http discover nci nih govcellminer Network organization of






- Slides: 14

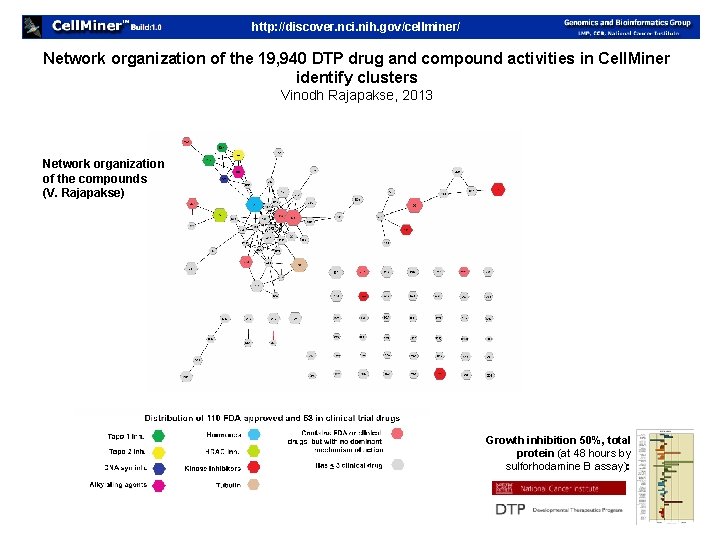
http: //discover. nci. nih. gov/cellminer/ Network organization of the 19, 940 DTP drug and compound activities in Cell. Miner identify clusters Vinodh Rajapakse, 2013 Network organization of the compounds (V. Rajapakse) Growth inhibition 50%, total protein (at 48 hours by sulforhodamine B assay):

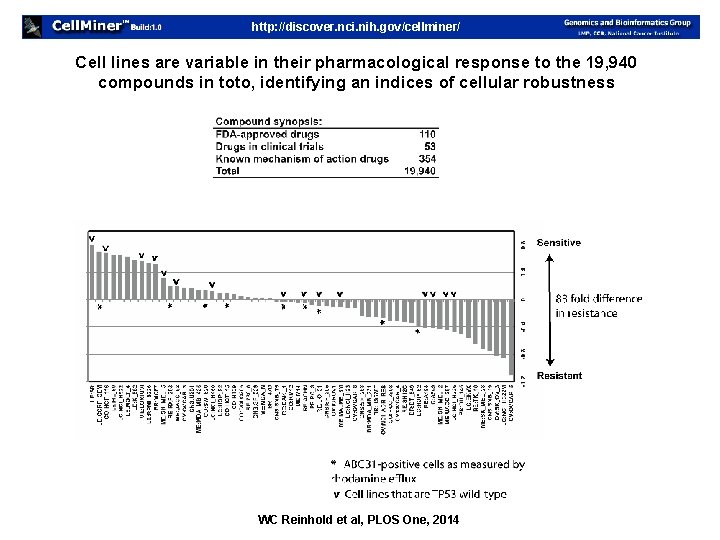
http: //discover. nci. nih. gov/cellminer/ Cell lines are variable in their pharmacological response to the 19, 940 compounds in toto, identifying an indices of cellular robustness WC Reinhold et al, PLOS One, 2014

http: //discover. nci. nih. gov/cellminer/ Applications of the molecular pharmacological data and tools: Four transcript expression examples

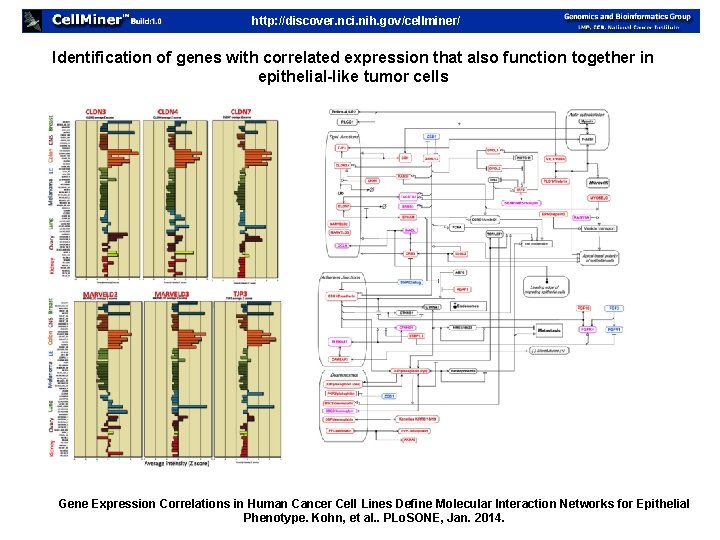
http: //discover. nci. nih. gov/cellminer/ Identification of genes with correlated expression that also function together in epithelial-like tumor cells Gene Expression Correlations in Human Cancer Cell Lines Define Molecular Interaction Networks for Epithelial Phenotype. Kohn, et al. . PLo. SONE, Jan. 2014.

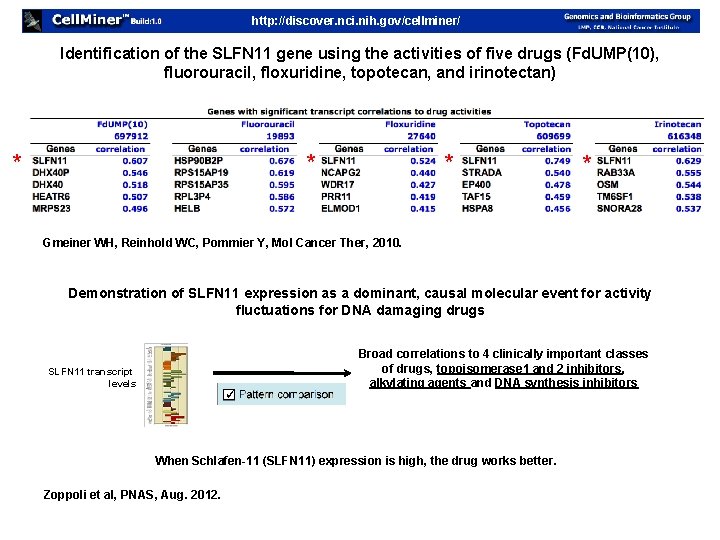
http: //discover. nci. nih. gov/cellminer/ Identification of the SLFN 11 gene using the activities of five drugs (Fd. UMP(10), fluorouracil, floxuridine, topotecan, and irinotectan) * * Gmeiner WH, Reinhold WC, Pommier Y, Mol Cancer Ther, 2010. Demonstration of SLFN 11 expression as a dominant, causal molecular event for activity fluctuations for DNA damaging drugs SLFN 11 transcript levels ✓ Broad correlations to 4 clinically important classes of drugs, topoisomerase 1 and 2 inhibitors, alkylating agents and DNA synthesis inhibitors When Schlafen-11 (SLFN 11) expression is high, the drug works better. Zoppoli et al, PNAS, Aug. 2012.

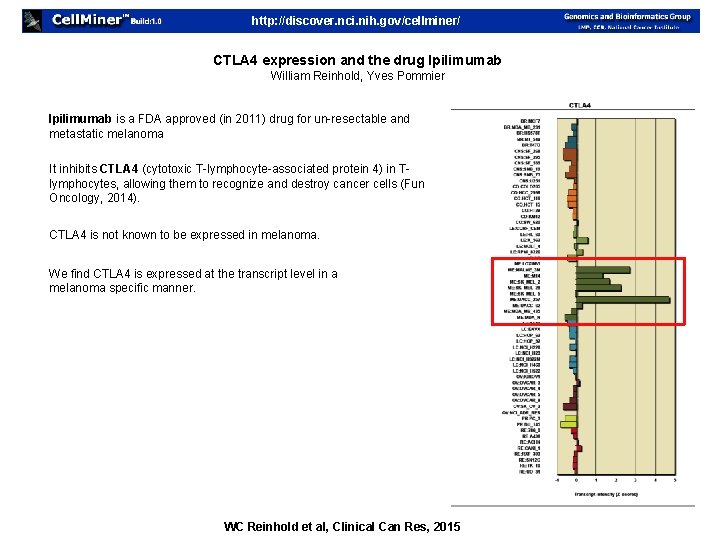
http: //discover. nci. nih. gov/cellminer/ CTLA 4 expression and the drug Ipilimumab William Reinhold, Yves Pommier Ipilimumab is a FDA approved (in 2011) drug for un-resectable and metastatic melanoma It inhibits CTLA 4 (cytotoxic T-lymphocyte-associated protein 4) in Tlymphocytes, allowing them to recognize and destroy cancer cells (Funt, Oncology, 2014). CTLA 4 is not known to be expressed in melanoma. We find CTLA 4 is expressed at the transcript level in a melanoma specific manner. WC Reinhold et al, Clinical Can Res, 2015

http: //discover. nci. nih. gov/cellminer/ RUNX 1 and CBFB expression identification of the HIV TAT antagonist Ro 5 -3335 as a candidate core binding factor (CBF) leukemia treatment CBFB RUNX 1 and CBFB expression Identified Ro 5 -3335 (NSC 743504) as a candidate treatment. Cunningham et al showed causal linkage between the Ro 5 -3335 and the genes. 1. Ro 5 -3335 Interacts with RUNX 1 and CBFβ directly. 2. Ro 5 -3335 repress RUNX 1/CBFB-dependent transactivation in reporter assays. 3. Ro 5 -3335 repress RUNX 1 -dependent hematopoiesis in zebrafish. 4. Ro 5 -3335 preferentially kills human CBF leukemia cell lines. 5. Ro 5 -3335 rescued the AML preleukemic phenotype in RUNX 1–ETO transgenic zebrafish. 6. Ro 5 -3335 reduced leukemia burden in a mouse CBFB–MYH 11 leukemia model. Cunningham et al, PNAS, Aug. 2012

http: //discover. nci. nih. gov/cellminer/ A systems pharmacological application of the data

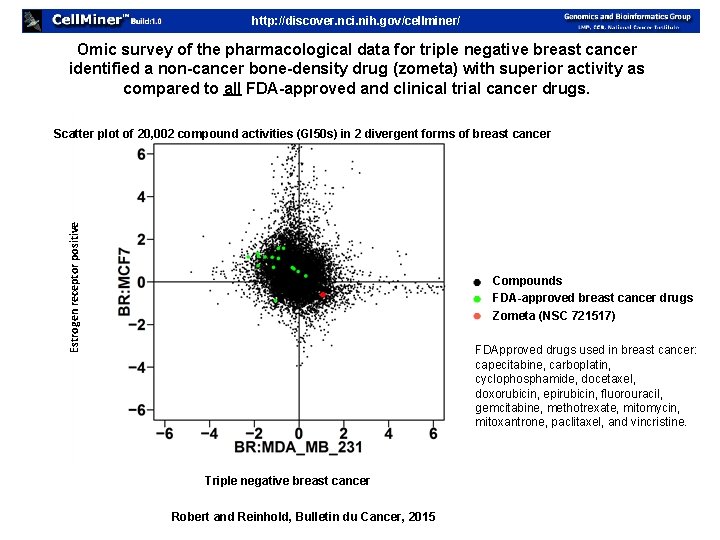
http: //discover. nci. nih. gov/cellminer/ Omic survey of the pharmacological data for triple negative breast cancer identified a non-cancer bone-density drug (zometa) with superior activity as compared to all FDA-approved and clinical trial cancer drugs. Estrogen receptor positive Scatter plot of 20, 002 compound activities (GI 50 s) in 2 divergent forms of breast cancer Compounds FDA-approved breast cancer drugs Zometa (NSC 721517) FDApproved drugs used in breast cancer: capecitabine, carboplatin, cyclophosphamide, docetaxel, doxorubicin, epirubicin, fluorouracil, gemcitabine, methotrexate, mitomycin, mitoxantrone, paclitaxel, and vincristine. Triple negative breast cancer Robert and Reinhold, Bulletin du Cancer, 2015

http: //discover. nci. nih. gov/cellminer/ Molecular Data integration

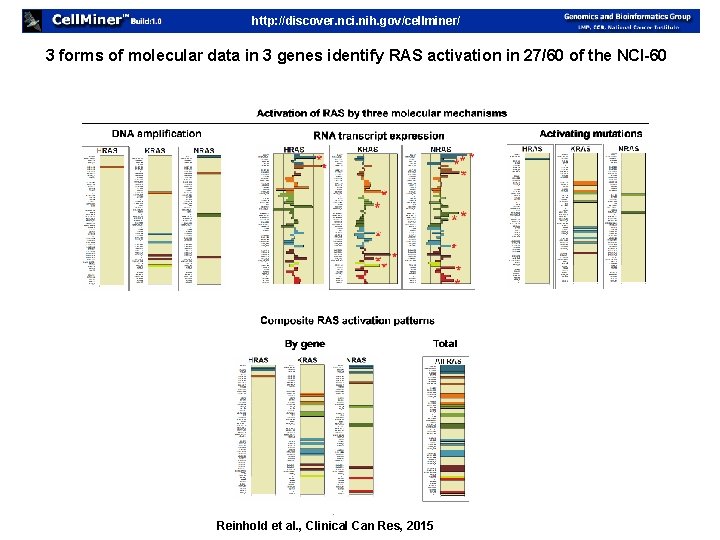
http: //discover. nci. nih. gov/cellminer/ 3 forms of molecular data in 3 genes identify RAS activation in 27/60 of the NCI-60 Reinhold et al. , Clinical Can Res, 2015

http: //discover. nci. nih. gov/cellminer/ 3 forms of molecular data predict PTEN knockdown in 8/60 cell lines, and demonstrate its association to drug activity. Reactivates PTEN An AKT inhibitor Reinhold et al. , Clinical Can Res, 2015

http: //discover. nci. nih. gov/cellminer/ 4 forms of molecular data in 3 genes provide the rationale for the microsatellite instability phenotype found in 8/9 cell lines demonstrated to have that instability Reinhold, et al. , submitted

The GBG is open to collaborations. Bldg. 37, 5041 http: //discover. nci. nih. gov/ wcr@mail. nih. gov William Reinhold Vinodh Rajapakse Margot Sunshine Barry Zeeberg Sudhir Varma Yves Pommier