Hubba Hub Objects AnalyzerA Framework of Interactome Hubs

Hubba: Hub Objects Analyzer—A Framework of Interactome Hubs Identification for Network Biology C. Y. Lin, C. H. Chin, H. H. Wu, S. H. Chen, C. W. Ho, and M. T. Ko. Web Server Issue, NAR, 2008 -08 -14 Hsin-Hung Wu

Outline • Network Summary • What Does Hubba Do • Hubba Methods • Web Demo
USA Traffic Network
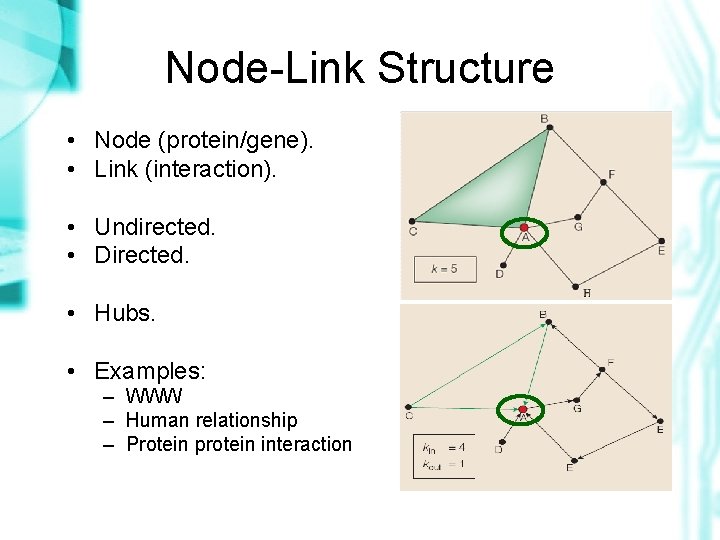
Node-Link Structure • Node (protein/gene). • Link (interaction). • Undirected. • Directed. • Hubs. • Examples: – WWW – Human relationship – Protein protein interaction
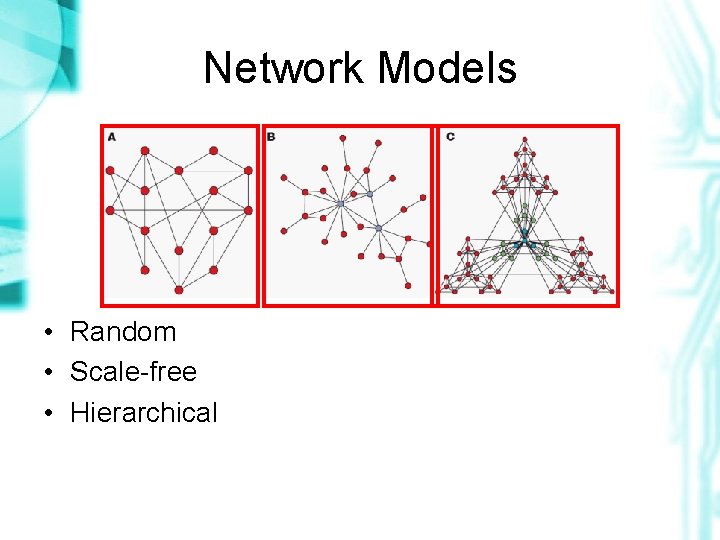
Network Models • Random • Scale-free • Hierarchical
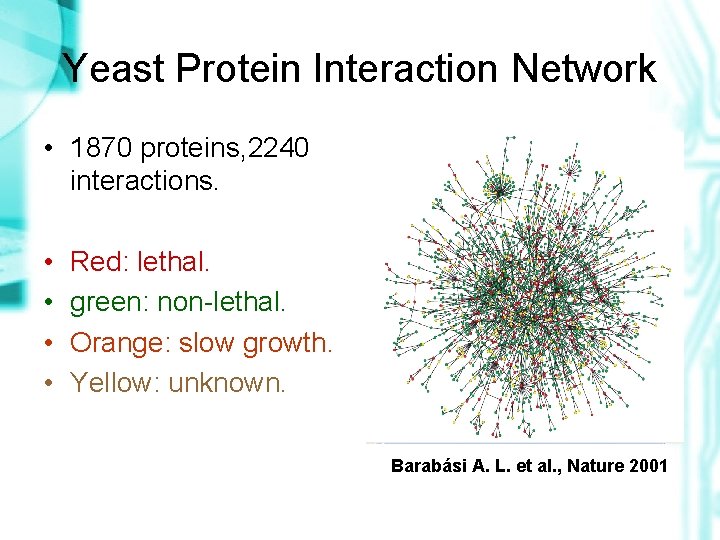
Yeast Protein Interaction Network • 1870 proteins, 2240 interactions. • • Red: lethal. green: non-lethal. Orange: slow growth. Yellow: unknown. Barabási A. L. et al. , Nature 2001

Modules in Biological Networks • Hub (Essential protein/gene); • Pathway; • Complex. • They are thought to be an important role. • Scientists are eager to decipher them in a biological network.
Hubba (http: //hub. iis. sinica. edu. tw/Hubba/)

What Does Hubba Do? • A web-based service. • It is designed to explore the essential nodes in a network. • Exploring the essential nodes by 6 methods. • Visualizing the resulted graphs in a user-friendly mode.

Hubba Methods • Degree • Bottle. Neck • Edge Percolation Component (EPC) • Subgraph Centrality (SC) • Maximum Neighborhood Component (MNC) • Density of Maximum Neighborhood Component (DMNC) • Double Screening Scheme (DSS)
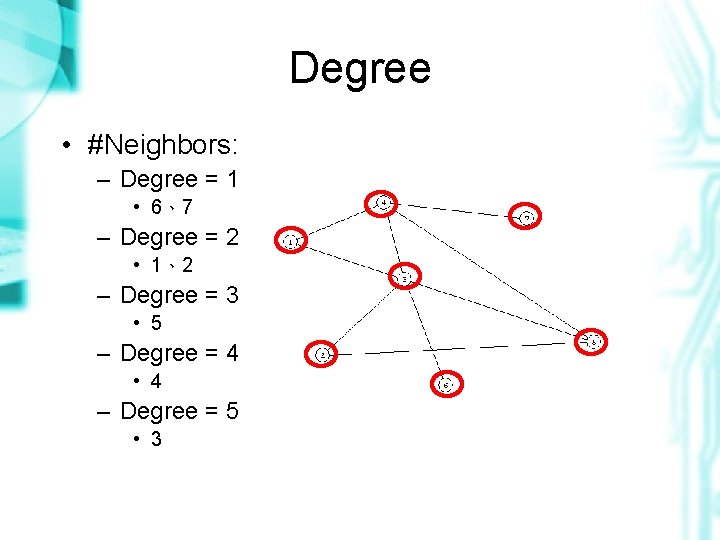
Degree • #Neighbors: – Degree = 1 • 6、7 – Degree = 2 • 1、2 – Degree = 3 • 5 – Degree = 4 • 4 – Degree = 5 • 3
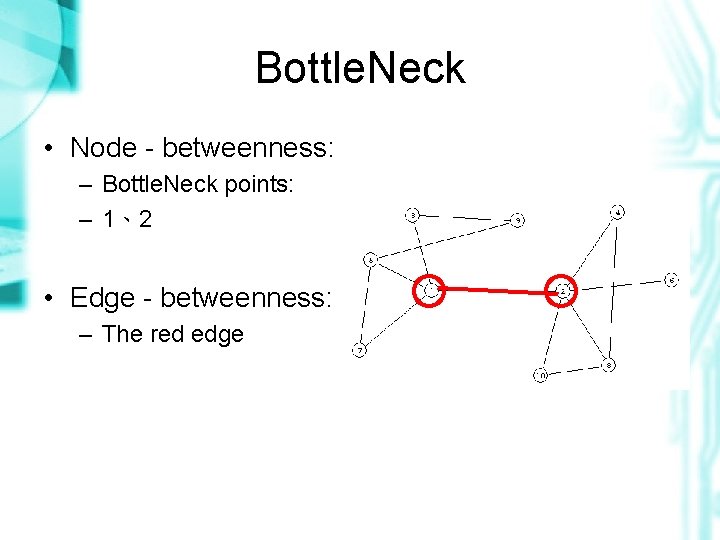
Bottle. Neck • Node - betweenness: – Bottle. Neck points: – 1、2 • Edge - betweenness: – The red edge

Edge Percolation Component (EPC) • The flow: – An original graph G – Let G’ be a realization of the random edge removing from G with a probability p. – If nodes v and w are connected in G’, set dvw be 1, otherwise set dvw be 0. – Repeat the upper procedure. – Define the Cvw the average of dvw over realizations. – The size of percolated component containing node v, Sv, is defined to be the sum of Cvw over nodes w. – The score of node v, EPC(v) , is defined to be Sv.
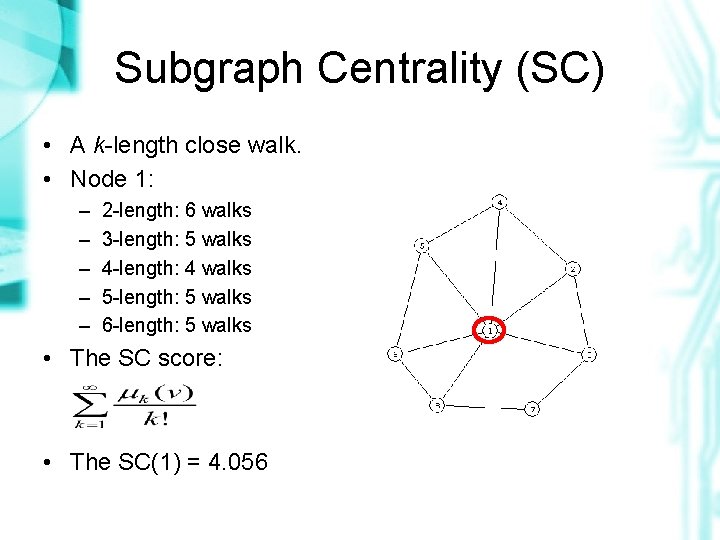
Subgraph Centrality (SC) • A k-length close walk. • Node 1: – – – 2 -length: 6 walks 3 -length: 5 walks 4 -length: 4 walks 5 -length: 5 walks 6 -length: 5 walks • The SC score: • The SC(1) = 4. 056

Maximum Neighborhood Component (MNC) • A node v (SRB 4), a subnetwork N(v). • The score of node v, MNC(v): – The size of the maximum connected component of N(v)

Density of Maximum Neighborhood Component (DMNC) • Node v • MNC(v) – Let N be the #node – E be the #edge • The score of node v, DMNC(v), is defined as:

Double Screening Scheme (DSS) • To reflect mixed characteristic of essential proteins with the above 6 methods. • For 2 scoring methods: – 2 n top ranked proteins by scoring method A are selected. – The 2 n proteins are ranked by scoring method B. – Output the n top ranked proteins.

DSS in Hubba • MNC and DMNC are selected: – Choose the MNC top rank group – Re-rank the selected nodes in DMNC. – Output the DSS result.
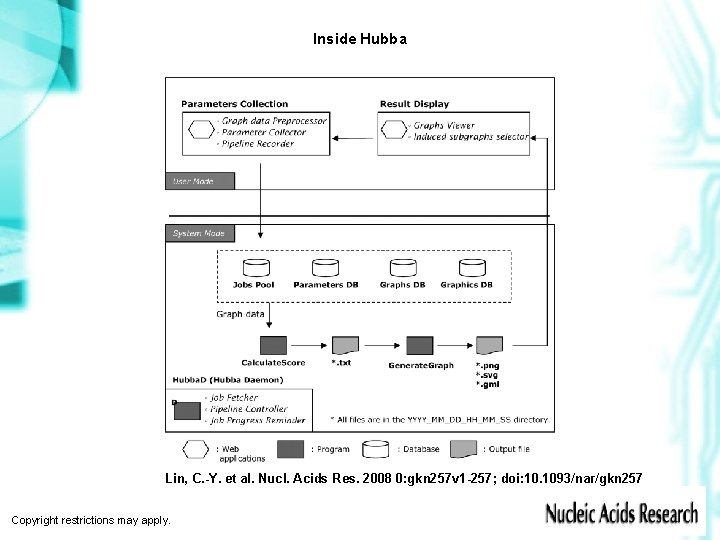
Inside Hubba Lin, C. -Y. et al. Nucl. Acids Res. 2008 0: gkn 257 v 1 -257; doi: 10. 1093/nar/gkn 257 Copyright restrictions may apply.
- Slides: 19