http www imgt org IMGT the global reference

http: //www. imgt. org IMGT, the global reference in immunogenetics and immunoinformatics http: //www. imgt. org Sofia Kossida IMGT Director Professor, Montpellier Universtity, IGH CNRS, Montpellier, France Marie-Paule Lefranc IMGT Founder and Director Emeritus Professor, Montpellier University, IGH CNRS, Montpellier, France February 10, 2015 Montpellier OMIS days
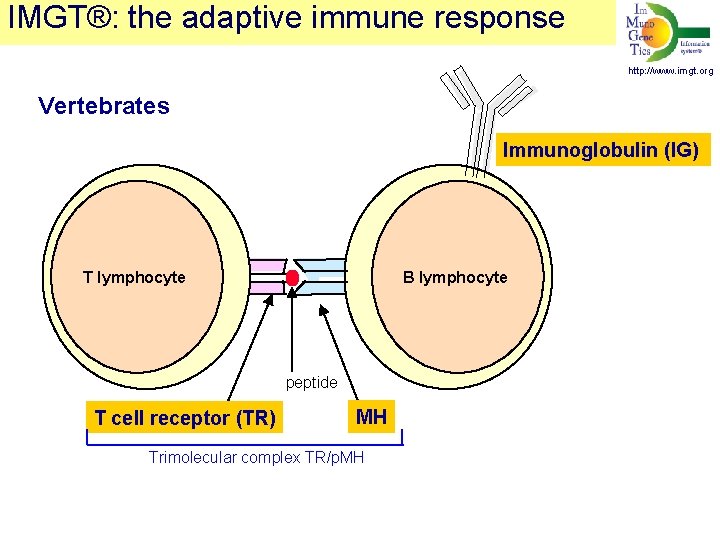
IMGT®: the adaptive immune response http: //www. imgt. org Vertebrates Immunoglobulin (IG) T lymphocyte B lymphocyte peptide T cell receptor (TR) MH Trimolecular complex TR/p. MH
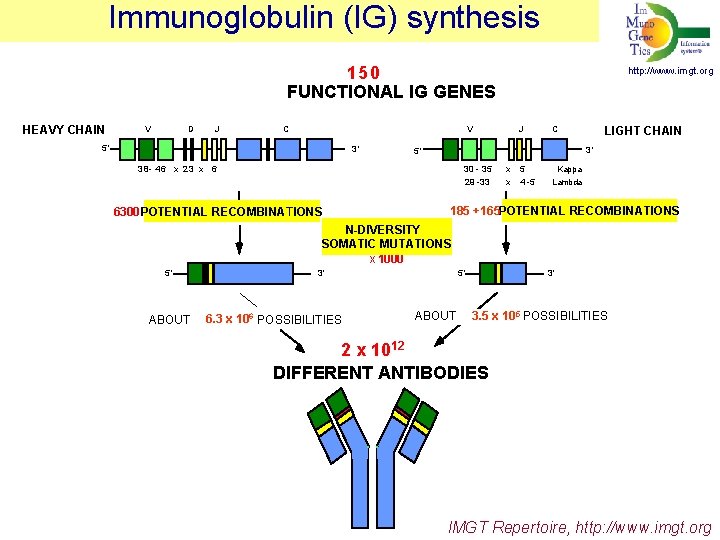
Immunoglobulin (IG) synthesis 150 FUNCTIONAL IG GENES HEAVY CHAIN V D J C V 5' 3' http: //www. imgt. org http: //imgt. cines. fr J C LIGHT CHAIN 3' 5' 38 - 46 x 23 x 6 30 - 35 x 5 Kappa 29 - -33 x 4 -5 Lambda 6300 POTENTIAL RECOMBINATIONS 185 +165 POTENTIAL RECOMBINATIONS N-DIVERSITY SOMATIC MUTATIONS x 1000 5' ABOUT 3' 6 6. 3 x 10 POSSIBILITIES 5' ABOUT 3' 5 POSSIBILITIES 3. 5 x 10 2 x 1012 DIFFERENT ANTIBODIES IMGT Repertoire, http: //www. imgt. org
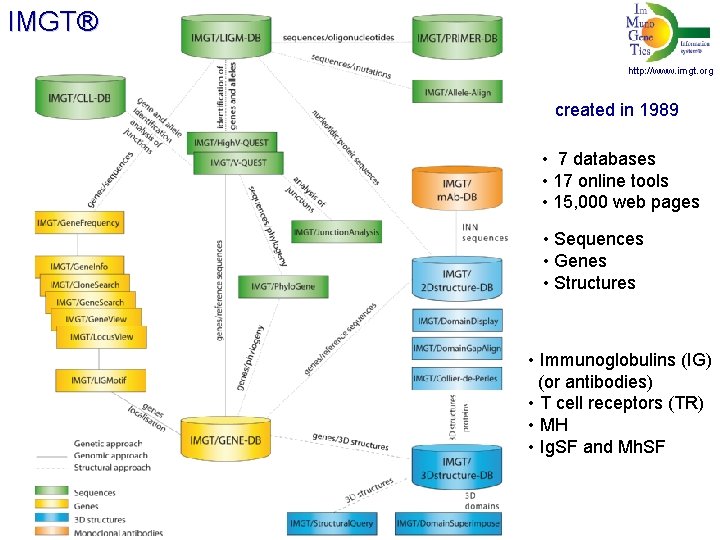
IMGT® http: //www. imgt. org created in 1989 • 7 databases • 17 online tools • 15, 000 web pages • Sequences • Genes • Structures • Immunoglobulins (IG) (or antibodies) • T cell receptors (TR) • MH • Ig. SF and Mh. SF
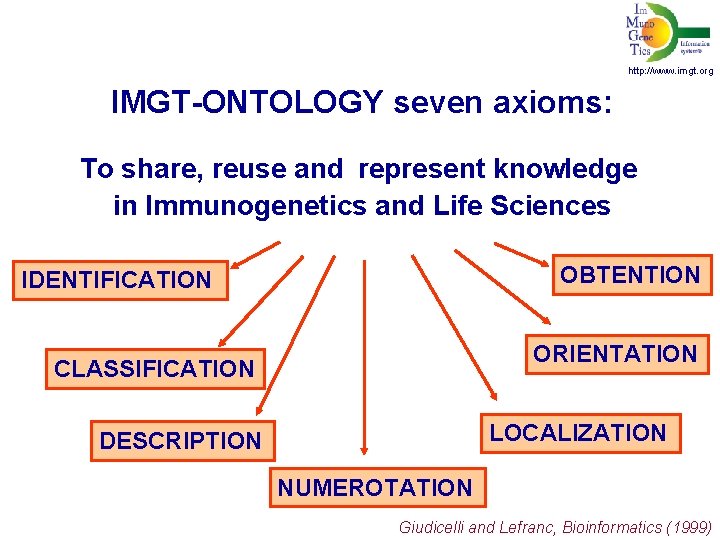
http: //www. imgt. org IMGT-ONTOLOGY seven axioms: To share, reuse and represent knowledge in Immunogenetics and Life Sciences OBTENTION IDENTIFICATION ORIENTATION CLASSIFICATION LOCALIZATION DESCRIPTION NUMEROTATION Giudicelli and Lefranc, Bioinformatics (1999)
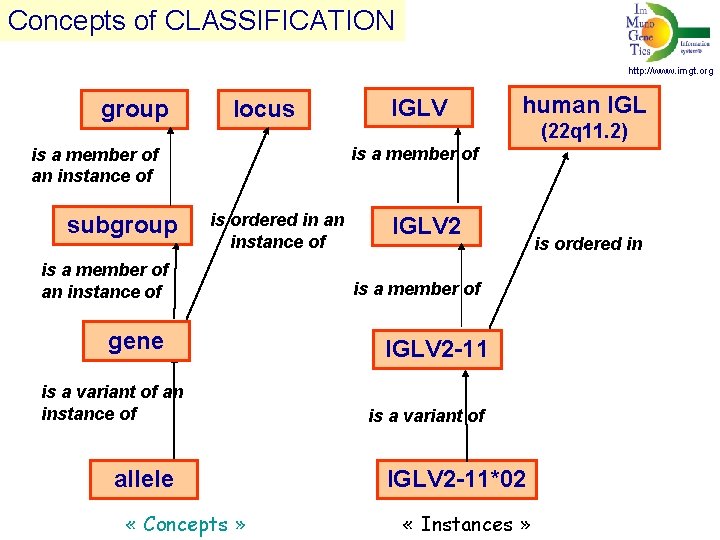
Concepts of CLASSIFICATION http: //www. imgt. org group locus human IGL (22 q 11. 2) is a member of an instance of subgroup IGLV is ordered in an instance of is a member of an instance of gene is a variant of an instance of allele « Concepts » IGLV 2 is a member of IGLV 2 -11 is a variant of IGLV 2 -11*02 « Instances » is ordered in

Concepts of CLASSIFICATION http: //www. imgt. org 1. group, subgroup, gene, allele 2. nomenclature for the V, D, J and C genes 3. immunoglobulin (IG) and T cell receptor (TR) genes 4. IMGT gene names approved by HUGO Nomenclature Committee (HGNC) in 1999. 5. IMGT alleles validated by WHO-IUIS/IMGT-NC IMGT/GENE-DB: international reference database for IG and TR genes (direct links from NCBI Gene) and alleles.

Concepts of DESCRIPTION http: //www. imgt. org IMGT labels (capital letters) IMGT Prototypes (table) IMGT Scientific chart, http: //www. imgt. org

Concepts of DESCRIPTION http: //www. imgt. org 1. standardized IMGT labels 2. used in all IMGT® databases and tools • nucleotide and amino acid sequences (IMGT/LIGM-DB…) • 2 D and 3 D structures (IMGT/3 Dstructure-DB…). 3. Sequence Ontology (SO) includes IMGT labels. 4. IMGT® databases can be queried using labels (a big ‘plus’ compared to generalist databases).
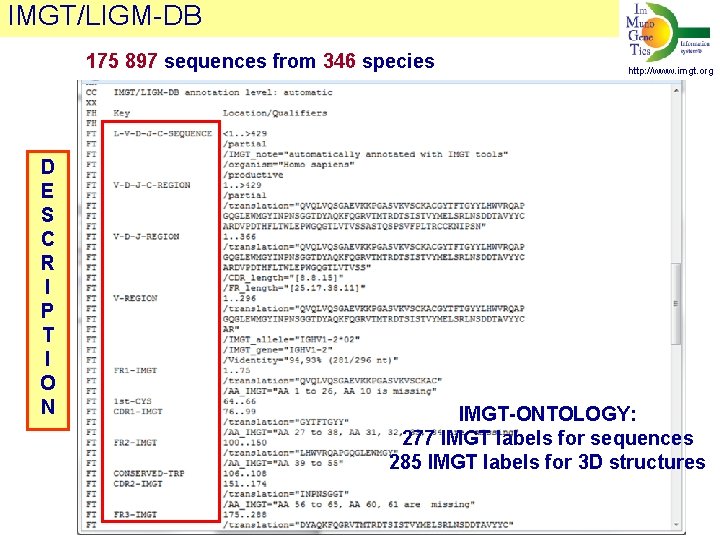
IMGT/LIGM-DB 175 897 sequences from 346 species D E S C R I P T I O N http: //www. imgt. org IMGT-ONTOLOGY: 277 IMGT labels for sequences 285 IMGT labels for 3 D structures
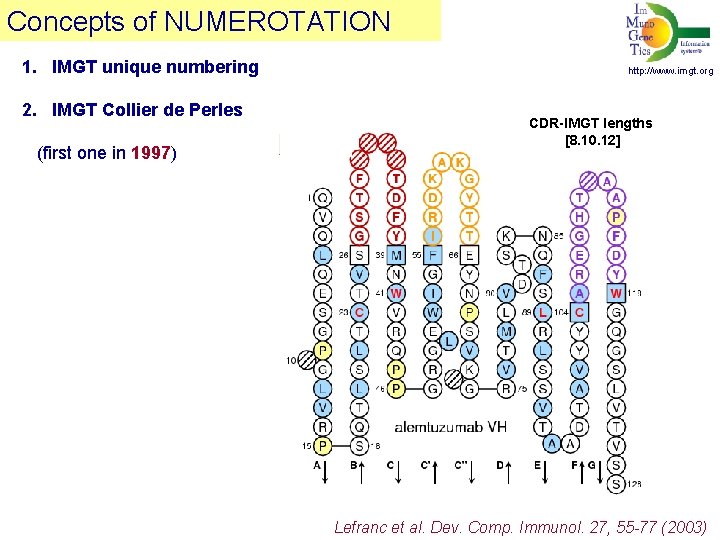
Concepts of NUMEROTATION 1. IMGT unique numbering 2. IMGT Collier de Perles (first one in 1997) http: //www. imgt. org CDR-IMGT lengths [8. 10. 12] Lefranc et al. Dev. Comp. Immunol. 27, 55 -77 (2003)
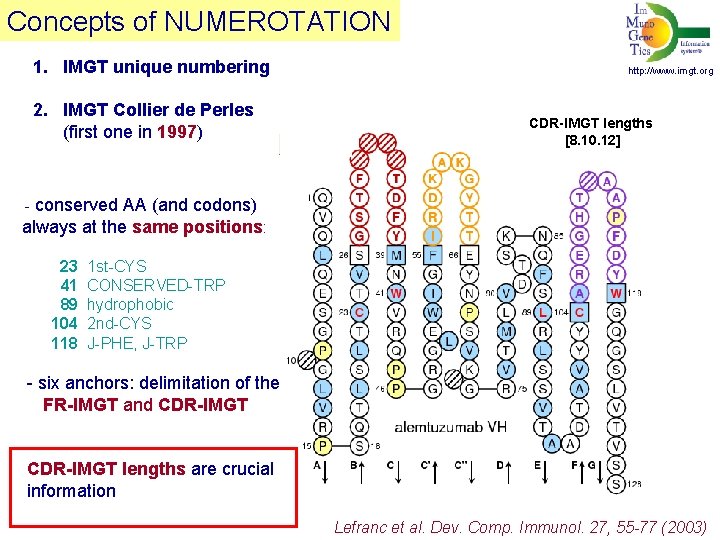
Concepts of NUMEROTATION 1. IMGT unique numbering 2. IMGT Collier de Perles (first one in 1997) http: //www. imgt. org CDR-IMGT lengths [8. 10. 12] - conserved AA (and codons) always at the same positions: 23 1 st-CYS 41 CONSERVED-TRP 89 hydrophobic 104 2 nd-CYS 118 J-PHE, J-TRP - six anchors: delimitation of the FR-IMGT and CDR-IMGT lengths are crucial information Lefranc et al. Dev. Comp. Immunol. 27, 55 -77 (2003)
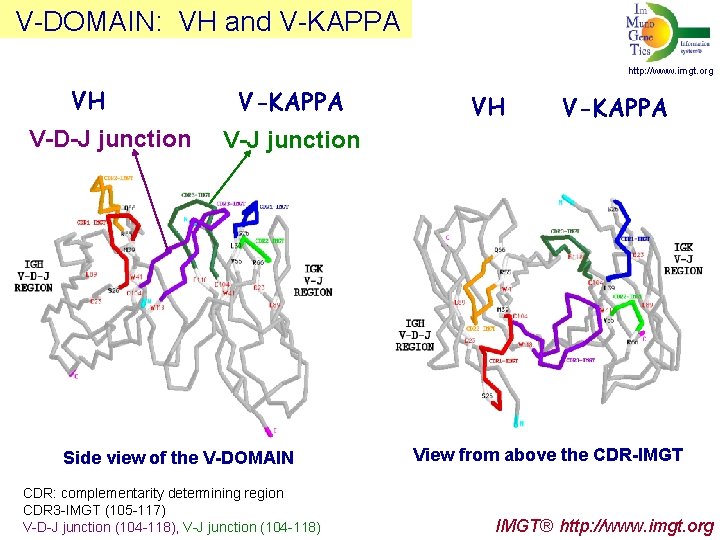
V-DOMAIN: VH and V-KAPPA http: //www. imgt. org VH V-D-J junction V-KAPPA VH V-KAPPA V-J junction Side view of the V-DOMAIN CDR: complementarity determining region CDR 3 -IMGT (105 -117) V-D-J junction (104 -118), V-J junction (104 -118) View from above the CDR-IMGT IMGT® http: //www. imgt. org
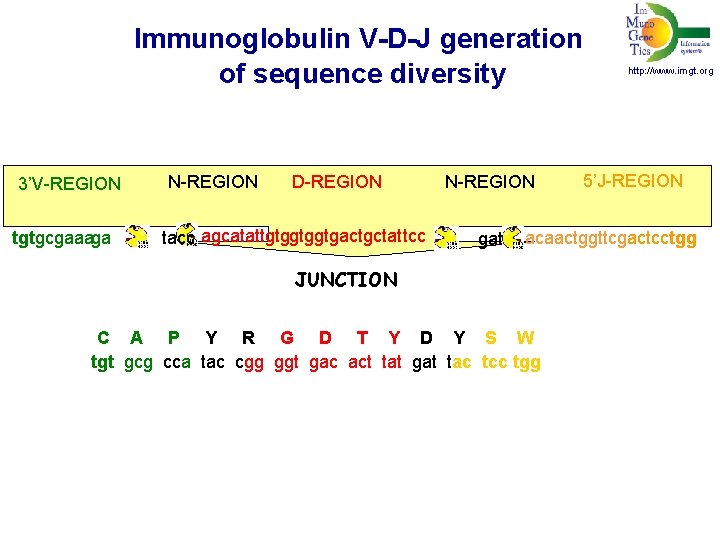
Immunoglobulin V-D-J generation of sequence diversity 3’V-REGION tgtgcgaaaga N-REGION D-REGION tacc agcatattgtggtggtgactgctat tcc N-REGION http: //www. imgt. org 5’J-REGION gatt acaactggttcgactcctgg JUNCTION C A P Y R G D T Y D Y S W tgtgcgcca ggggtgactactat tgt gcg cca tac cgg ggt gac act tat gat tac tcc tgg
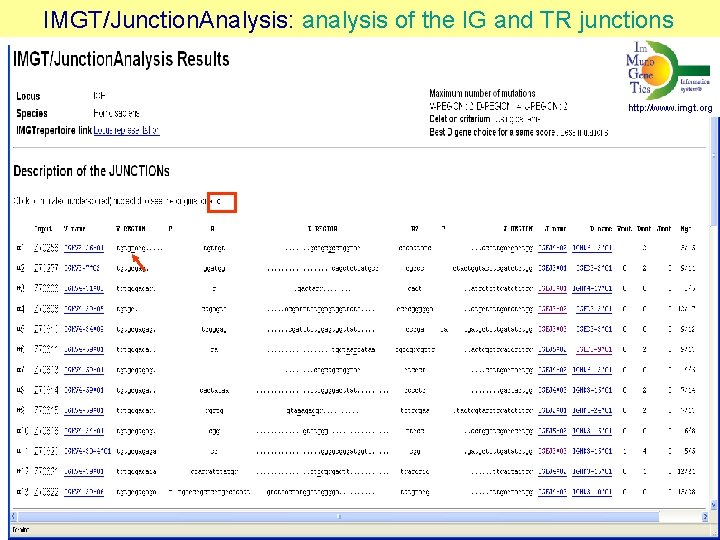
IMGT/Junction. Analysis: analysis of the IG and TR junctions http: //www. imgt. org Yousfi Monod et al. Bioinformatics 20, i 379 -i 385 (200
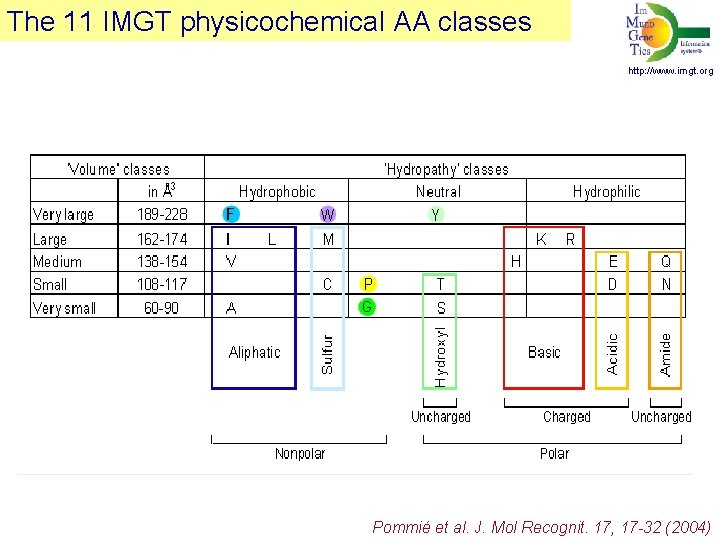
The 11 IMGT physicochemical AA classes http: //www. imgt. org Pommié et al. J. Mol Recognit. 17, 17 -32 (2004)
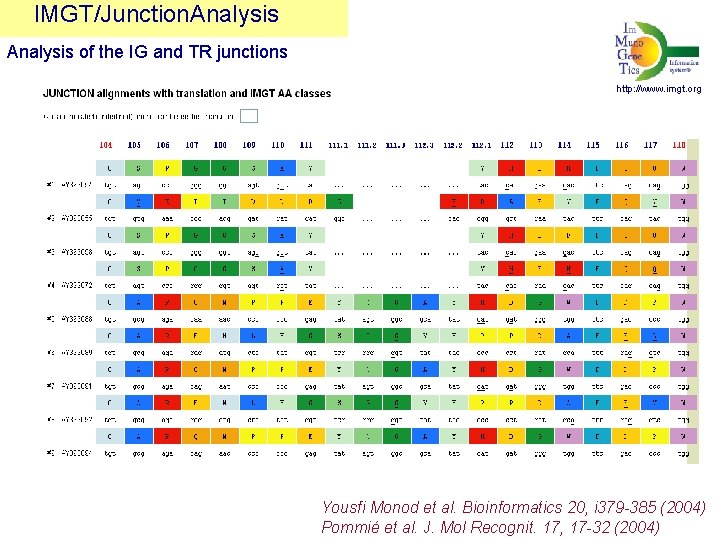
IMGT/Junction. Analysis of the IG and TR junctions http: //www. imgt. org Yousfi Monod et al. Bioinformatics 20, i 379 -385 (2004) Pommié et al. J. Mol Recognit. 17, 17 -32 (2004)
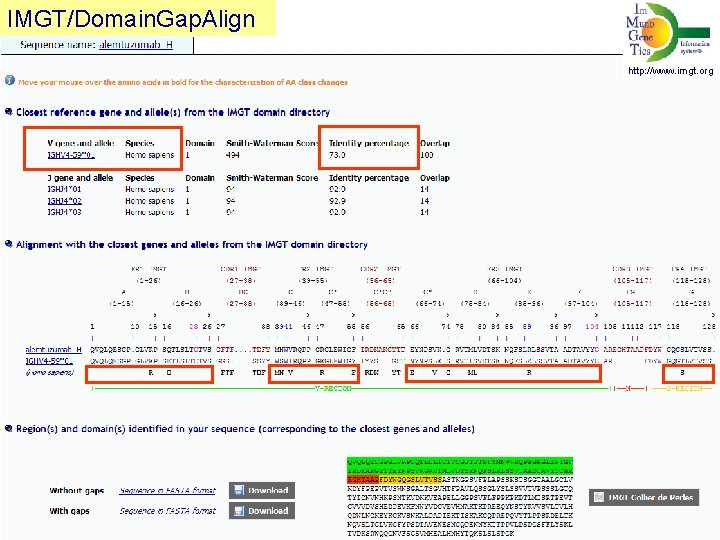
IMGT/Domain. Gap. Align V-REGION identity percent http: //www. imgt. org
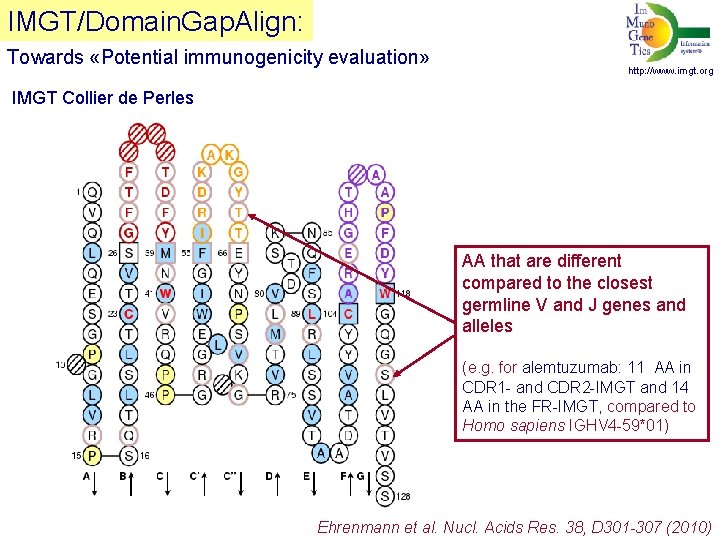
IMGT/Domain. Gap. Align: Towards «Potential immunogenicity evaluation» http: //www. imgt. org IMGT Collier de Perles AA that are different compared to the closest germline V and J genes and alleles (e. g. for alemtuzumab: 11 AA in CDR 1 - and CDR 2 -IMGT and 14 AA in the FR-IMGT, compared to Homo sapiens IGHV 4 -59*01) Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)
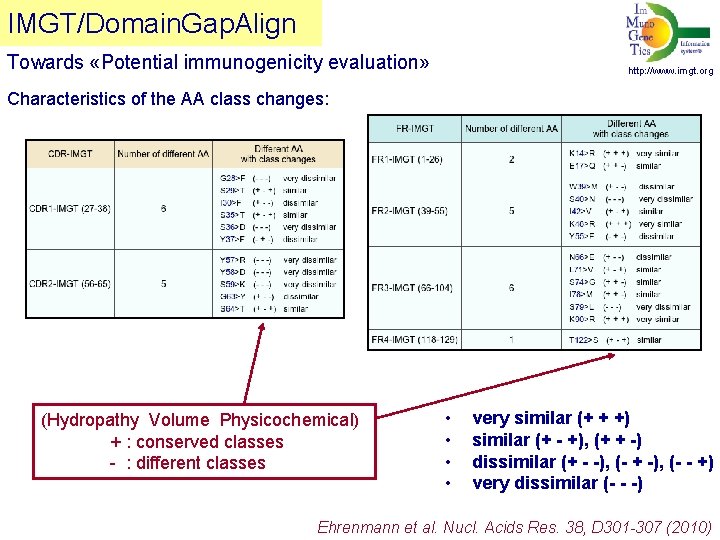
IMGT/Domain. Gap. Align Towards «Potential immunogenicity evaluation» http: //www. imgt. org Characteristics of the AA class changes: (Hydropathy Volume Physicochemical) + : conserved classes - : different classes • • very similar (+ + +) similar (+ - +), (+ + -) dissimilar (+ - -), (- + -), (- - +) very dissimilar (- - -) Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)
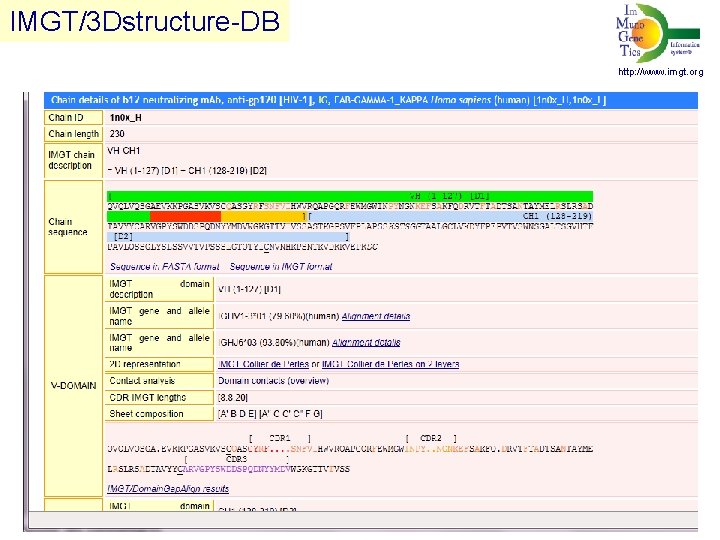
IMGT/3 Dstructure-DB http: //www. imgt. org
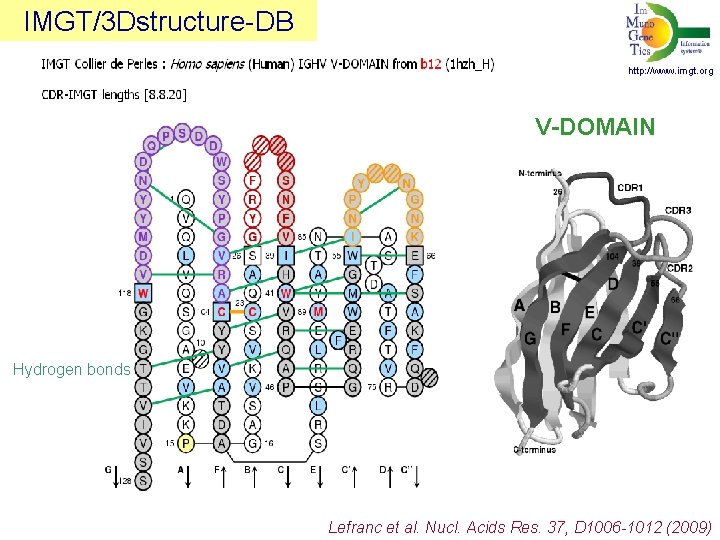
IMGT/3 Dstructure-DB http: //www. imgt. org V-DOMAIN Hydrogen bonds Lefranc et al. Nucl. Acids Res. 37, D 1006 -1012 (2009)
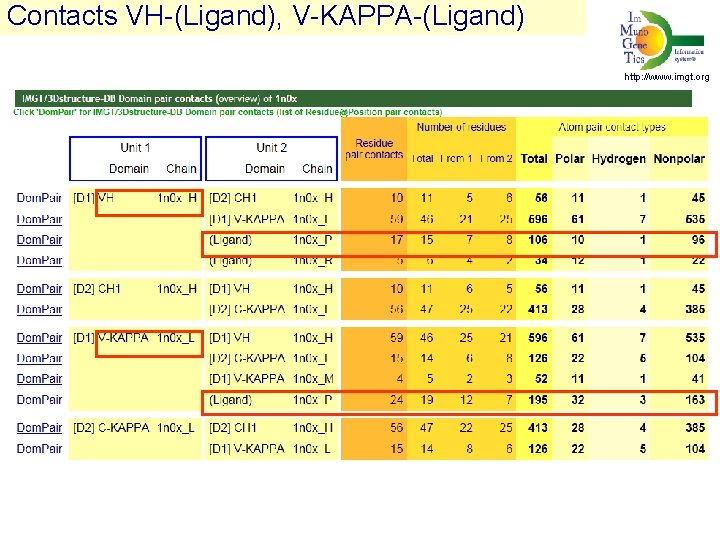
Contacts VH-(Ligand), V-KAPPA-(Ligand) http: //www. imgt. org
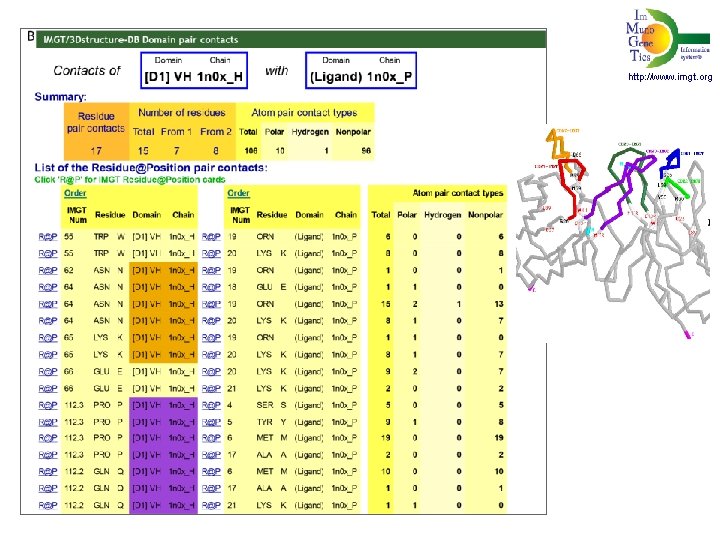
http: //www. imgt. org
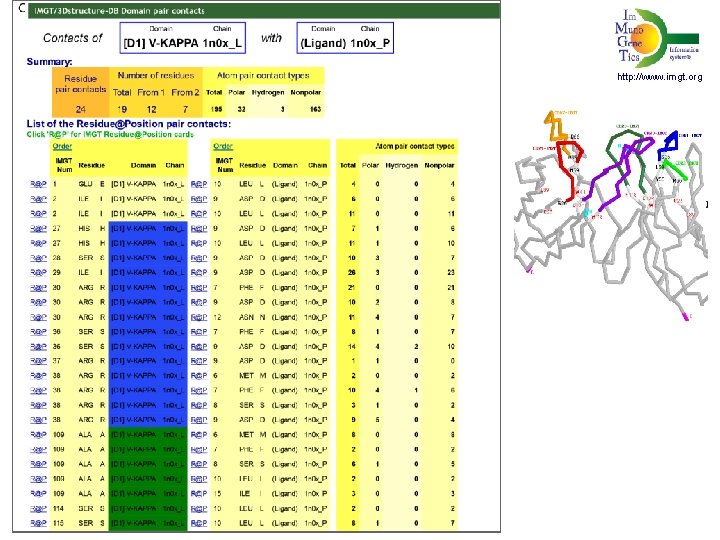
http: //www. imgt. org
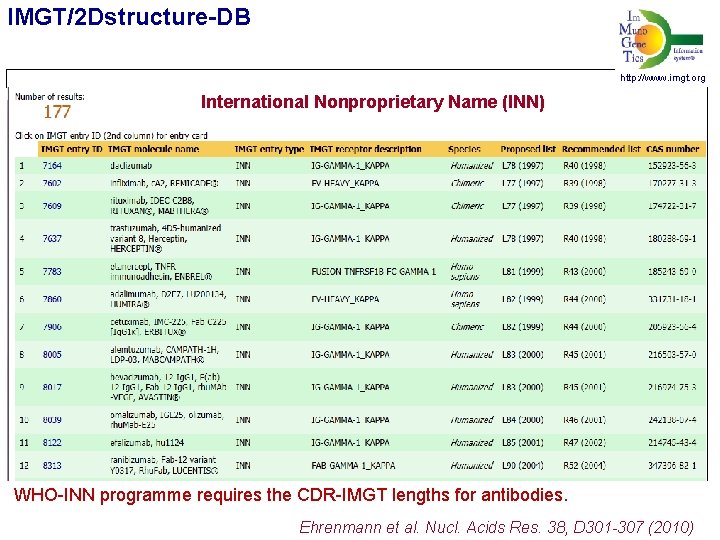
IMGT/2 Dstructure-DB http: //www. imgt. org International Nonproprietary Name (INN) WHO-INN programme requires the CDR-IMGT lengths for antibodies. Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)

IMGT/2 Dstructure-DB http: //www. imgt. org Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)
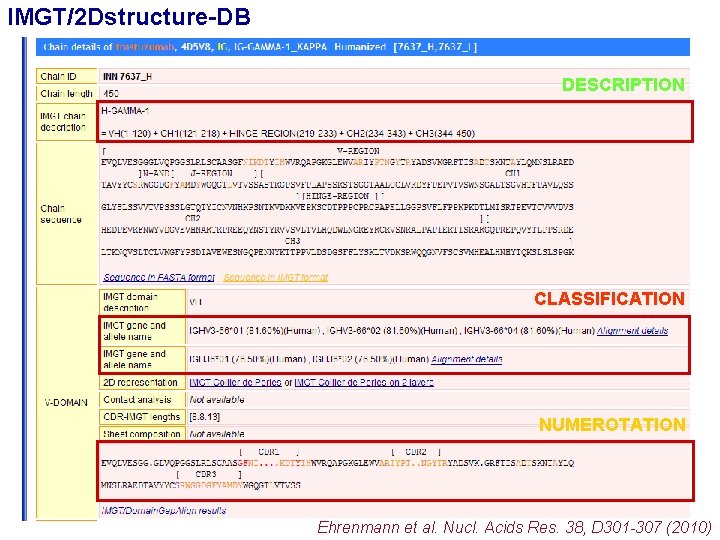
IMGT/2 Dstructure-DB DESCRIPTION CLASSIFICATION NUMEROTATION Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)
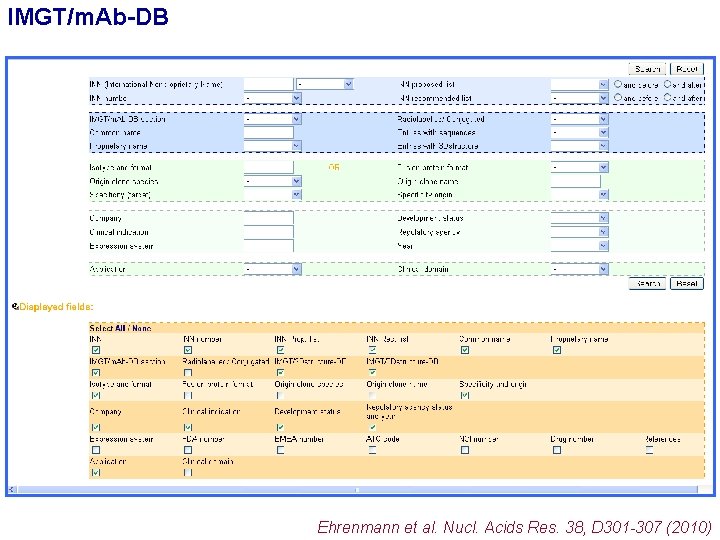
IMGT/m. Ab-DB Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)
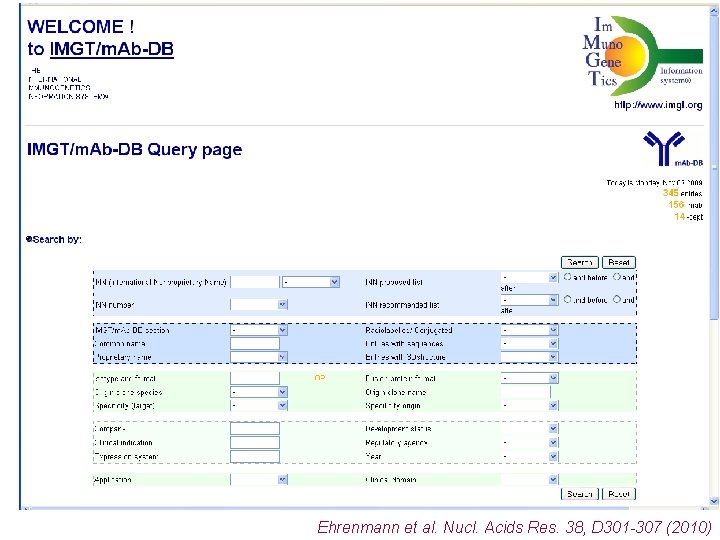
Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)
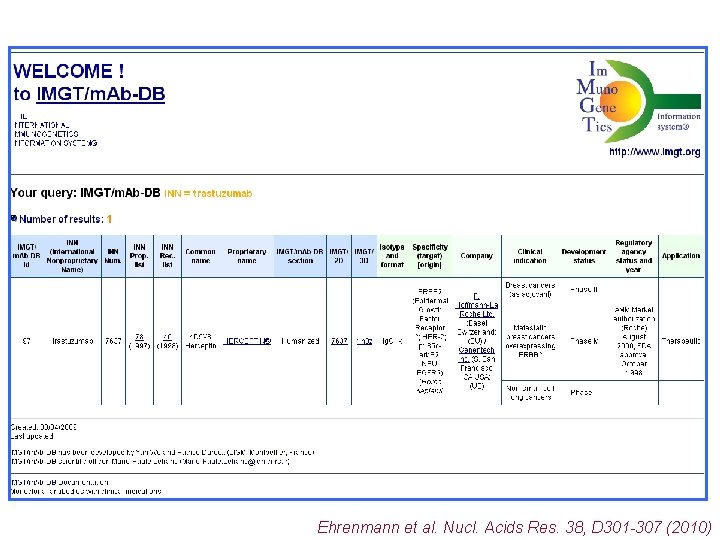
Ehrenmann et al. Nucl. Acids Res. 38, D 301 -307 (2010)

Acknowledgements http: //www. imgt. org Bio. STIC-LR ACI IMPbio GIS AGENAE Plan Pluri-Formation Université Montpellier 2 ANR FLAVORES ANR BIOSYS GIS IBi. SA Grand Plateau Technique Régional Languedoc-Roussillon GPTR «Immuno. Grid» , 6 th PCRDT, STREPS IST and the companies that support the IMGT efforts of standardization.

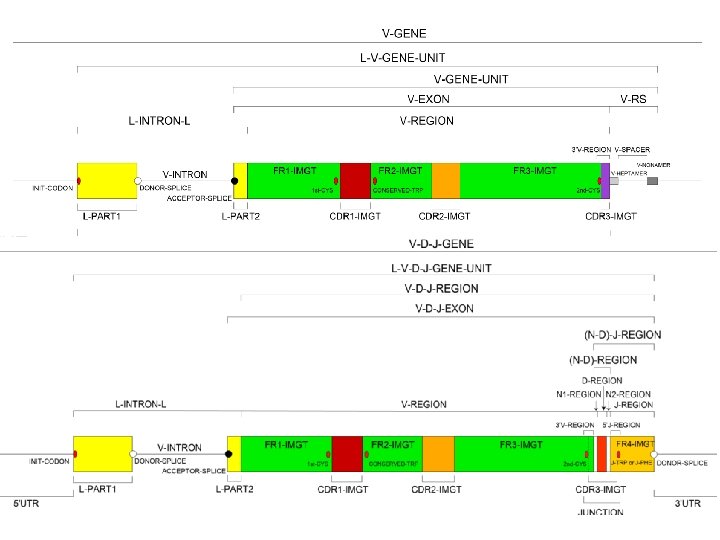
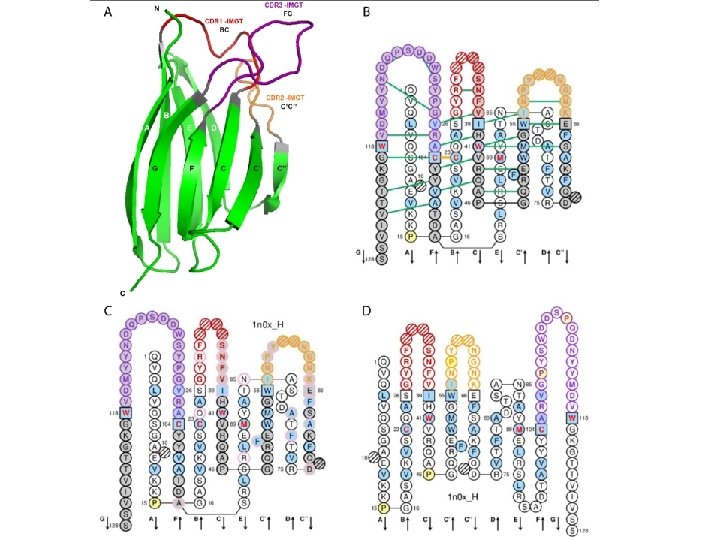
- Slides: 35