How to Build a MOD Lincoln Stein Cold

How to Build a MOD Lincoln Stein Cold Spring Harbor Laboratory

What’s a MOD? u Model Organism Database u Repository for reagents – Stocks, vectors, clones – Genetic & physical maps u Large-scale data sets – Genome – EST sets, microarray results, 2 -cell hybrid interactions u Literature u Ontologies & Nomenclature u Meetings, announcements
Worm. Base Web Site
Worm. Base Tour: Looking for MAP Kinase
mek-2 Studies Found RNAi Phenotype a Genetic Locus: & Exprmek-2 Pattern
mek-2 RNAi Phenotype
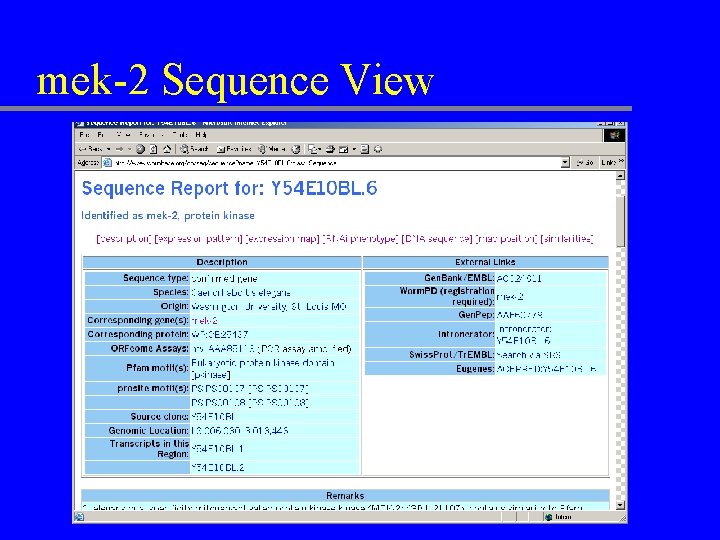
mek-2 Sequence View
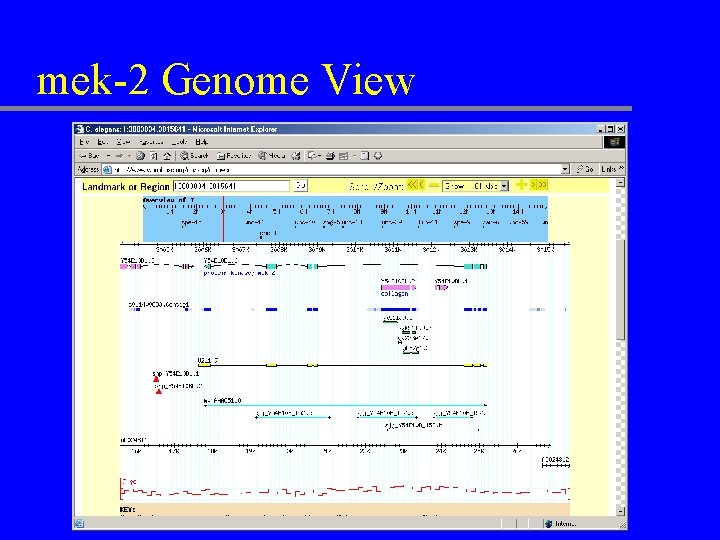
mek-2 Genome View

Elegans/Briggsae Synteny
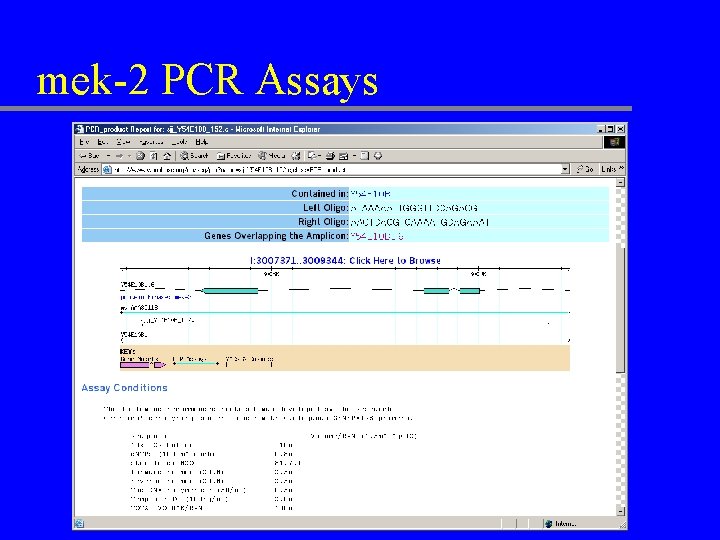
mek-2 PCR Assays
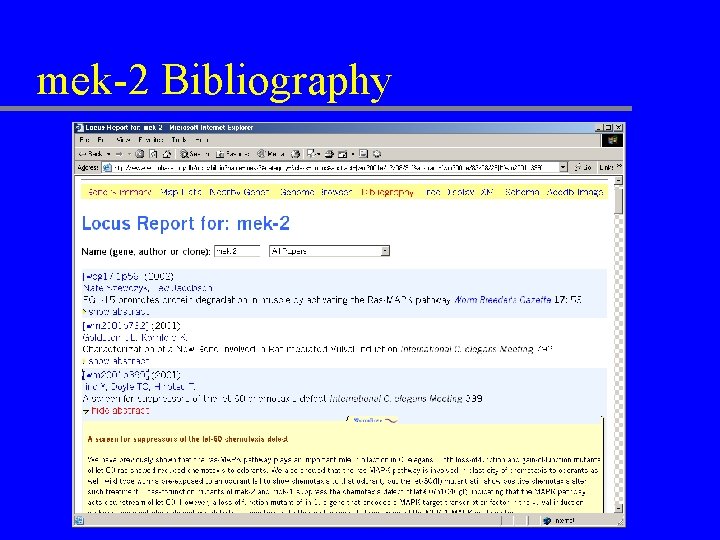
mek-2 Bibliography
mek-2 Citation
VBx Neuroanatomy
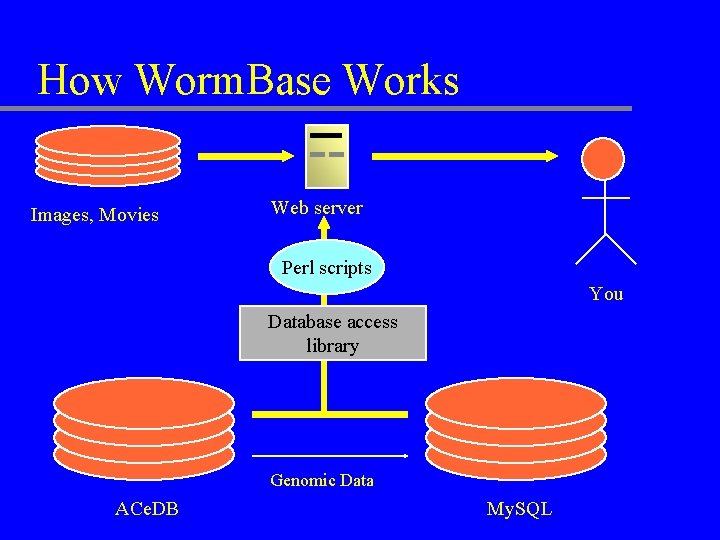
How Worm. Base Works Images, Movies Web server Perl scripts You Database access library Genomic Data ACe. DB My. SQL
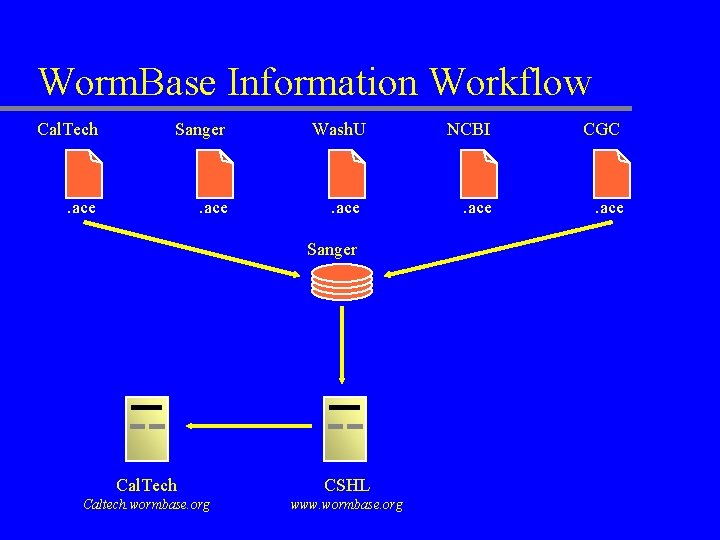
Worm. Base Information Workflow Cal. Tech Sanger . ace Wash. U . ace Sanger Cal. Tech CSHL Caltech. wormbase. org www. wormbase. org NCBI CGC . ace
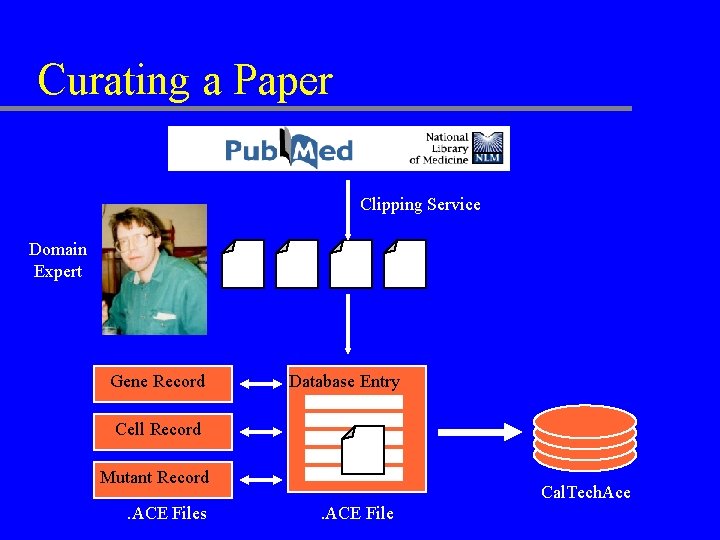
Curating a Paper Clipping Service Domain Expert Gene Record Database Entry Cell Record Mutant Record. ACE Files Cal. Tech. Ace. ACE File

Can You Reuse Worm. Base Software for your Favorite Organism? No!

Sorry Charlie u Wormbase website difficult to install u Data model nematode-centric u Curators tools very process-specific u Customization difficult u Software documentation uneven u Standard operating procedure documentation uneven

MOD Redux u SGD, MGD, Fly. Base, TAIR… u The same basic idea as Worm. Base u Implementation entirely different u Wheel reinvented many times u Little software sharing u This madness must stop!

The GMOD Project u Portable, open source software to support model organism databases u Multiple MODs involved – Worm, fly, yeast, mouse, arabidopsis, rat, monocot, [fugu], [E. coli] u Funded by NIH as of June 2002 – Programmers, coordinator, quarterly meetings http: //www. gmod. org
GMOD Home Page
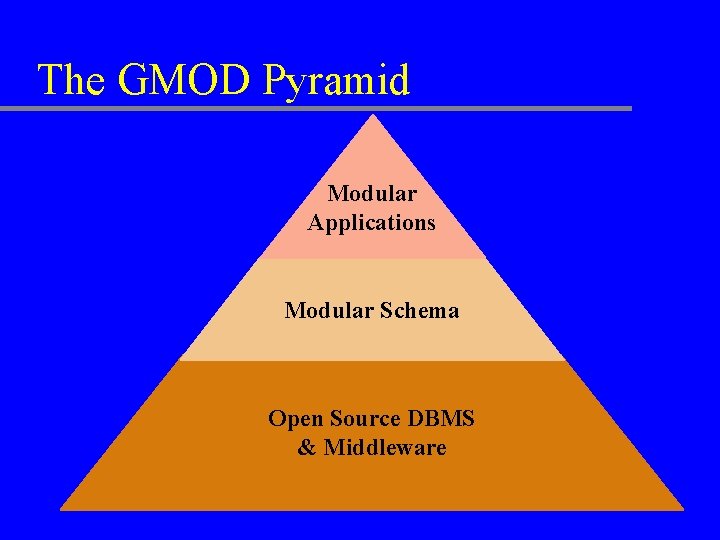
The GMOD Pyramid Modular Applications Modular Schema Open Source DBMS & Middleware
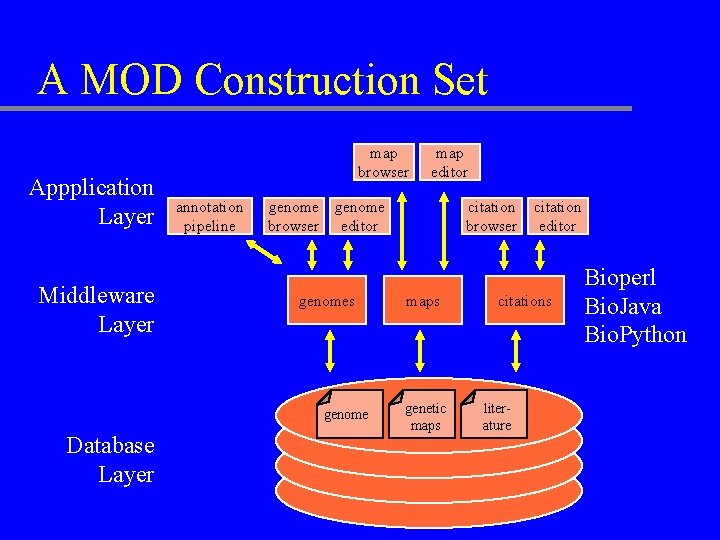
A MOD Construction Set Appplication Layer Middleware Layer map browser annotation pipeline genome browser editor genomes genome Database Layer map editor citation browser maps genetic maps citation editor citations literature Bioperl Bio. Java Bio. Python

Current GMOD Packages u Chado modular schema u Apollo genome annotation editor u Gbrowse generic genome browser u Pub. Search literature curation editor u CMAP comparative map browser u Lab. Doc standard operating procedure editor

Chado – Modular Schema u Immediate goal: common schema for use by Fly. Base and Worm. Base u Ontology Driven – Small number of generic tables e. g. “feature” – Controlled vocabulary names subtypes & describes relationships among them » e. g. “transcript fg 83. 2 encodes protein fp 1803” – Detail tables provide further information on subtypes
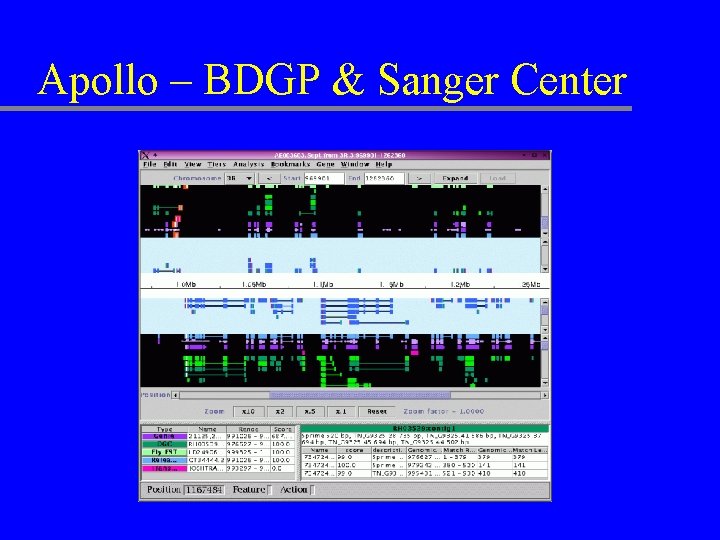
Apollo – BDGP & Sanger Center

Apollo Data adapters u Parser -> data models -> display u Existing data adapters – – GAME XML GFF Ensembl CGI server DAS u Write your own data adapter! – Extend Abstract. Data. Adapter class – Display options defined in config file

Who is Using Apollo? u BDGP – Reannotated Drosophila genome u Bristol-Myers Squibb – Launching Apollo from web browser via mime types u GNF – JDBC adapter layer over Bio. SQL u Biogen – View human genome alignment between public and Biogen internal database – Connected BLAT pipeline to Apollo u HGMP-RC Fugu Genomics group – Displaying annotations on fugu scaffolds

Pub. Search – TAIR & Rat. DB
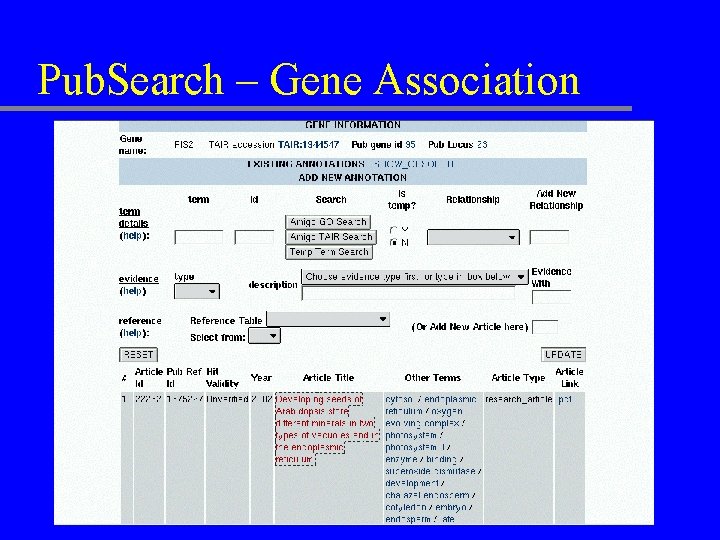
Pub. Search – Gene Association
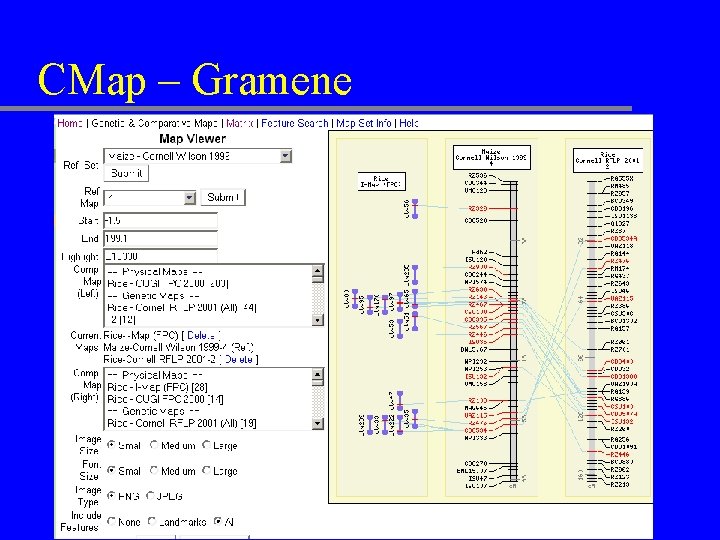
CMap – Gramene
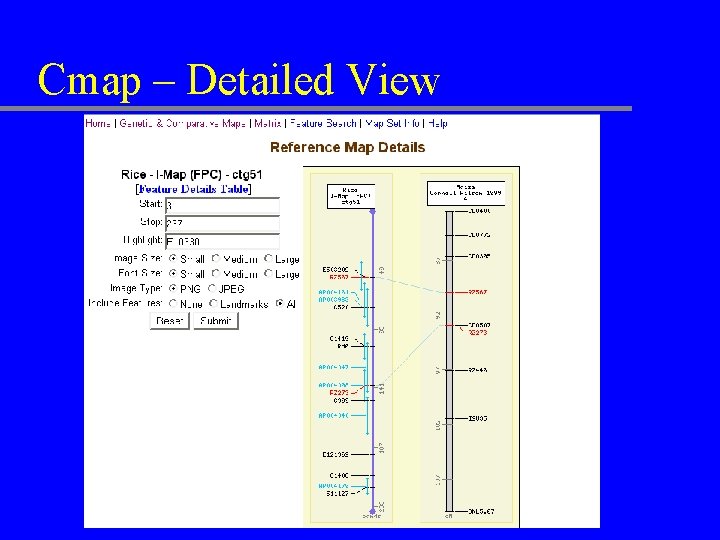
Cmap – Detailed View

GBrowse – Worm. Base

GBrowse – Zoomed in

GBrowse – Zoomed Way In

GBrowse – Zoomed Way In

GBrowse – Keyword Search
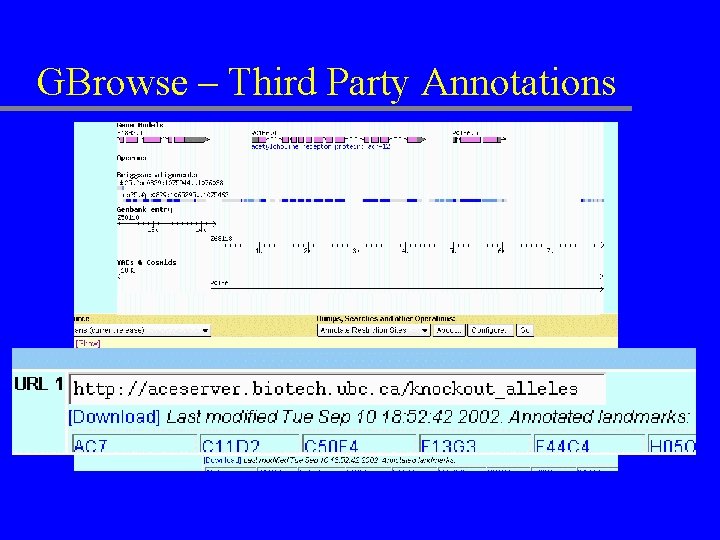
GBrowse – Third Party Annotations

Sequence dumps & other reports

Extensively Customizable u End-user – Turn tracks on and off, change order, change packing & labeling attributes (stored in cookie) u Data provider – Change fonts, colors, text. – Change overview – genetic map, contigs, coverage, karyotype. – Define new tracks using simple config file. – Tinker with track appearance to hearts content.

Adding a New Track (a) Create a GFF file named “deletions. gff” Chr 1 targeted deletion 1293224 1294901. . . Deletion d 101 k 2 Chr 1 targeted deletion 8239811 8241116. . . Deletion d 680 k 2 Chr 2 targeted deletion 5866382 5866500. . . Deletion d 007 k 2 (b) Run the load_gff. pl script > load_gff. pl –d example_database deletions. gff Loading features… Done. 3 features loaded. (c) Add a new track “stanza” to the gbrowse configuration file [Knockout] feature = deletion glyph = span fgcolor = red key = Knockouts link = http: //example. org/cgi-bin/knockout_details? $name citation = These are deletion knockouts produced by the example knockout consortium (http: //example. org/knockouts. html)
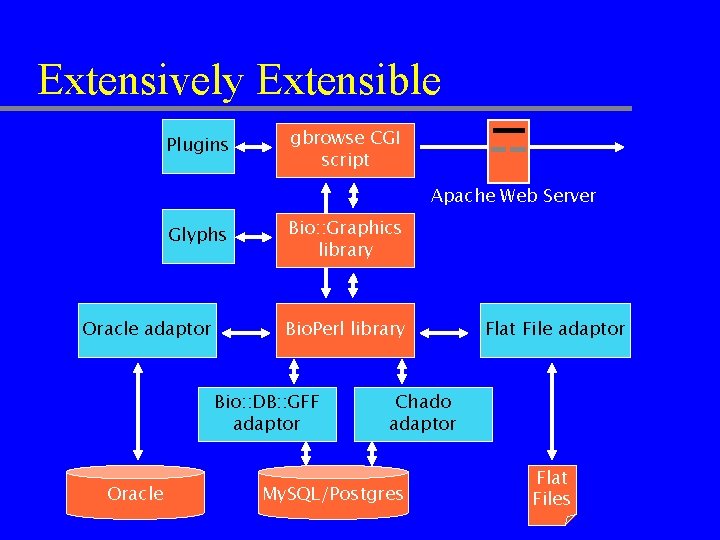
Extensively Extensible Plugins gbrowse CGI script Apache Web Server Glyphs Oracle adaptor Bio: : Graphics library Bio. Perl library Bio: : DB: : GFF adaptor Oracle Flat File adaptor Chado adaptor My. SQL/Postgres Flat Files
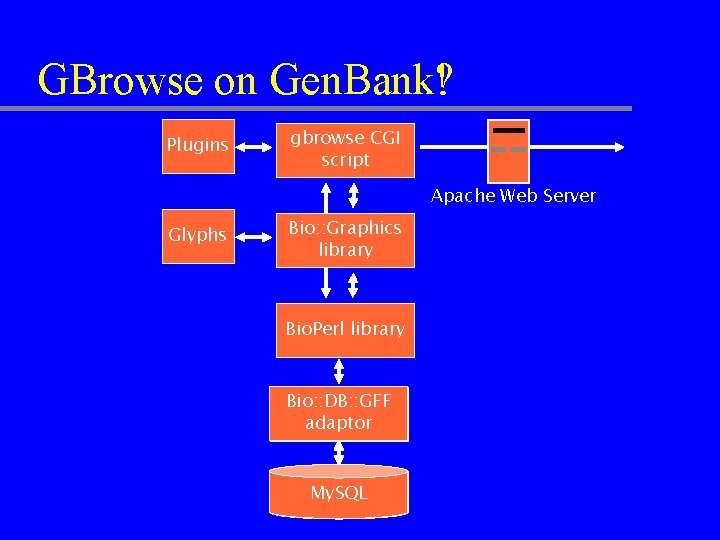
GBrowse on Gen. Bank! Gen. Bank? Plugins gbrowse CGI script Apache Web Server Glyphs Bio: : Graphics library Bio. Perl library Bio: : DB: : GFF Gen. Bank adaptor Proxy Adaptor My. SQL Gen. Bank
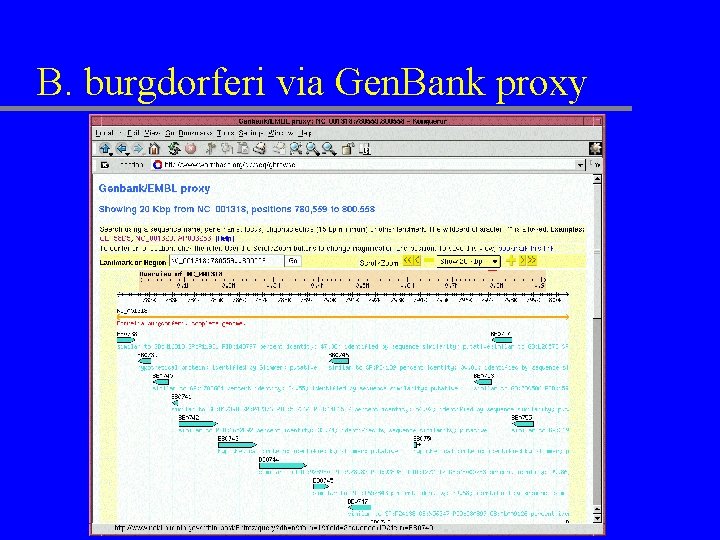
B. burgdorferi via Gen. Bank proxy

Who is Using GBrowse? u GMOD Members – Worm. Base, Fly. Base, Rat. DB u HGMP-RC Fugu genomics group u KEGG (multiple microorganisms) u Ingenium AG (mouse) u Bristoll-Myers Squibb (drosophila) u Texas A&M University (salmonella) u Institute of Systems Biology (human)

Coming Soon to www. gmod. org u Biopipe – genome annotation pipeline u Insertional mutagenesis analysis pipeline u Tree browser u Pathway browsers u Generic MOD web site framework

Joining GMOD u Go to www. gmod. org u Examine software matrix u Find a project or suggest new one u Contact Scott Cain: cain@cshl. org – Or mail gmod-dev@lists. sourceforge. net

Credits CSHL Adrian Arva Shuly Avraham Scott Cain Ken Clark Allen Day Harvard David Emmert Stan Letovsky BDGP Nomi Harris Suzanna Lewis Chris Mungall John Richter Sheng. Qiang Shu Colin Weil EBI Michele Clamp Stephen Searle Carnegie Institute Sue Rhee Danny Yoo http: //www. gmod. org
- Slides: 48