How species richness and total abundance constrain the
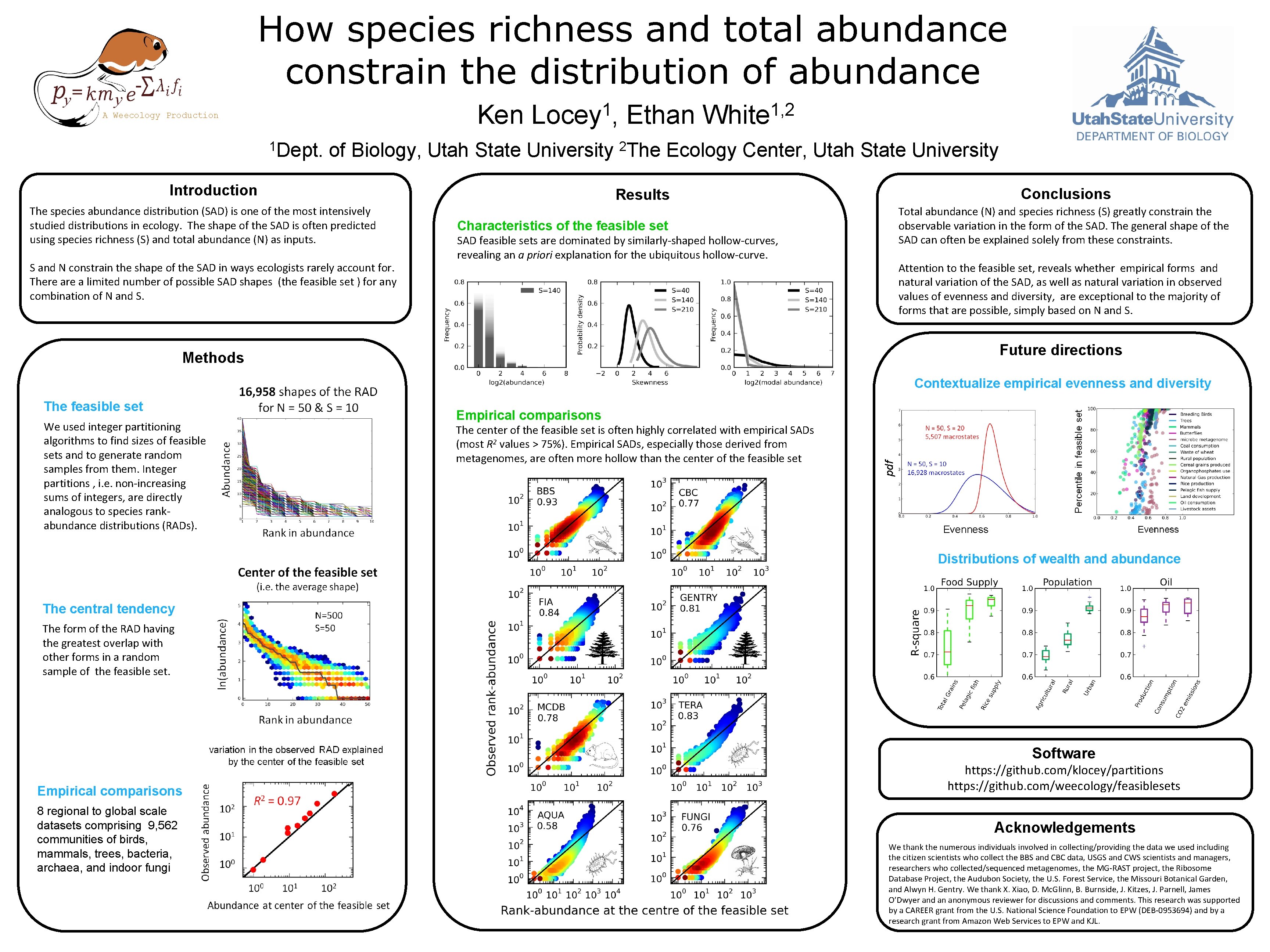
How species richness and total abundance constrain the distribution of abundance Ken A Weecology Production 1 Dept. 1 Locey , of Biology, Utah State University Introduction The species abundance distribution (SAD) is one of the most intensively studied distributions in ecology. The shape of the SAD is often predicted using species richness (S) and total abundance (N) as inputs. S and N constrain the shape of the SAD in ways ecologists rarely account for. There a limited number of possible SAD shapes (the feasible set ) for any combination of N and S. Ethan 2 The 1, 2 White Ecology Center, Utah State University Conclusions Results Total abundance (N) and species richness (S) greatly constrain the observable variation in the form of the SAD. The general shape of the SAD can often be explained solely from these constraints. Characteristics of the feasible set SAD feasible sets are dominated by similarly-shaped hollow-curves, revealing an a priori explanation for the ubiquitous hollow-curve. Attention to the feasible set, reveals whether empirical forms and natural variation of the SAD, as well as natural variation in observed values of evenness and diversity, are exceptional to the majority of forms that are possible, simply based on N and S. Future directions Methods Contextualize empirical evenness and diversity We used integer partitioning algorithms to find sizes of feasible sets and to generate random samples from them. Integer partitions , i. e. non-increasing sums of integers, are directly analogous to species rankabundance distributions (RADs). Empirical comparisons The center of the feasible set is often highly correlated with empirical SADs (most R 2 values > 75%). Empirical SADs, especially those derived from metagenomes, are often more hollow than the center of the feasible set pdf The feasible set Evenness Distributions of wealth and abundance The central tendency The form of the RAD having the greatest overlap with other forms in a random sample of the feasible set. Software Empirical comparisons 8 regional to global scale datasets comprising 9, 562 communities of birds, mammals, trees, bacteria, archaea, and indoor fungi https: //github. com/klocey/partitions https: //github. com/weecology/feasiblesets Acknowledgements We thank the numerous individuals involved in collecting/providing the data we used including the citizen scientists who collect the BBS and CBC data, USGS and CWS scientists and managers, researchers who collected/sequenced metagenomes, the MG-RAST project, the Ribosome Database Project, the Audubon Society, the U. S. Forest Service, the Missouri Botanical Garden, and Alwyn H. Gentry. We thank X. Xiao, D. Mc. Glinn, B. Burnside, J. Kitzes, J. Parnell, James O’Dwyer and an anonymous reviewer for discussions and comments. This research was supported by a CAREER grant from the U. S. National Science Foundation to EPW (DEB-0953694) and by a research grant from Amazon Web Services to EPW and KJL.
- Slides: 1