Human Genome Analysis Slides freely downloadable from Lectures

(Human Genome Analysis) Slides freely downloadable from Lectures. Gerstein. Lab. org & “tweetable” (via @markgerstein). See last slide for more info. 1 1 - Mark Gerstein, Yale (c) Mark Gerstein, 2002, Lectures. Gerstein. Lab. org Yale, bioinfo. mbb. yale. edu Large-scale Transcriptome Mining: Building Interpretative Models while Protecting Individual Privacy
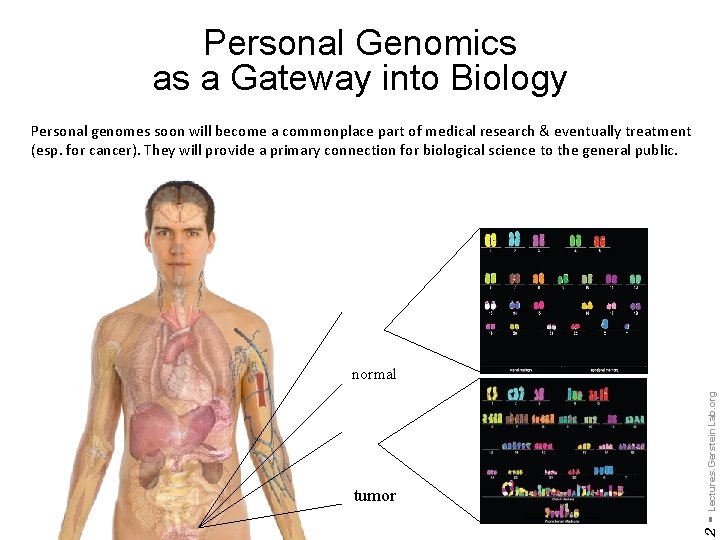
Personal Genomics as a Gateway into Biology Personal genomes soon will become a commonplace part of medical research & eventually treatment (esp. for cancer). They will provide a primary connection for biological science to the general public. 2 - tumor Lectures. Gerstein. Lab. org normal
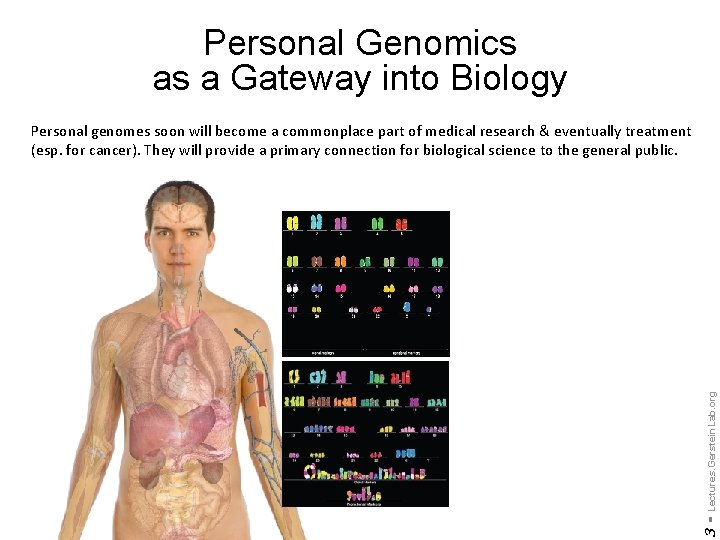
Personal Genomics as a Gateway into Biology 3 - Lectures. Gerstein. Lab. org Personal genomes soon will become a commonplace part of medical research & eventually treatment (esp. for cancer). They will provide a primary connection for biological science to the general public.
4 - Lectures. Gerstein. Lab. org
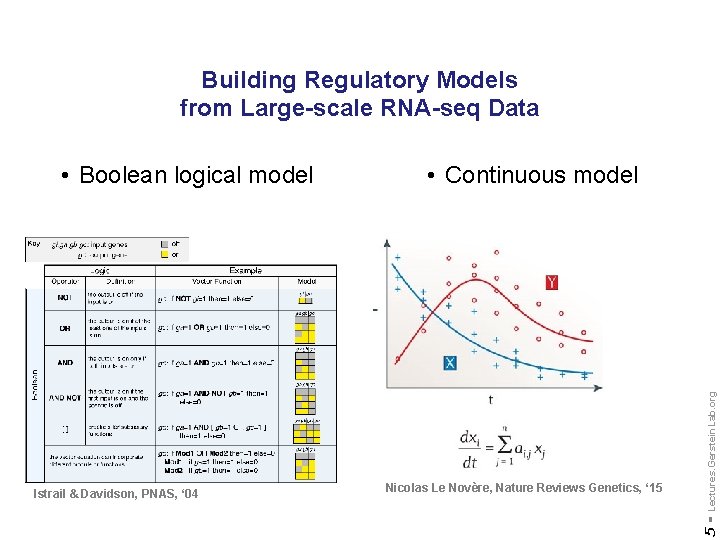
Building Regulatory Models from Large-scale RNA-seq Data Nicolas Le Novère, Nature Reviews Genetics, ‘ 15 Lectures. Gerstein. Lab. org Istrail & Davidson, PNAS, ‘ 04 • Continuous model 5 - • Boolean logical model

Privacy Aspects of Large-scale RNA-seq Analysis 6 - Lectures. Gerstein. Lab. org • Large magnitude of RNA-seq data generated - ENCODE, mod. ENCODE, TCGA, GTEx, Roadmap, psych. ENCODE, etc. • Mostly the data is about the phenotype (e. g. , cancer gene expression), but the individual information often comes along as collateral - Maybe we can separate private info but couple it with the public presentation?

• The Dilemma of Genomic Privacy Lectures. Gerstein. Lab. org - Fundamental, inherited info that’s very private vs the need for large-scale data • Large-scale Mining of RNA-sharing to enable med. research - Current Social & Tech Approaches seq to Determine State • Issues: burdensome security, Space Models inconsistencies + ways the solutions - Using dimensionality reduction to have been partially "hacked” help determine internal & external • Strawman Hybrid Soc-Tech drivers Proposal (Cloud Enclaves. - Decoupling expression changes into Quantifying Leaks, & Closely those driven by worm-fly conserved Coupled priv. -public data) genes vs species-specific ones. Also, Conserved genes have similar RNA-seq: How to Publicly canonical patterns (i. PDPs) in Share Some of it contrast to species specific ones (Ex - Removing SNVs in reads using MRF of ribosomal v signaling genes) - Quantifying & removing variant info - In human cell cycle, only conserved from expression levels + e. QTLs genes show matching periodic - Linking Attack using extreme pattern expression levels 7 - • Large-scale Transcriptome Mining: Building Interpretative Models while Protecting Individual Privacy

The Conundrum of Genomic Privacy: Is it a Problem? Yes Genetic Exceptionalism : genome is potentially very revealing about one’s identity & characteristics Shifting societal foci No one really cares about your genes You might not care [Klitzman & Sweeney ('11), J Genet Couns 20: 98 l; Greenbaum & Gerstein ('09), New Sci. (Sep 23) ] 8 - No Lectures. Gerstein. Lab. org • Most discussion of Identification Risk but what about Characterization Risk? - Finding you were in study X vs identifying that you have trait Y from studying your identified genome

Tricky Privacy Considerations in Personal Genomics - Could your genetic data give rise to a product line? • Culture Clash: Genomics historically has been a proponent of “open data” but not clear personal genomics fits this • Ethically challenged history of genetics [D Greenbaum & M Gerstein (’ 08). Am J. Bioethics; D Greenbaum & M Gerstein, Hartford Courant, 10 Jul. '08 ; SF Chronicle, 2 Nov. '08; Greenbaum et al. PLOS CB (‘ 11) ; Greenbaum & Gerstein ('13), The Scientist; Photo from NY Times] Lectures. Gerstein. Lab. org - Genomic sequence very revealing about one’s children. Is true consent possible? - Once put on the web it can’t be taken back • Ownership of the data & what consent means (Hela) 9 - • Personal Genomic info. essentially meaningless currently but will it be in 20 yrs? 50 yrs?

The Other Side of the Coin: Why we should share 10 - [Yale Law Roundtable (‘ 10). Comp. in Sci. & Eng. 12: 8; D Greenbaum & M Gerstein (‘ 09). Am. J. Bioethics; D Greenbaum & M Gerstein (‘ 10). SF Chronicle, May 2, Page E-4; Greenbaum et al. PLOS CB (‘ 11)] Lectures. Gerstein. Lab. org • Sharing helps speed research - Large-scale mining of this information is important for medical research - Privacy is cumbersome, particularly for big data - Sharing is important for reproducible research • Sharing is useful for education

The Dilemma 11 - • What is acceptable risk? What is acceptable data leakage? Can we quantify leakage? • Cost Benefit Analysis: how helpful is identifiable data in genomic research v. potential harm from a breach? • The individual (harmed? ) v the collective (benefits) - But do sick patients care about their privacy? • Maybe a we need a few "test pilots” (ala PGP)? - Sports stars & celebrities? Lectures. Gerstein. Lab. org [Economist, 15 Aug ‘ 15]

[Seringhaus & Gerstein ('09), Hart. Courant (Jun 5); Greenbaum & Gerstein ('11), NY Times (6 Oct)] 12 - • Sharing & "peer-production" is central to success of many new ventures, with the same risks as in genomics • We confront privacy risks every day we access the internet • (. . . or is the genome more exceptional & fundamental? ) Lectures. Gerstein. Lab. org Genomics has similar "Big Data" Dilemma in the Rest of Society

Current Social & Technical Solutions • Consents • “Protected” distribution of data (db. GAP) • Local computes on secure computer • Issues - Non-uniformity of consents & paperwork [Greenbuam et al ('04), Nat. Biotech; Greenbaum & Gerstein ('13), The Scientist] 13 - - Encryption & computer security creates burdensome requirements on data sharing & large scale analysis - Many schemes get “hacked” Lectures. Gerstein. Lab. org • Different international norms, leading to confusion

Privacy Hacks • Personalized genomic data generation is booming • “Detection of genome in a mixture” - Individuals give consent to participate but request anonymity 14 - • Larger and more datasets leads to more realistic risks of linking attacks, that may be much more damaging than detection of genome in a mixture attacks • Main focus is on protecting variants Lectures. Gerstein. Lab. org • HAPMAP, Personal genome project, 1000 Genomes…

Lectures. Gerstein. Lab. org 15 - Cross correlated small set of identifiable IMDB movie database rating records with large set of “anonymized” Netflix customer ratings

Strawman Hybrid Social & Tech Proposed Solution? • Technology to make things easier - Cloud computing & enclaves (eg solution of Genomics England) • Technological barriers shouldn't create a social incentive for “hacking” - Lightweight, freely accessible secondary datasets coupled to underlying variants - Selection of stub & "test pilot" datasets for benchmarking - Develop programs on public stubs on your laptop, then move the program to the cloud for private production run [D Greenbaum, M Gerstein (‘ 11). Am J Bioeth 11: 39. Greenbaum & Gerstein, The Scientist ('13)] Lectures. Gerstein. Lab. org - Genetic Licensure & training for individuals (similar to medical license, drivers license) • Quantifying Leakage & allowing a small amounts of it (eg photos of eye color) • Careful separation & coupling of private & public data 16 - • Fundamentally, researchers have to keep genetic secrets

• The Dilemma of Genomic Privacy Lectures. Gerstein. Lab. org - Fundamental, inherited info that’s very private vs the need for large-scale data • Large-scale Mining of RNA-sharing to enable med. research - Current Social & Tech Approaches seq to Determine State • Issues: burdensome security, Space Models inconsistencies + ways the solutions - Using dimensionality reduction to have been partially "hacked” help determine internal & external • Strawman Hybrid Soc-Tech drivers Proposal (Cloud Enclaves. - Decoupling expression changes into Quantifying Leaks, & Closely those driven by worm-fly conserved Coupled priv. -public data) genes vs species-specific ones. Also, Conserved genes have similar RNA-seq: How to Publicly canonical patterns (i. PDPs) in Share Some of it contrast to species specific ones (Ex - Removing SNVs in reads using MRF of ribosomal v signaling genes) - Quantifying & removing variant info - In human cell cycle, only conserved from expression levels + e. QTLs genes show matching periodic - Linking Attack using extreme pattern expression levels 17 - • Large-scale Transcriptome Mining: Building Interpretative Models while Protecting Individual Privacy
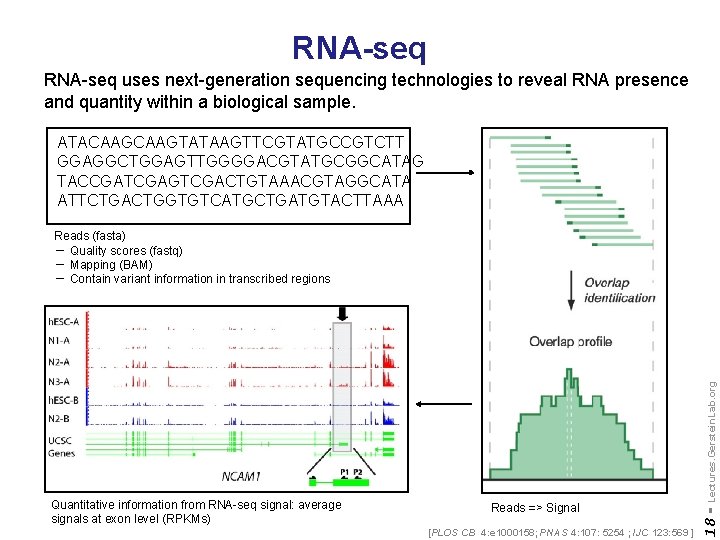
RNA-seq uses next-generation sequencing technologies to reveal RNA presence and quantity within a biological sample. ATACAAGTATAAGTTCGTATGCCGTCTT GGAGGCTGGAGTTGGGGACGTATGCGGCATAG TACCGATCGAGTCGACTGTAAACGTAGGCATA ATTCTGACTGGTGTCATGCTGATGTACTTAAA Reads => Signal [PLOS CB 4: e 1000158; PNAS 4: 107: 5254 ; IJC 123: 569 ] 18 - Quantitative information from RNA-seq signal: average signals at exon level (RPKMs) Lectures. Gerstein. Lab. org Reads (fasta) - Quality scores (fastq) - Mapping (BAM) - Contain variant information in transcribed regions
![Light-weight formats Mapping coordinates without variants (MRF) 19 - [Bioinformatics 27: 281] Reads (linked Light-weight formats Mapping coordinates without variants (MRF) 19 - [Bioinformatics 27: 281] Reads (linked](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-19.jpg)
Light-weight formats Mapping coordinates without variants (MRF) 19 - [Bioinformatics 27: 281] Reads (linked via ID, 10 X larger than mapping coord. ) Lectures. Gerstein. Lab. org • Some lightweight format clearly separate public & private info. , aiding exchange • Files become much smaller • Distinction between formats to compute on and those to archive with – become sharper with big data
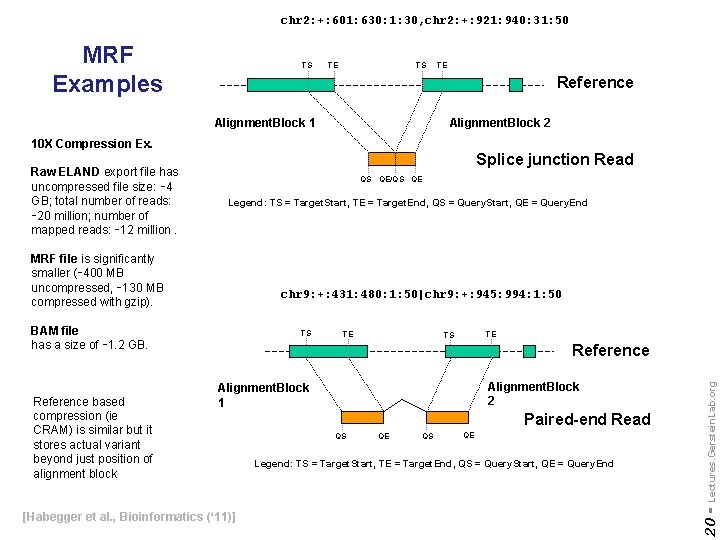
chr 2: +: 601: 630: 1: 30, chr 2: +: 921: 940: 31: 50 MRF Examples TS TE Reference Alignment. Block 1 Alignment. Block 2 10 X Compression Ex. Splice junction Read QS QE/QS QE Legend: TS = Target. Start, TE = Target. End, QS = Query. Start, QE = Query. End MRF file is significantly smaller (∼ 400 MB uncompressed, ∼ 130 MB compressed with gzip). chr 9: +: 431: 480: 1: 50|chr 9: +: 945: 994: 1: 50 BAM file has a size of ∼ 1. 2 GB. TE TE TS Reference Alignment. Block 2 Alignment. Block 1 [Habegger et al. , Bioinformatics (‘ 11)] Paired-end Read QS QE Legend: TS = Target. Start, TE = Target. End, QS = Query. Start, QE = Query. End Lectures. Gerstein. Lab. org Reference based compression (ie CRAM) is similar but it stores actual variant beyond just position of alignment block TS 20 - Raw ELAND export file has uncompressed file size: ∼ 4 GB; total number of reads: ∼ 20 million; number of mapped reads: ∼ 12 million.

• The Dilemma of Genomic Privacy Lectures. Gerstein. Lab. org - Fundamental, inherited info that’s very private vs the need for large-scale data • Large-scale Mining of RNA-sharing to enable med. research - Current Social & Tech Approaches seq to Determine State • Issues: burdensome security, Space Models inconsistencies + ways the solutions - Using dimensionality reduction to have been partially "hacked” help determine internal & external • Strawman Hybrid Soc-Tech drivers Proposal (Cloud Enclaves. - Decoupling expression changes into Quantifying Leaks, & Closely those driven by worm-fly conserved Coupled priv. -public data) genes vs species-specific ones. Also, Conserved genes have similar RNA-seq: How to Publicly canonical patterns (i. PDPs) in Share Some of it contrast to species specific ones (Ex - Removing SNVs in reads using MRF of ribosomal v signaling genes) - Quantifying & removing variant info - In human cell cycle, only conserved from expression levels + e. QTLs genes show matching periodic - Linking Attack using extreme pattern expression levels 21 - • Large-scale Transcriptome Mining: Building Interpretative Models while Protecting Individual Privacy
![22 - [Biometrics 68(1) 1– 11] • e. QTLs are genomic loci that contribute 22 - [Biometrics 68(1) 1– 11] • e. QTLs are genomic loci that contribute](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-22.jpg)
22 - [Biometrics 68(1) 1– 11] • e. QTLs are genomic loci that contribute to variation in m. RNA expression levels • e. QTLs provide insights on transcription regulation, and the molecular basis of phenotypic outcomes • e. QTL mapping can be done with RNA-Seq data Lectures. Gerstein. Lab. org e. QTL Mapping Using RNA-Seq Data
![Information Content and Predictability [Harmanciet al. Nat. Meth. (in revision)] Information Content and Predictability [Harmanciet al. Nat. Meth. (in revision)]](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-23.jpg)
Information Content and Predictability [Harmanciet al. Nat. Meth. (in revision)]

Representative Expression, Genotype, e. QTL Datasets • m. RNA sequencing for 462 individuals • Publicly available. Quantification for protein coding genes • Approximately 3, 000 cis-e. QTL (FDR<0. 05) • Genotypes are available from the 1000 Genomes Project
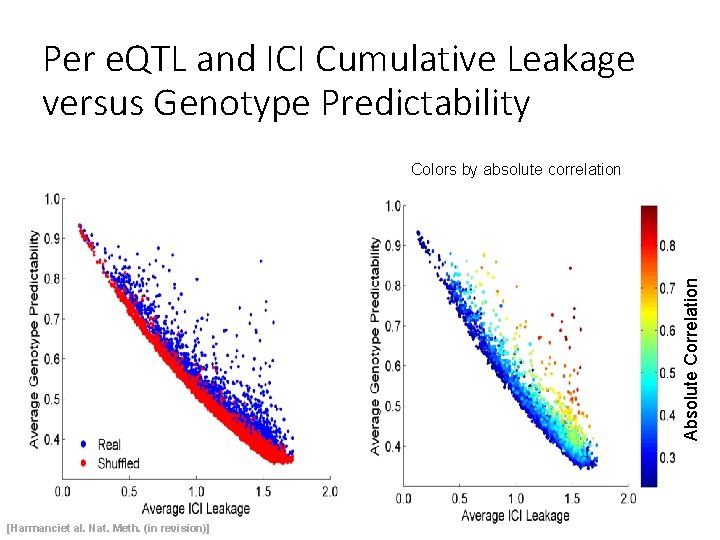
Per e. QTL and ICI Cumulative Leakage versus Genotype Predictability Absolute Correlation Colors by absolute correlation [Harmanciet al. Nat. Meth. (in revision)]
![Cumulative Leakage versus Joint Predictability [Harmanciet al. Nat. Meth. (in revision)] Cumulative Leakage versus Joint Predictability [Harmanciet al. Nat. Meth. (in revision)]](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-26.jpg)
Cumulative Leakage versus Joint Predictability [Harmanciet al. Nat. Meth. (in revision)]
![Linking Attack Scenario [Harmanciet al. Nat. Meth. (in revision)] Linking Attack Scenario [Harmanciet al. Nat. Meth. (in revision)]](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-27.jpg)
Linking Attack Scenario [Harmanciet al. Nat. Meth. (in revision)]
![Steps in Instantiation of a (Mock) Linking Attack [Harmanciet al. Nat. Meth. (in revision)] Steps in Instantiation of a (Mock) Linking Attack [Harmanciet al. Nat. Meth. (in revision)]](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-28.jpg)
Steps in Instantiation of a (Mock) Linking Attack [Harmanciet al. Nat. Meth. (in revision)]
![[Harmanciet al. Nat. Meth. (in revision)] [Harmanciet al. Nat. Meth. (in revision)]](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-29.jpg)
[Harmanciet al. Nat. Meth. (in revision)]
![Extremity based linking with homozygous genotypes [Harmanciet al. Nat. Meth. (in revision)] Attacker can Extremity based linking with homozygous genotypes [Harmanciet al. Nat. Meth. (in revision)] Attacker can](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-30.jpg)
Extremity based linking with homozygous genotypes [Harmanciet al. Nat. Meth. (in revision)] Attacker can estimate the reliability of linkings Sensitivity: Fraction of correctly linked Individuals among all individuals PPV: Fraction of correctly linked individuals among selected individuals

• The Dilemma of Genomic Privacy Lectures. Gerstein. Lab. org - Fundamental, inherited info that’s very private vs the need for large-scale data • Large-scale Mining of RNA-sharing to enable med. research - Current Social & Tech Approaches seq to Determine State • Issues: burdensome security, Space Models inconsistencies + ways the solutions - Using dimensionality reduction to have been partially "hacked” help determine internal & external • Strawman Hybrid Soc-Tech drivers Proposal (Cloud Enclaves. - Decoupling expression changes into Quantifying Leaks, & Closely those driven by worm-fly conserved Coupled priv. -public data) genes vs species-specific ones. Also, Conserved genes have similar RNA-seq: How to Publicly canonical patterns (i. PDPs) in Share Some of it contrast to species specific ones (Ex - Removing SNVs in reads using MRF of ribosomal v signaling genes) - Quantifying & removing variant info - In human cell cycle, only conserved from expression levels + e. QTLs genes show matching periodic - Linking Attack using extreme pattern expression levels 31 - • Large-scale Transcriptome Mining: Building Interpretative Models while Protecting Individual Privacy
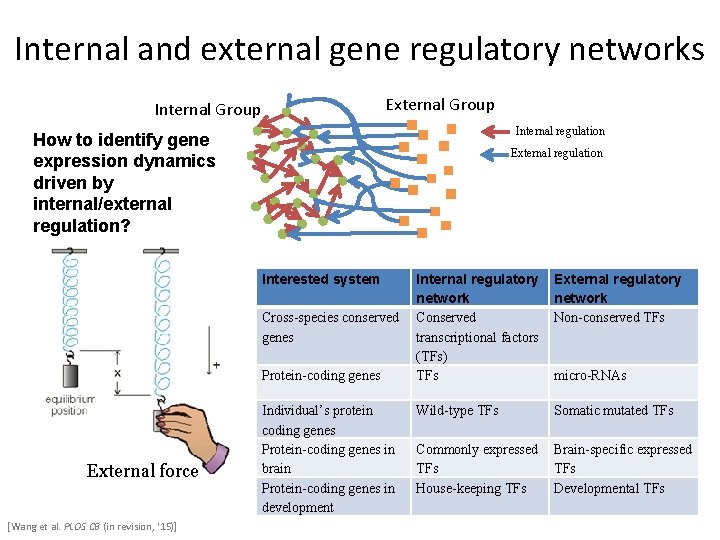
Internal and external gene regulatory networks External Group Internal regulation How to identify gene expression dynamics driven by internal/external regulation? External regulation Interested system Cross-species conserved genes Protein-coding genes External force [Wang et al. PLOS CB (in revision, ‘ 15)] Individual’s protein coding genes Protein-coding genes in brain Protein-coding genes in development Internal regulatory network Conserved transcriptional factors (TFs) TFs External regulatory network Non-conserved TFs Wild-type TFs Somatic mutated TFs Commonly expressed TFs House-keeping TFs Brain-specific expressed TFs Developmental TFs micro-RNAs
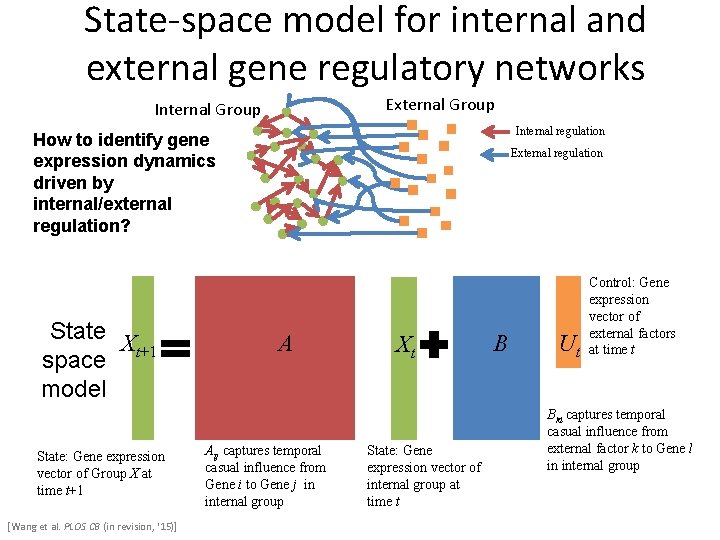
State-space model for internal and external gene regulatory networks External Group Internal regulation How to identify gene expression dynamics driven by internal/external regulation? State Xt+1 space model State: Gene expression vector of Group X at time t+1 [Wang et al. PLOS CB (in revision, ‘ 15)] External regulation A Aij captures temporal casual influence from Gene i to Gene j in internal group Xt State: Gene expression vector of internal group at time t B U Control: Gene expression vector of external factors t at time t Bkl captures temporal casual influence from external factor k to Gene l in internal group
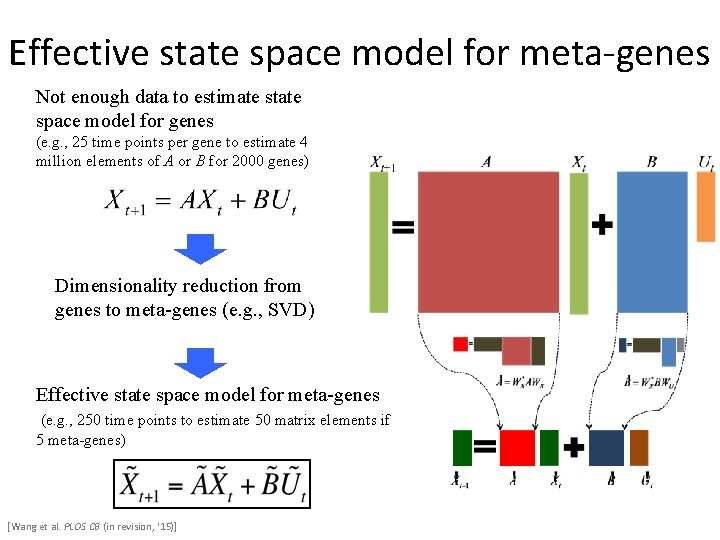
Effective state space model for meta-genes Not enough data to estimate state space model for genes (e. g. , 25 time points per gene to estimate 4 million elements of A or B for 2000 genes) Dimensionality reduction from genes to meta-genes (e. g. , SVD) Effective state space model for meta-genes (e. g. , 250 time points to estimate 50 matrix elements if 5 meta-genes) [Wang et al. PLOS CB (in revision, ‘ 15)]
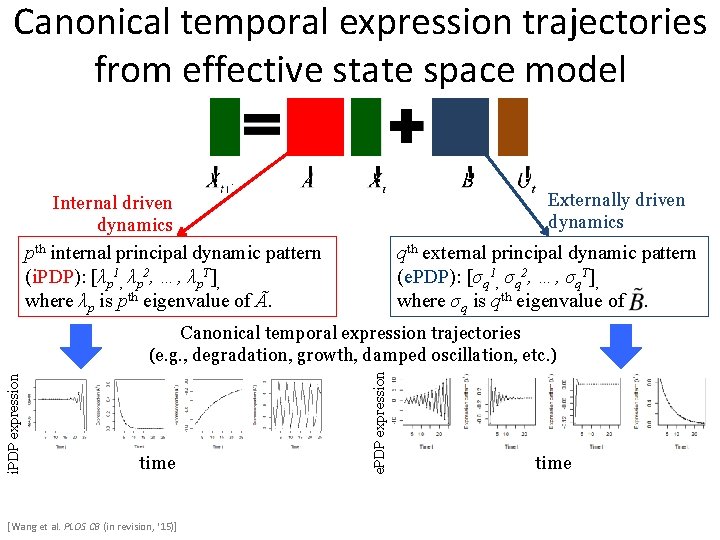
Canonical temporal expression trajectories from effective state space model Externally driven dynamics Internal driven dynamics pth internal principal dynamic pattern (i. PDP): [λp 1, λp 2, …, λp. T], where λp is pth eigenvalue of Ã. qth external principal dynamic pattern (e. PDP): [σq 1, σq 2, …, σq. T], where σq is qth eigenvalue of. time [Wang et al. PLOS CB (in revision, ‘ 15)] e. PDP expression i. PDP expression Canonical temporal expression trajectories (e. g. , degradation, growth, damped oscillation, etc. ) time
![[Wang et al. PLOS CB (in revision, ‘ 15)] A. Gene state-space model Flowchart [Wang et al. PLOS CB (in revision, ‘ 15)] A. Gene state-space model Flowchart](http://slidetodoc.com/presentation_image/5524a5648431822c49ad44f5d401e8df/image-36.jpg)
[Wang et al. PLOS CB (in revision, ‘ 15)] A. Gene state-space model Flowchart C. Meta-gene state-space model +c 3 +c 4 x. EXT=d 1 +d 3 Genes of U +c 2 time [λp 1, λp 2, …, λp. T] [σq 1, σq 2, …, σq. T] time +d 2 +d 4 D. Internal/External Principal Dynamic Patterns (PDPs) time … x. INT=c 1 Meta-genes of X E. Gene’s internal (INT) and external (EXT) driven expression dynamics composed of PDPs time Meta-genes of U Xt+1=AXt+BUt Genes of X B. Dimensionality Reduction / / / Internal regulation among genes/meta-genes Group X by A/Ã External regulation from genes/meta-genes in Group U to genes/meta-genes in Group X by B/ Genes/Meta-genes in Group X / Genes/Meta-genes in Group U
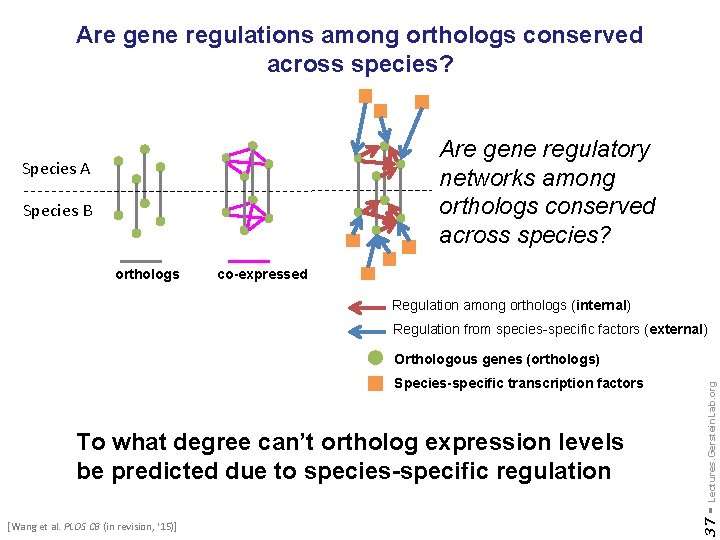
Are gene regulations among orthologs conserved across species? Are gene regulatory networks among orthologs conserved across species? Species A Species B orthologs co-expressed Regulation among orthologs (internal) Regulation from species-specific factors (external) To what degree can’t ortholog expression levels be predicted due to species-specific regulation [Wang et al. PLOS CB (in revision, ‘ 15)] 37 - Species-specific transcription factors Lectures. Gerstein. Lab. org Orthologous genes (orthologs)
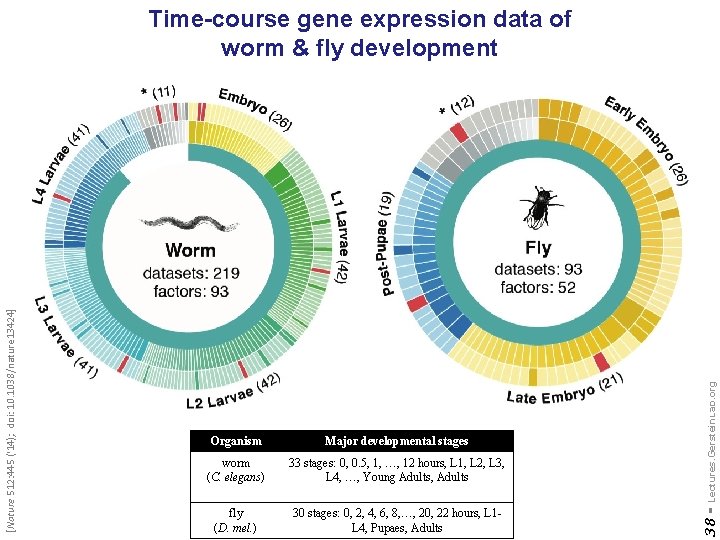
Major developmental stages worm (C. elegans) 33 stages: 0, 0. 5, 1, …, 12 hours, L 1, L 2, L 3, L 4, …, Young Adults, Adults fly (D. mel. ) 30 stages: 0, 2, 4, 6, 8, …, 20, 22 hours, L 1 L 4, Pupaes, Adults Lectures. Gerstein. Lab. org Organism 38 - [Nature 512: 445 ('14); doi: 10. 1038/nature 13424] Time-course gene expression data of worm & fly development
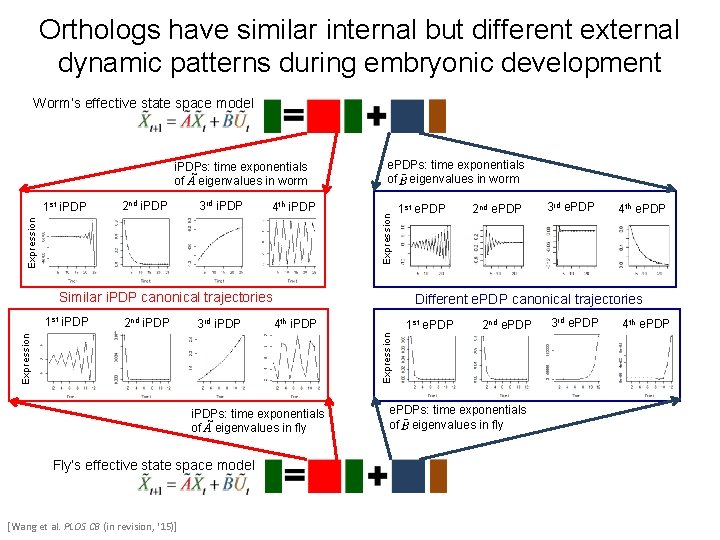
Orthologs have similar internal but different external dynamic patterns during embryonic development Worm’s effective state space model 2 nd i. PDP 3 rd i. PDP 4 th i. PDP Expression 1 st i. PDP e. PDPs: time exponentials of eigenvalues in worm Expression i. PDPs: time exponentials of à eigenvalues in worm Similar i. PDP canonical trajectories 2 nd i. PDP 3 rd i. PDP 2 nd e. PDP 4 th i. PDP Expression i. PDPs: time exponentials of à eigenvalues in fly Fly’s effective state space model [Wang et al. PLOS CB (in revision, ‘ 15)] 3 rd e. PDP 4 th e. PDP Different e. PDP canonical trajectories 1 st e. PDP 2 nd e. PDP Expression 1 st i. PDP 1 st e. PDPs: time exponentials of eigenvalues in fly 3 rd e. PDP 4 th e. PDP
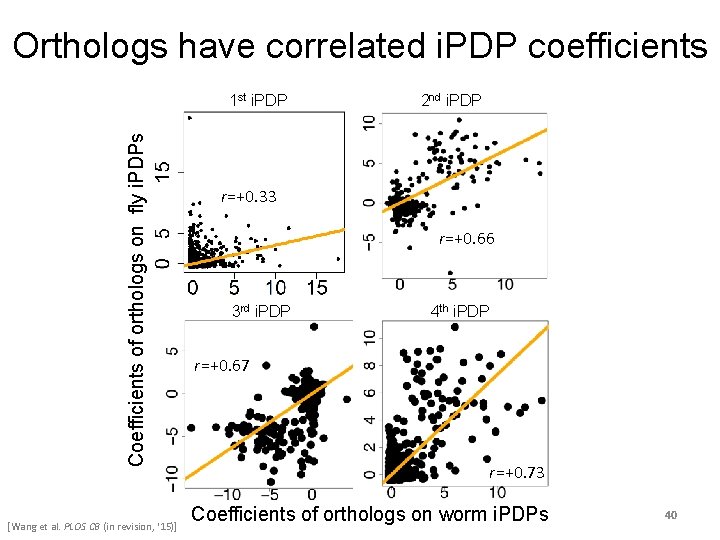
Orthologs have correlated i. PDP coefficients Coefficients of orthologs on fly i. PDPs 1 st i. PDP [Wang et al. PLOS CB (in revision, ‘ 15)] 2 nd i. PDP r=+0. 33 r=+0. 66 3 rd i. PDP 4 th i. PDP r=+0. 67 r=+0. 73 Coefficients of orthologs on worm i. PDPs 40
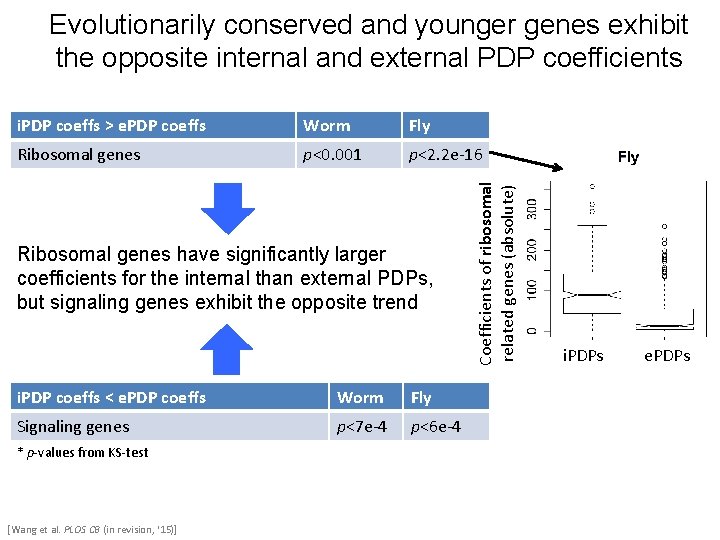
Evolutionarily conserved and younger genes exhibit the opposite internal and external PDP coefficients Worm Fly Ribosomal genes p<0. 001 p<2. 2 e-16 Ribosomal genes have significantly larger coefficients for the internal than external PDPs, but signaling genes exhibit the opposite trend i. PDP coeffs < e. PDP coeffs Worm Fly Signaling genes p<7 e-4 p<6 e-4 * p-values from KS-test [Wang et al. PLOS CB (in revision, ‘ 15)] Coefficients of ribosomal related genes (absolute) i. PDP coeffs > e. PDP coeffs i. PDPs e. PDPs
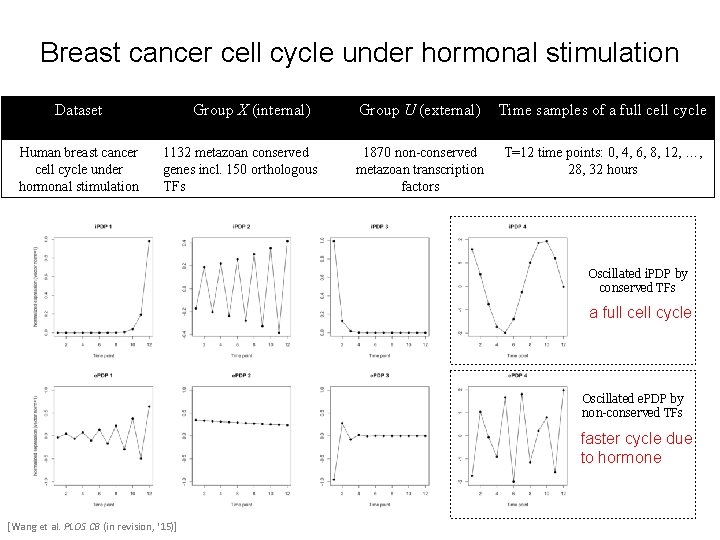
Breast cancer cell cycle under hormonal stimulation Dataset Human breast cancer cell cycle under hormonal stimulation Group X (internal) 1132 metazoan conserved genes incl. 150 orthologous TFs Group U (external) Time samples of a full cell cycle 1870 non-conserved metazoan transcription factors T=12 time points: 0, 4, 6, 8, 12, …, 28, 32 hours Oscillated i. PDP by conserved TFs a full cell cycle Oscillated e. PDP by non-conserved TFs faster cycle due to hormone [Wang et al. PLOS CB (in revision, ‘ 15)]

• The Dilemma of Genomic Privacy Lectures. Gerstein. Lab. org - Fundamental, inherited info that’s very private vs the need for large-scale data • Large-scale Mining of RNA-sharing to enable med. research - Current Social & Tech Approaches seq to Determine State • Issues: burdensome security, Space Models inconsistencies + ways the solutions - Using dimensionality reduction to have been partially "hacked” help determine internal & external • Strawman Hybrid Soc-Tech drivers Proposal (Cloud Enclaves. - Decoupling expression changes into Quantifying Leaks, & Closely those driven by worm-fly conserved Coupled priv. -public data) genes vs species-specific ones. Also, Conserved genes have similar RNA-seq: How to Publicly canonical patterns (i. PDPs) in Share Some of it contrast to species specific ones (Ex - Removing SNVs in reads using MRF of ribosomal v signaling genes) - Quantifying & removing variant info - In human cell cycle, only conserved from expression levels + e. QTLs genes show matching periodic - Linking Attack using extreme pattern expression levels 43 - • Large-scale Transcriptome Mining: Building Interpretative Models while Protecting Individual Privacy

Acknowledgements DREISS. gersteinlab. org D Wang, F He, S Maslov privacy papers. gersteinlab. org/subject/ D Greenbaum Priva. Seq. gersteinlab. org L Habegger, A Sboner, TA Gianoulis, J Rozowsky, A Agarwal, M Snyder Hiring Postdocs. See gersteinlab. org/jobs ! 44 - RSEQtools. gersteinlab. org Lectures. Gerstein. Lab. org A Harmanci
Default Theme • Default Outline Level 1 45 - Lectures. Gerstein. Lab. org - Level 2

More Information on this Talk SUBJECT: Networks DESCRIPTION: NOTES: This PPT should work on mac & PC. Paper references in the talk were mostly from Papers. Gerstein. Lab. org. PERMISSIONS: This Presentation is copyright Mark Gerstein, Yale University, 2010. Please read permissions statement at http: //www. gersteinlab. org/misc/permissions. html. Feel free to use images in the talk with PROPER acknowledgement (via citation to relevant papers or link to gersteinlab. org). 46 - Lectures. Gerstein. Lab. org PHOTOS & IMAGES. For thoughts on the source and permissions of many of the photos and clipped images in this presentation see http: //streams. gerstein. info. In particular, many of the images have particular EXIF tags, such as kwpotppt , that can be easily queried from flickr, viz: http: //www. flickr. com/photos/mbgmbg/tags/kwpotppt.
- Slides: 46