10 10 31 10 11 32 11 10 33 11 11 Input A B rule 00 01 10 11 Output 00 01 00 00 00 01 02 01 01 00 00 A(B), B(A) 03 10 11 01 00 00 01 10 10 01 01 10 11 12 01 11 00 10 13 01 11 20 10 00 21 10 00 11 01 22 11 01 10 00 23 11 01 30 10 10 31 10 10 11 11 32 11 11 10 10 33 11 11 VL Netzwerke, WS 2007/08 Edda Klipp A B N=2, k=1 In 0 1 rule out 0011 0101 0123 2 A B 1 “Experimente…. ” In AB 00 01 10 11 Out AB 01 11 00 10 18

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Random Boolean Networks If the rules for updating states are unknown select rules randomly Rule 0 Rule 2 Rule 1 N nodes VL Netzwerke, WS 2007/08 ½ p. N (N-1) edges Edda Klipp 19

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Kauffman’s NK Boolean Networks An NK automaton is an autonomous random network of N Boolean logic elements. Each element has K inputs and one output. The signals at inputs and outputs take binary (0 or 1) values. The Boolean elements of the network and the connections between elements are chosen in a random manner. There are no external inputs to the network. The number of elements N is assumed to be large. S. A. Kauffman, 1969, J Theor Biol. Metabolic Stability and Epigenesis in Randomly Constructed Genetic Nets S. A. Kauffman. The Origins of Order: Self-Organization and Selection in Evolution, Oxford University Press, New York, 1993. S. A. Kauffman, 2003, PNAS, Random Boolean Network Models and the Yeast Transcriptional Network VL Netzwerke, WS 2007/08 Edda Klipp 20

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Kauffman’s NK Boolean Networks An automaton operates in discrete time. The set of the output signals of the Boolean elements at a given moment of time characterizes a current state of an automaton. During an automaton operation, the sequence of states converges to a cyclic attractor. The states of an attractor can be considered as a "program" of an automaton operation. The number of attractors M and the typical attractor length L are important characteristics of NK automata. VL Netzwerke, WS 2007/08 Edda Klipp 21

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Kauffman’s Boolean Network Fundamental question: require metabolic stability and epigenesis the genetic regulatory circuits to be precisely constructed? ? Has fortunate evolutionary history selected only nets of highly ordered circuits which alone insure metabolic stability; Or are stability and epigenesis, even in nets of randomly interconnected regulatory circuits, to be expected as the probable consequence of as yet unknown mathematical laws? Are living things more akin to precisely programmed automata selected by evolution, or to randomly assembled automata…? Note: cellular differentiation despite identical sets of genes VL Netzwerke, WS 2007/08 Edda Klipp 22

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Kauffman’s Boolean Network VL Netzwerke, WS 2007/08 Edda Klipp 23

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Further Properties K connections: 22 K Boolean input functions Nets are free of external inputs. Once, connections and rules are selected, they remain constant and the time evolution is deterministic. Earlier work by Walker and Ashby (1965): same Boolean functions for all genes: Choice of Boolean function affects length of cycles: “and” yields short cycles, “exclusive or” yields cycles of immense length VL Netzwerke, WS 2007/08 Edda Klipp 24

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Further Properties: Cycles State of the net: Row listing the present value of all N elements (0 or 1) Finite number of states (2 N) as system passes along a sequence of states from an arbitrary initial state, it must eventually re-enter a state previously passed a cycle Cycle length: number of states on a re-enterant cycle of behavior Cycle of length 1 – equilibrial state Transient (or run-in) length: number of state between initial states and entering the cycle Confluent: set of states leading to or being part of a cycle VL Netzwerke, WS 2007/08 Edda Klipp 25

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Further Properties: Number of Cycles Such a net must contain at least one cycle, it may have more. There number can be counted just be releasing the net from different initial states No state can diverge on to two different states, no state can be on two different cycles VL Netzwerke, WS 2007/08 Edda Klipp 26

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Further Properties: Number of Cycles (a) A net of three binary elements, each of which receives inputs from the other two. The Boolean function assigned to each element is shown beside the element. (b) All possible states of the 3 -element net are shown in the left 3 x 8 matrix below T. The subsequent state of the net at time T+ 1, shown in the matrix on the right, is derived from the inputs and functions shown in (a). (c) A kimatograph showing the sequence of state transitions leading into a state cycle of length 3. All states lie on one confluent. There are three run-ins to the single state cycle. VL Netzwerke, WS 2007/08 Edda Klipp 27

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example: Net with N=10 Periodic attractor (yellow) and basin of attraction (cyan) VL Netzwerke, WS 2007/08 Edda Klipp 28

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example: Net with N=10 The entire state space of an RBN with 10 nodes. Note: Self connections do not appear so a period-1 attractor appears to have no outputs although each network state must have exactly one output. VL Netzwerke, WS 2007/08 Edda Klipp 29

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Further Properties: Distance compares two states of the net Can be defined as the number of genes with different values in two states. For example N=5: state (00000) and state (00111) differ in the value of three elements This is used as measure of dissimilarity between - subsequent states on a transient - subsequent states on a cycle - cycles VL Netzwerke, WS 2007/08 Edda Klipp 30

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Totally Connected Nets, K=N Is like random mapping of a finite set of numbers into itself. Expected length of cycle is E. g. net with N=200 2200 states expected cycle length 2100 ~ 1030 Compare to Hubbel’s age of the universe: 1023 If every transition would take only a second…. Such networks are biologically impossible VL Netzwerke, WS 2007/08 Edda Klipp 31

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics One Connected Nets, K=1 Either one cycle of length N Or a number of disconnected cycles for the full systems state cycles lengths are lowest common multiples of the individual loop lenghts the state cycle length becomes easily very large Again biologically not feasible VL Netzwerke, WS 2007/08 Edda Klipp 32

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Two Connected Nets, K=2 Kauffman studied networks of N= 15, 50, 64, …, 400, 1024, . . , 8191 Nets of 1000 elements possess 21000~10300 states 16 Boolean functions Study of cycle length (surprisingly short) VL Netzwerke, WS 2007/08 Edda Klipp 33
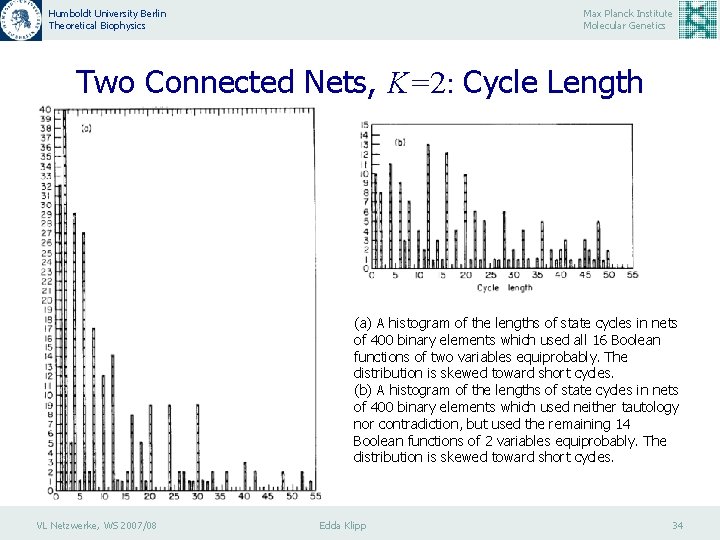
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Two Connected Nets, K=2: Cycle Length (a) A histogram of the lengths of state cycles in nets of 400 binary elements which used all 16 Boolean functions of two variables equiprobably. The distribution is skewed toward short cycles. (b) A histogram of the lengths of state cycles in nets of 400 binary elements which used neither tautology nor contradiction, but used the remaining 14 Boolean functions of 2 variables equiprobably. The distribution is skewed toward short cycles. VL Netzwerke, WS 2007/08 Edda Klipp 34
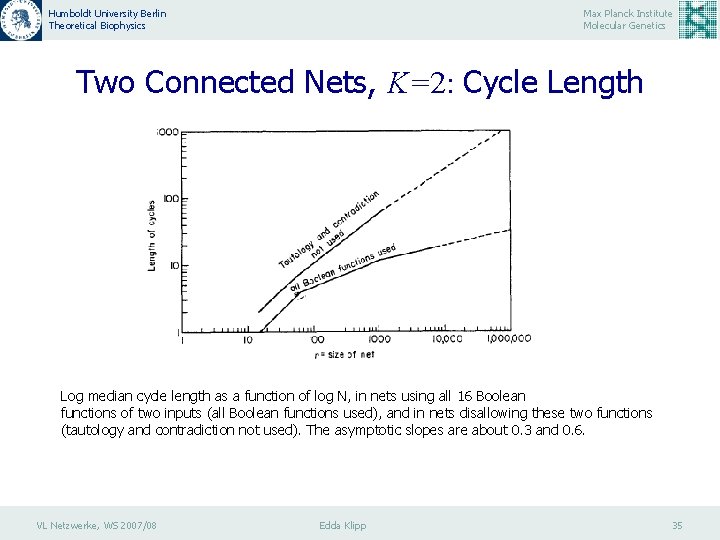
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Two Connected Nets, K=2: Cycle Length Log median cycle length as a function of log N, in nets using all 16 Boolean functions of two inputs (all Boolean functions used), and in nets disallowing these two functions (tautology and contradiction not used). The asymptotic slopes are about 0. 3 and 0. 6. VL Netzwerke, WS 2007/08 Edda Klipp 35
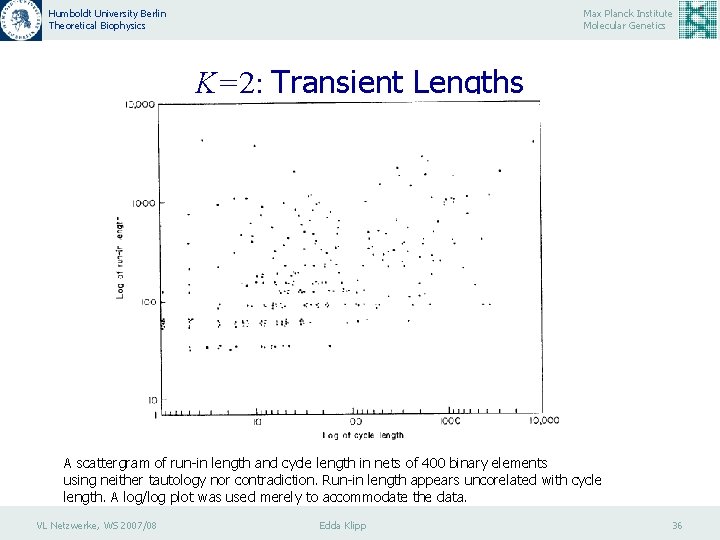
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics K=2: Transient Lengths A scattergram of run-in length and cycle length in nets of 400 binary elements using neither tautology nor contradiction. Run-in length appears uncorelated with cycle length. A log/log plot was used merely to accommodate the data. VL Netzwerke, WS 2007/08 Edda Klipp 36
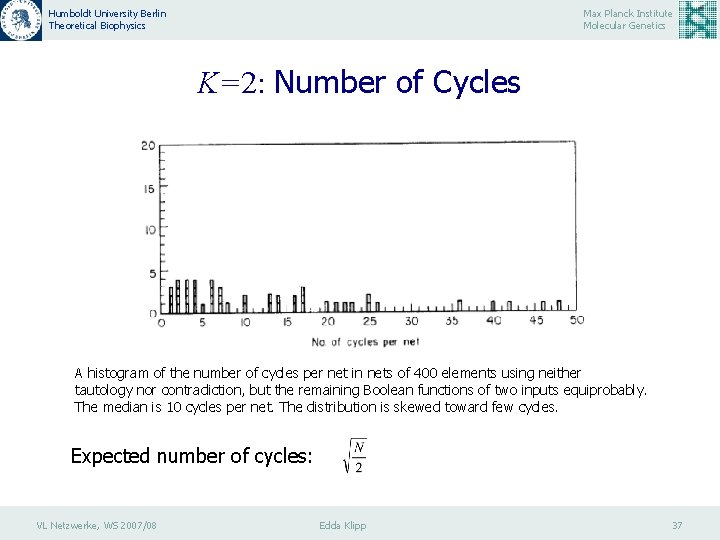
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics K=2: Number of Cycles A histogram of the number of cycles per net in nets of 400 elements using neither tautology nor contradiction, but the remaining Boolean functions of two inputs equiprobably. The median is 10 cycles per net. The distribution is skewed toward few cycles. Expected number of cycles: VL Netzwerke, WS 2007/08 Edda Klipp 37

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics K=2: Activity After release from an arbitrary initial state: Number of elements changing their state per state transition decreases Example: net of 100 elements first step: about 0. 4 N elements change exponential decay of this number minimum activity 0 to 0. 25 N On a cycle: 0 to 35 of 100 elements change most genes are constant during a cycle Bis hier 12. Nov 2007 VL Netzwerke, WS 2007/08 Edda Klipp 38

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Noise One unit of noise may be introduced by arbitrarily changing the value of a single gene for one time moment. The system may return to the cycle perturbed or run into a different cycle. In a net of size N there are just N states which differ from any state in the value of just one gene Consider a net with several cycles: By perturbing all states on each cycle (distance 1) one obtains a matrix listing all cycles and how often they are reached from another one. By dividing all cells by the rows totals transition probabilities The matrix is a Markov chain. VL Netzwerke, WS 2007/08 Edda Klipp 39

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Noise: for the Example Boolean network a b c d Cycle 1 0000 0001 0101 0010 0000 0011 0000 0100 0001 0101 0110 0000 0111 0000 a(t+1) = a(t) b(t+1) = (not c(t)) and d(t) c(t+1) = a(t) and b(t) d(t+1) = not c(t) Transition Matrix VL Netzwerke, WS 2007/08 Steady state: 0101 C 2 C 1 ¾ ¼ Cycle 2 1000 1001 1101 1010 1000 1011 1000 1100 1011 1101 1111 1110 1010 1111 1010 Cycle: 1000 1001 1111 1010 1000 C 2 ¼ ¾ Edda Klipp 40

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Noise (a) A matrix listing the 30 cycles of one net and the total number of times one unit of perturbation shifted the net from each cycle to each cycle. The system generally returns to the cycle perturbed. Division of the value in each cell of the matrix by the total of its row yields the matrix of transition probabilities between modes of behavior which constitute a Markov chain. The transition probabilities between cycles may be asymmetric. (b) Transitions between cycles in the net shown in (a). The solid arrows are the most probable transition to a cycle other than the cycle perturbed, the dotted arrows are the second most probable. The remaining transitions are not shown. Cycles 2, 7, 5 and 15 form an ergodic set into which the remaining cycles flow. If all the transitions between cycles are included, the ergodic set of cycles becomes: 1, 2, 3, 5, 6, 12, 13, 15, 16. The remainder are transient cycles leading into this single ergodic set-. VL Netzwerke, WS 2007/08 Edda Klipp 41
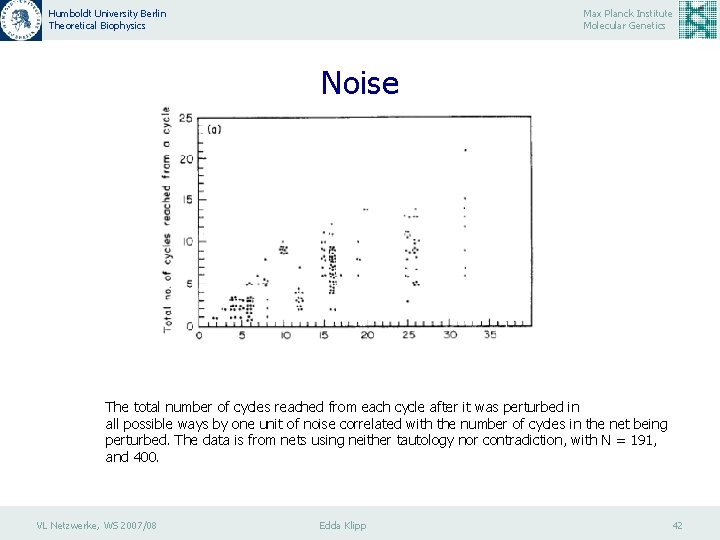
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Noise The total number of cycles reached from each cycle after it was perturbed in all possible ways by one unit of noise correlated with the number of cycles in the net being perturbed. The data is from nets using neither tautology nor contradiction, with N = 191, and 400. VL Netzwerke, WS 2007/08 Edda Klipp 42
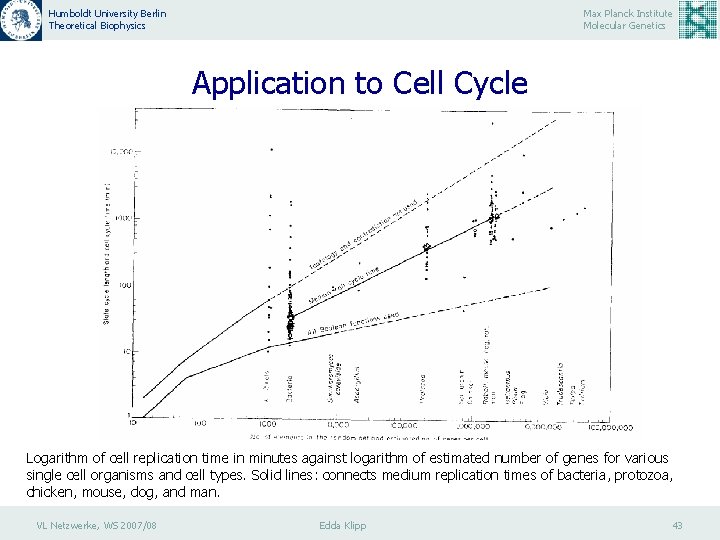
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Application to Cell Cycle Logarithm of cell replication time in minutes against logarithm of estimated number of genes for various single cell organisms and cell types. Solid lines: connects medium replication times of bacteria, protozoa, chicken, mouse, dog, and man. VL Netzwerke, WS 2007/08 Edda Klipp 43
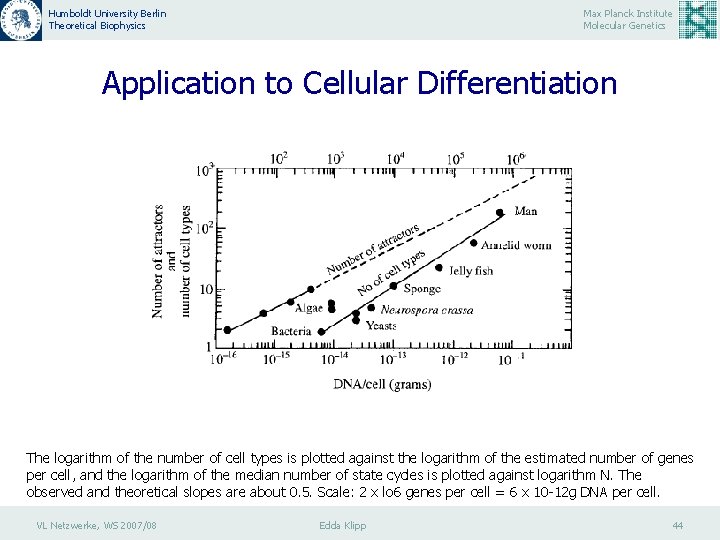
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Application to Cellular Differentiation The logarithm of the number of cell types is plotted against the logarithm of the estimated number of genes per cell, and the logarithm of the median number of state cycles is plotted against logarithm N. The observed and theoretical slopes are about 0. 5. Scale: 2 x lo 6 genes per cell = 6 x 10 -12 g DNA per cell. VL Netzwerke, WS 2007/08 Edda Klipp 44
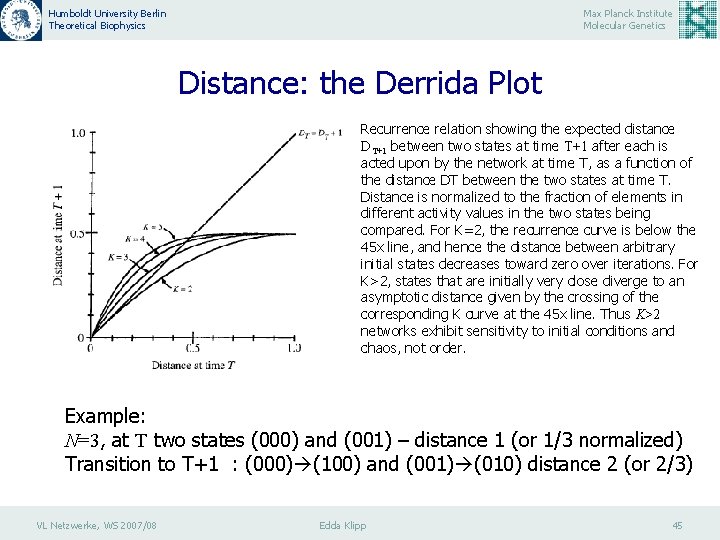
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Distance: the Derrida Plot Recurrence relation showing the expected distance DT+1 between two states at time T+1 after each is acted upon by the network at time T, as a function of the distance DT between the two states at time T. Distance is normalized to the fraction of elements in different activity values in the two states being compared. For K=2, the recurrence curve is below the 45 x line, and hence the distance between arbitrary initial states decreases toward zero over iterations. For K>2, states that are initially very close diverge to an asymptotic distance given by the crossing of the corresponding K curve at the 45 x line. Thus K>2 networks exhibit sensitivity to initial conditions and chaos, not order. Example: N=3, at T two states (000) and (001) – distance 1 (or 1/3 normalized) Transition to T+1 : (000) (100) and (001) (010) distance 2 (or 2/3) VL Netzwerke, WS 2007/08 Edda Klipp 45
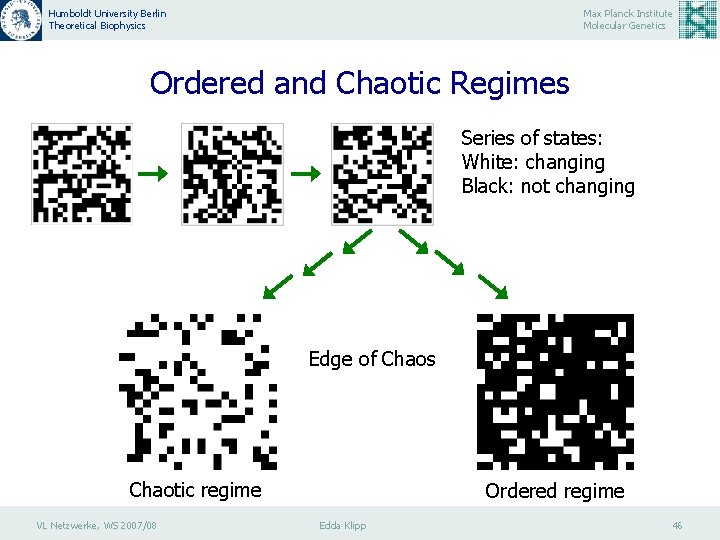
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Ordered and Chaotic Regimes Series of states: White: changing Black: not changing Edge of Chaos Chaotic regime VL Netzwerke, WS 2007/08 Ordered regime Edda Klipp 46

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Kauffman’s NK Boolean Networks Dependence of Behavior on Degree K K large (K = N): the behavior is essentially stochastic. The successive states are random with respect to the preceding ones. The "programs" are very sensitive to minimal disturbances (a minimal disturbance is a change of an output of a particular element during an automaton operation) and to mutations (changes in Boolean element types and in network connections). The attractor lengths L are very large: L ~ 2 N/2. The number of attractors M is of the order of N. If the connection degree K is decreased, this stochastic type of behavior is still observed, until K ~ 2. VL Netzwerke, WS 2007/08 Edda Klipp 47

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Kauffman’s NK Boolean Networks Dependence of Behavior on Degree K At K ~ 2 the network behavior changes drastically. The sensitivity to minimal disturbances is small. The mutations create typically only slight variations an automaton dynamics. Only some rare mutations evoke the radical, cascading changes in the automata "programs". The attractor length L and the number of attractors M are of the order of 1/2 N. This is the behavior at the edge of chaos, at the borderland between chaos and order. VL Netzwerke, WS 2007/08 Edda Klipp 48

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Dynamics – Scale Free Nets VL Netzwerke, WS 2007/08 Edda Klipp 49
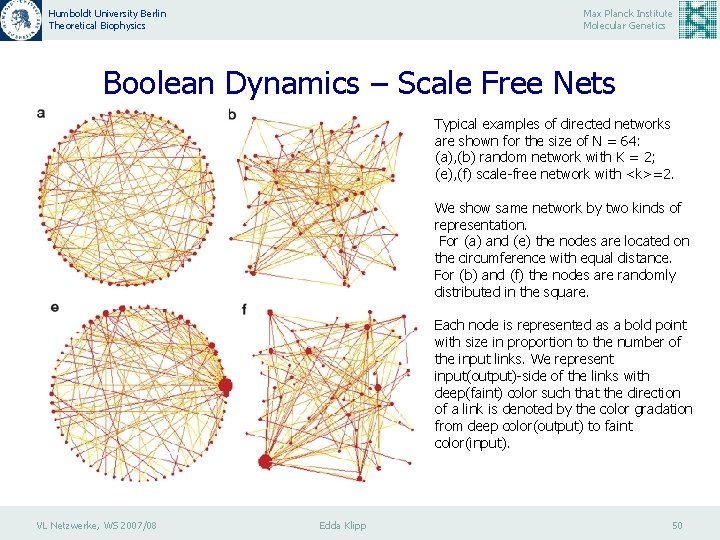
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Dynamics – Scale Free Nets Typical examples of directed networks are shown for the size of N = 64: (a), (b) random network with K = 2; (e), (f) scale-free network with <k>=2. We show same network by two kinds of representation. For (a) and (e) the nodes are located on the circumference with equal distance. For (b) and (f) the nodes are randomly distributed in the square. Each node is represented as a bold point with size in proportion to the number of the input links. We represent input(output)-side of the links with deep(faint) color such that the direction of a link is denoted by the color gradation from deep color(output) to faint color(input). VL Netzwerke, WS 2007/08 Edda Klipp 50
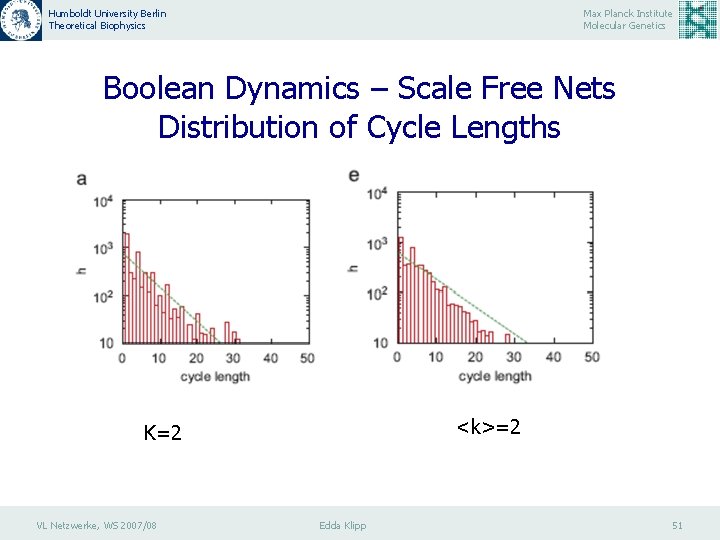
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Dynamics – Scale Free Nets Distribution of Cycle Lengths <k>=2 K=2 VL Netzwerke, WS 2007/08 Edda Klipp 51
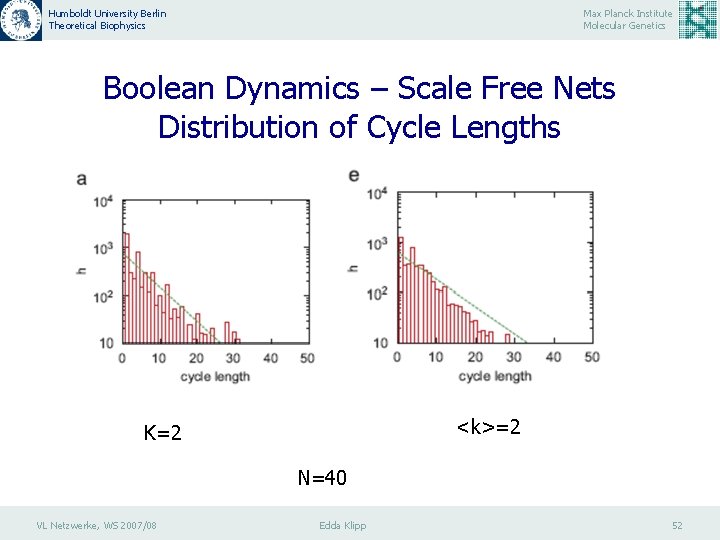
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Dynamics – Scale Free Nets Distribution of Cycle Lengths <k>=2 K=2 N=40 VL Netzwerke, WS 2007/08 Edda Klipp 52
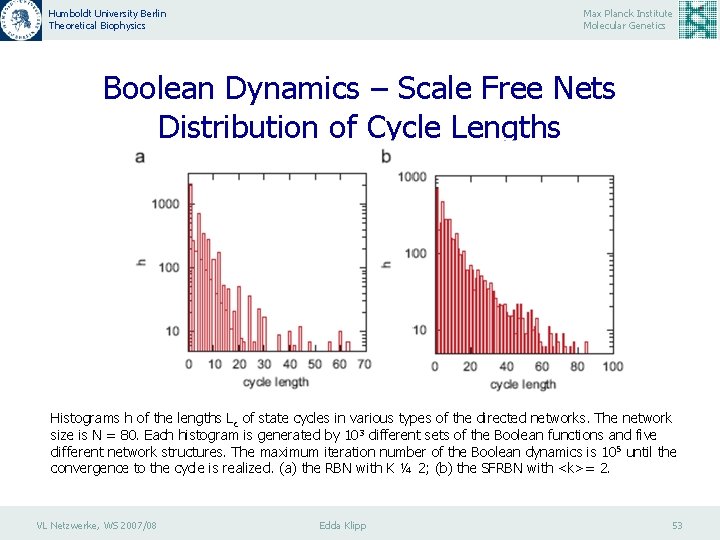
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Dynamics – Scale Free Nets Distribution of Cycle Lengths Histograms h of the lengths Lc of state cycles in various types of the directed networks. The network size is N = 80. Each histogram is generated by 103 different sets of the Boolean functions and five different network structures. The maximum iteration number of the Boolean dynamics is 10 5 until the convergence to the cycle is realized. (a) the RBN with K ¼ 2; (b) the SFRBN with <k>= 2. VL Netzwerke, WS 2007/08 Edda Klipp 53
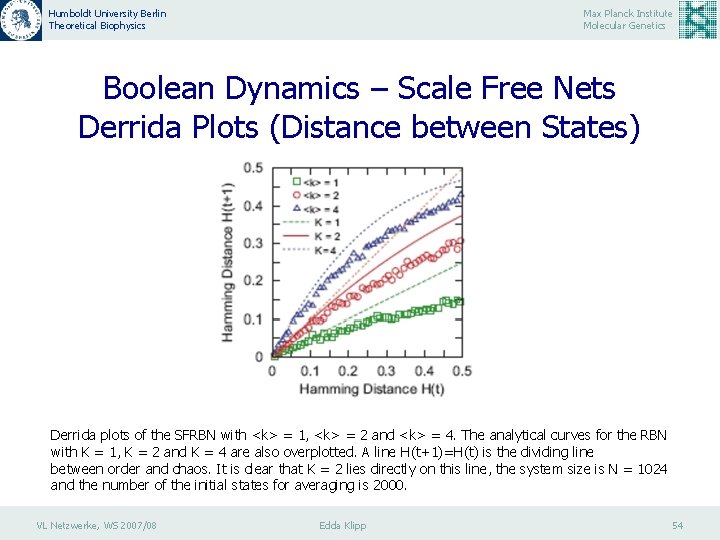
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Dynamics – Scale Free Nets Derrida Plots (Distance between States) Derrida plots of the SFRBN with <k> = 1, <k> = 2 and <k> = 4. The analytical curves for the RBN with K = 1, K = 2 and K = 4 are also overplotted. A line H(t+1)=H(t) is the dividing line between order and chaos. It is clear that K = 2 lies directly on this line, the system size is N = 1024 and the number of the initial states for averaging is 2000. VL Netzwerke, WS 2007/08 Edda Klipp 54
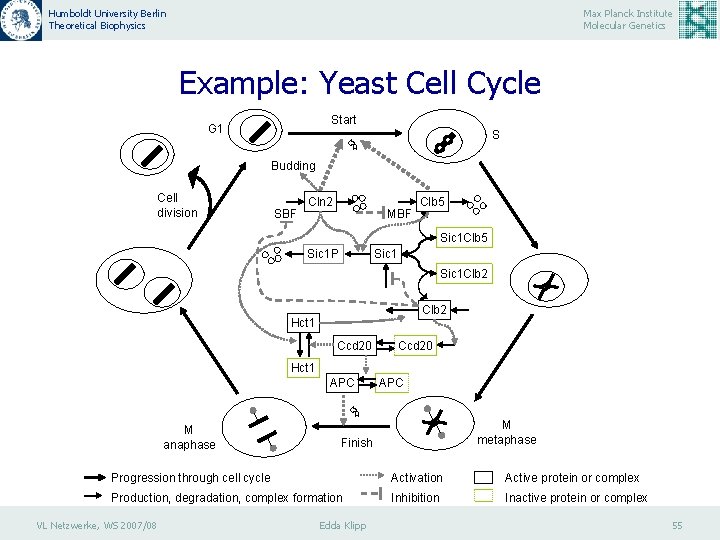
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Example: Yeast Cell Cycle Start G 1 S Budding Cell division SBF Cln 2 MBF Clb 5 Sic 1 P Sic 1 Clb 2 Hct 1 Ccd 20 Hct 1 APC M anaphase M metaphase Finish Progression through cell cycle Activation Active protein or complex Production, degradation, complex formation Inhibition Inactive protein or complex VL Netzwerke, WS 2007/08 Edda Klipp 55
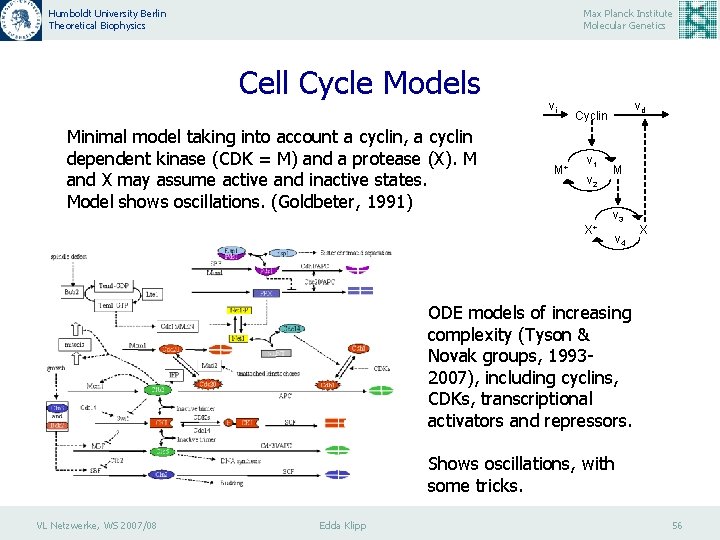
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Cell Cycle Models Minimal model taking into account a cyclin, a cyclin dependent kinase (CDK = M) and a protease (X). M and X may assume active and inactive states. Model shows oscillations. (Goldbeter, 1991) vi M+ vd Cyclin v 1 v 2 X+ M v 3 v 4 X ODE models of increasing complexity (Tyson & Novak groups, 19932007), including cyclins, CDKs, transcriptional activators and repressors. Shows oscillations, with some tricks. VL Netzwerke, WS 2007/08 Edda Klipp 56

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Yeast Cell Cycle – Data VL Netzwerke, WS 2007/08 Edda Klipp 57
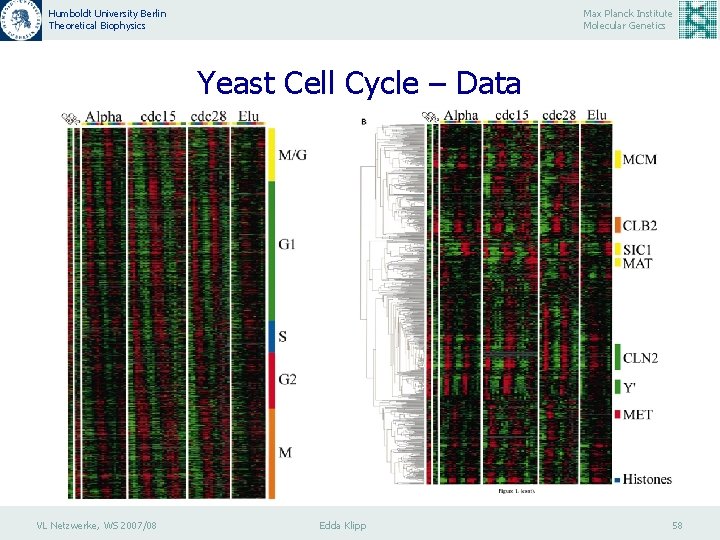
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Yeast Cell Cycle – Data VL Netzwerke, WS 2007/08 Edda Klipp 58

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Yeast Cell Cycle – Model VL Netzwerke, WS 2007/08 Edda Klipp 59
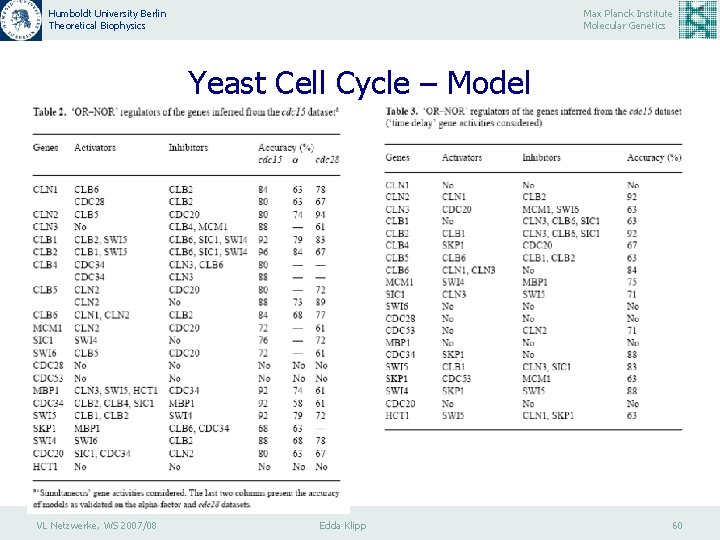
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Yeast Cell Cycle – Model VL Netzwerke, WS 2007/08 Edda Klipp 60
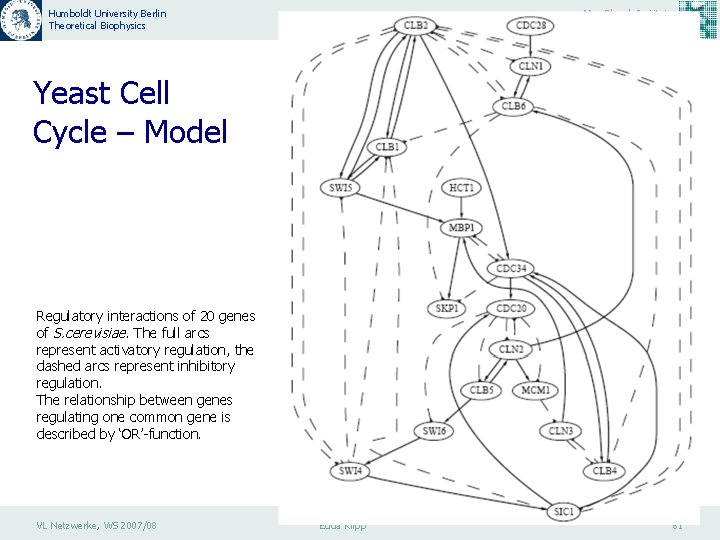
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Yeast Cell Cycle – Model Regulatory interactions of 20 genes of S. cerevisiae. The full arcs represent activatory regulation, the dashed arcs represent inhibitory regulation. The relationship between genes regulating one common gene is described by ‘OR’-function. VL Netzwerke, WS 2007/08 Edda Klipp 61

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Identification Given: Experimental data Demanded: Network connectivity and Boolean rules VL Netzwerke, WS 2007/08 Edda Klipp 62

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Identification VL Netzwerke, WS 2007/08 Edda Klipp 63

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Identification VL Netzwerke, WS 2007/08 Edda Klipp 64

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification Problem Formally, it is defined the identification problem. Relating to the identification problem, it is also defined the consistency problem, the counting problem and the enumeration problem. Let (Ij; Oj) be a pair of expression patterns of {v 1; … ; vn}, where Ij corresponds to the INPUT and Oj corresponds to the OUTPUT. We call the pair (Ij; Oj) an example. q A node vi in a Boolean network G(V; F) is consistent with an example (Ij; Oj) if Oj(vi) = fi(Ij(vi 1 ); … ; Ij(vik)) holds. q A Boolean network G(V; F) is consistent with (Ij; Oj) if all nodes are consistent with (Ij; Oj). q For a set of examples EX = {(I 1; O 1); (I 2; O 2); …; (Im; Om)}, network G(V; F) (resp. node vi) is consistent with EX if G(V; F) (resp. node vi) is consistent with all (Ij; Oj) for 1≤ j ≤ m. VL Netzwerke, WS 2007/08 Edda Klipp 65

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification Problem q CONSISTENCY: Given N (the number of nodes) and EX, decide whether or not there exists a Boolean network consistent with EX and output one if it exists; q COUNTING: Given N and EX, count the number of Boolean networks consistent with EX; q ENUMERATION: Given N and EX, output all the Boolean networks consistent with EX; q IDENTIFICATION: Given N and EX, decide whether or not there exists a unique Boolean network consistent with EX and output it if it exists. VL Netzwerke, WS 2007/08 Edda Klipp 66

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification Problem VL Netzwerke, WS 2007/08 Edda Klipp 67

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification: Algorithm for the consistency problem: The algorithm below is natural and conceptually very simple since it simply outputs Boolean functions consistent with given examples: (1) For each node vi V, execute STEP (2) If there exists a triplet (fi; vk; vh) satisfying Oj(vi) = fi(Ij(vk); Ij(vh)) for all j = 1; … ; m, output fi as a Boolean function assigned to vi and output vk; vh as input nodes to vi. In order to find a triplet (fi; vk; vh), we use a simple exhaustive search: for each pair of nodes (vk; vh) (k < h) and for each Boolean function f, we check whether or not Oj(vi) = f(Ij(vk); Ij(vh)) holds for all j. VL Netzwerke, WS 2007/08 Edda Klipp 68

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification: Algorithm For the enumeration problem: replace STEP (2) with the following: (2') Enumerate all triplets (fi; vk; vh) satisfying Oj(vi) = fi(Ij(vk); Ij(vh)) for all j = 1; … ; m. Then, any combination of triplets ((f 1; vk 1; vh 1); (f 2; vk 2; vh 2); … ; (fn; vkn; vhn)) can represent a consistent Boolean network. Of course, we carefully enumerate triplets since there exists more than two triplets which represent the same Boolean function (such as vk Λ vh and vh Λ vk). VL Netzwerke, WS 2007/08 Edda Klipp 69

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification: Algorithm For the counting problem: simply multiply the number of triplets consistent with each node. For the identification problem: replace STEP (2) with the following: (2") If there exists only one triplet (fi; vk; vh) satisfying Oj(vi) = fi(Ij(vk); Ij(vh)) for all j = 1; … ; m. output fi as a Boolean function assigned to vi and output vk; vh as input nodes to vi. VL Netzwerke, WS 2007/08 Edda Klipp 70

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Identification: Time Complexity There are Boolean functions with K input variables. There are (possible) combinations of input nodes per node. For each node, triplets are examined in the algorithm. For each triplets, m examples are examined. Therefore, pairs of Boolean functions and examples are examined in total. In order to examine one pair, O(K) time is required. Therefore, the algorithm works in time. Thus, the algorithm works in polynomial time for fixed K. Similarly, we can show that the algorithms for the counting problem and the identification problem work in polynomial time for fixed K. For any Boolean network of fixed K, O(log N) INPUT/OUTPUT pairs are sufficient with high probability. VL Netzwerke, WS 2007/08 Edda Klipp 71

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Identification: Reveal REVEAL. The results suggested that only a small number of state transition pairs (100 pairs from 1015) were sufficient for inferring Boolean networks with 50 nodes (genes) whose indegree (the number of input nodes to a node) was bounded by 3. VL Netzwerke, WS 2007/08 Edda Klipp 72

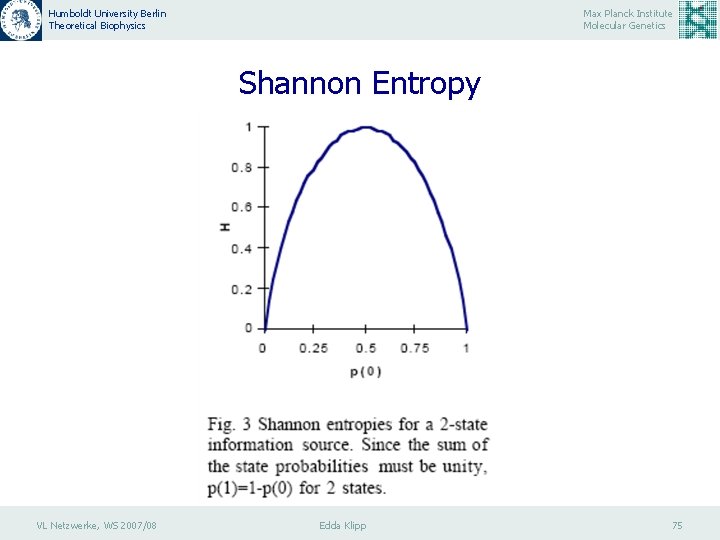
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Boolean Network Identification: Reveal Information theoretic principles of mutual information (M) analysis Information theory provides us with a quantitative information measure, the Shannon entropy, H. The Shannon entropy is defined in terms of the probability of observing a particular symbol or event, pi, within a given sequence (Shannon & Weaver, 1963), H= - S pi log pi. VL Netzwerke, WS 2007/08 Edda Klipp 73

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Shannon Entropy In a binary system, an element, X, may be in either of s=2 states, say on or off. Over a particular sequence of events, the sum of the probabilities of X being on, p(1) or off, p(0) must be equal to unity, therefore p(1)=1 -p(0), and H(X)=-p(0)*log[p(0)]-[1 -p(0)] *log[1 -p(0)]. H reaches its maximum when the on and off states are equiprobable, i. e. the system is using each information carrying state to its fullest possible extent. As one state becomes more probable than the other, H decreases - the system is becoming biased. In the limiting case, where one probability is unity (certainty) and the other(s) zero (impossibility), H is zero (no uncertainty - no freedom of choice - no information). The maximum entropy, Hmax, occurs when all states are equiprobable, i. e. p(0)=p(1) =1/2. Accordingly, Hmax=log(2). Entropies are commonly measured in “bits” (binary digits), when using the logarithm on base 2; e. g. Hmax=1 for a 2 state system. VL Netzwerke, WS 2007/08 Edda Klipp 74
Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Shannon Entropy VL Netzwerke, WS 2007/08 Edda Klipp 75

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Shannon Entropy Determination of H. a) Single element. Probabilities are calculated from frequency of on/off values of X and Y. VL Netzwerke, WS 2007/08 Edda Klipp 76

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Shannon Entropy: Co-Occurrence H(X)= - S pi log pi , H(Y)= - S pj log pj , and H(X, Y) = - S pi, j log pi, j Determination of H. b) Distribution of value pairs. H is calculated from the probabilities of co-occurrence. VL Netzwerke, WS 2007/08 Edda Klipp 77

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Conditional Entropies There are 2 conditional entropies which capture the relationship between the sequences of X and Y, H(X|Y) and H(Y|X). These are related as follows (Shannon & Weaver, 1963): H(X, Y) = H(Y|X) + H(X) = H(X|Y) + H(Y). In words, the uncertainty of X and the remaining uncertainty of Y given knowledge of X, H(Y|X), i. e. the information contained in Y that is not shared with X, sum to the entropy of the combination of X and Y. VL Netzwerke, WS 2007/08 Edda Klipp 78

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Mutual Information “Mutual information”, M(X, Y), also referred to as “rate of transmission” between an input/output channel pair (Shannon & Weaver, 1963) is defined as: M(X, Y) = H(Y) - H(Y|X) = H(X) - H(X|Y). The shared information between X and Y corresponds to the remaining information of X if we remove the information of X that is not shared with Y. Using the above equations, mutual information can be defined directly in terms of the original entropies; this formulation will be important for the considerations below: M(X, Y) = H(X) + H(Y) - H(X, Y). VL Netzwerke, WS 2007/08 Edda Klipp 79

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics Mutual Information Venn diagrams of information relationships. In each case, add the shaded portions of both squares to determine of the following: [H(X)+H(Y)], H(X, Y), and M(X, Y). The small corner rectangles represent information that X and Y have in common. H(Y) is shown smaller than H(X) and with the corner rectangle on the left instead of the right to indicate that X and Y are different, although they have some mutual information. VL Netzwerke, WS 2007/08 Edda Klipp 80

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics REVEAL Algorithm VL Netzwerke, WS 2007/08 Edda Klipp 81

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics REVEAL Algorithm 1. Identification of perfect input-output state pairs of connectivity k=1 Compute the mutual information of all input-output state vector pairs. The calculation of the mutual information values reveals that H(at+1; at)=H(at+1), i. e. at uniquely determines. Likewise, H(dt+1; ct)=H(dt+1), i. e. ct uniquely determines dt+1. For all other genes there is no perfect match. 2. Determination of the rules for the identified pairs at k=1. We retrieve the rules a(t+1)=a(t) and d(t+1)=not c(t) by the respective rule tables. 3. Identification of perfect input-output state pairs of connectivity k=2 If not all rules can be retrieved by k=1 we consider k=2, by comparing the output state vectors of the remaining genes all possible pairs of input state vectors. The calculation gives H(bt+1; ct, dt)=H(bt+1), i. e. the pair ct, dt determines bt+1. Likewise, H(ct+1; at, dt)=H(bt+1), i. e. the pair at, dt determines ct+1. 4. Determination of the rules for the identified pairs at k=2 We retrieve the rules b(t+1)= (not c(t)) and d(t) and likewise c(t+1)=a(t) and b(t). 5. Identification of perfect input-output state pairs of connectivity k=p 6. Determination of the rules for the identified pairs at k=p. Stop, if all genes have been assigned a rule, otherwise increment p and go to 5. VL Netzwerke, WS 2007/08 Edda Klipp 82

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics REVEAL Algorithm: Other example VL Netzwerke, WS 2007/08 Edda Klipp 83

Humboldt University Berlin Theoretical Biophysics Max Planck Institute Molecular Genetics REVEAL Algorithm: Other example VL Netzwerke, WS 2007/08 Edda Klipp 84
- Slides: 84