How does insulin work 1984 Pierre De Meyts











- Slides: 45

How does insulin work? (~1984) Pierre De Meyts = Chuck; head of research at Novo Nordsk 1 Aug

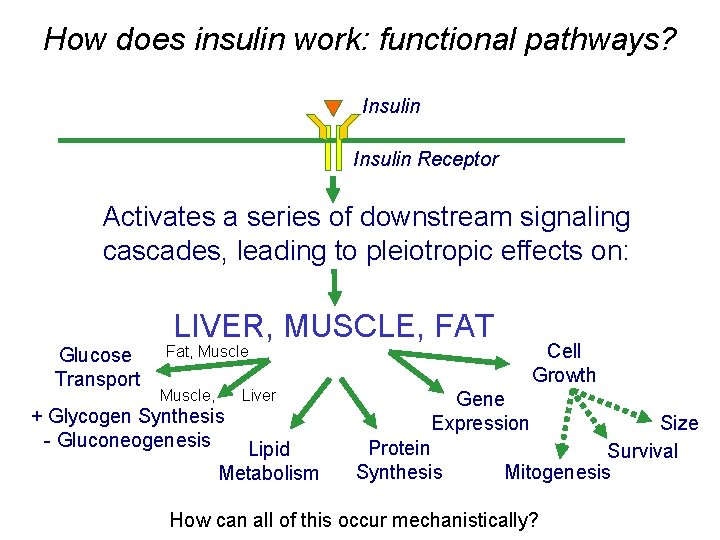
How does insulin work: functional pathways? Insulin Receptor Activates a series of downstream signaling cascades, leading to pleiotropic effects on: LIVER, MUSCLE, FAT Glucose Transport Cell Growth Fat, Muscle, Liver + Glycogen Synthesis - Gluconeogenesis Lipid Metabolism Gene Expression Protein Synthesis Size Survival Mitogenesis How can all of this occur mechanistically?

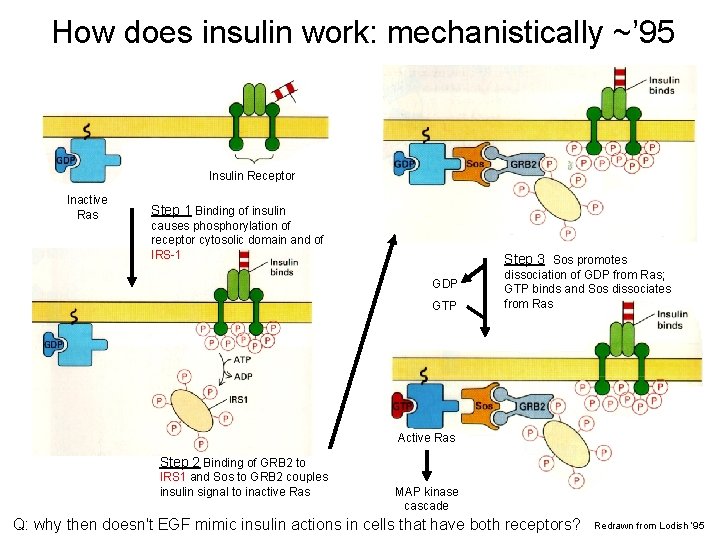
How does insulin work: mechanistically ~’ 95 Insulin Receptor Inactive Ras Step 1 Binding of insulin causes phosphorylation of receptor cytosolic domain and of IRS-1 Step 3 Sos promotes GDP GTP dissociation of GDP from Ras; GTP binds and Sos dissociates from Ras Active Ras Step 2 Binding of GRB 2 to IRS 1 and Sos to GRB 2 couples insulin signal to inactive Ras MAP kinase cascade Q: why then doesn't EGF mimic insulin actions in cells that have both receptors? Redrawn from Lodish ‘ 95

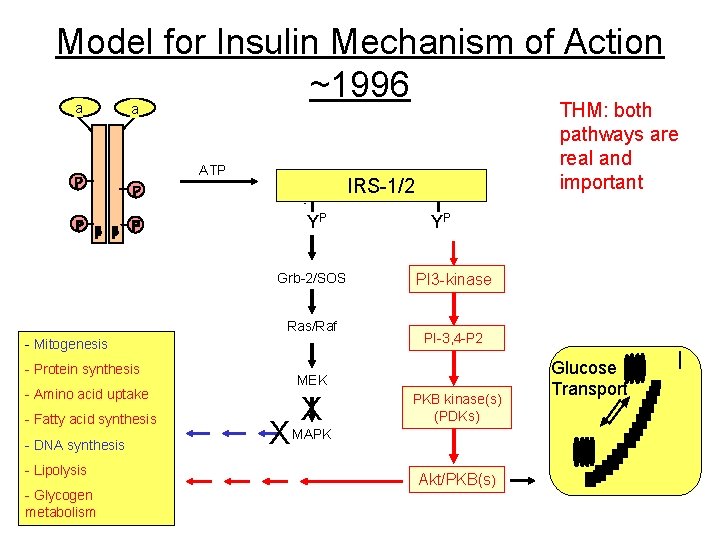
Model for Insulin Mechanism of Action ~1996 a a ATP IRS-1/2 YP Grb-2/SOS Ras/Raf - Mitogenesis - Protein synthesis - Amino acid uptake - Fatty acid synthesis - DNA synthesis - Lipolysis - Glycogen metabolism THM: both pathways are real and important YP PI 3 -kinase PI-3, 4 -P 2 MEK X X MAPK PKB kinase(s) (PDKs) Akt/PKB(s) Glucose Transport

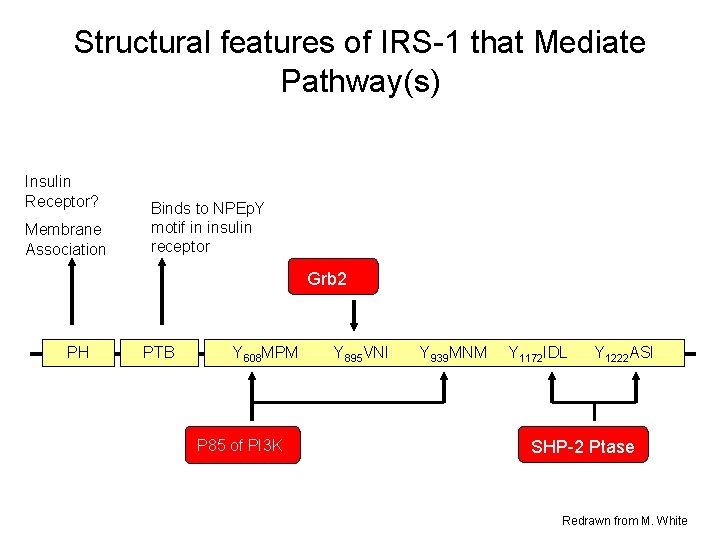
Structural features of IRS-1 that Mediate Pathway(s) Insulin Receptor? Membrane Association Binds to NPEp. Y motif in insulin receptor Grb 2 PH PTB Y 608 MPM P 85 of PI 3 K Y 895 VNI Y 939 MNM Y 1172 IDL Y 1222 ASI SHP-2 Ptase Redrawn from M. White

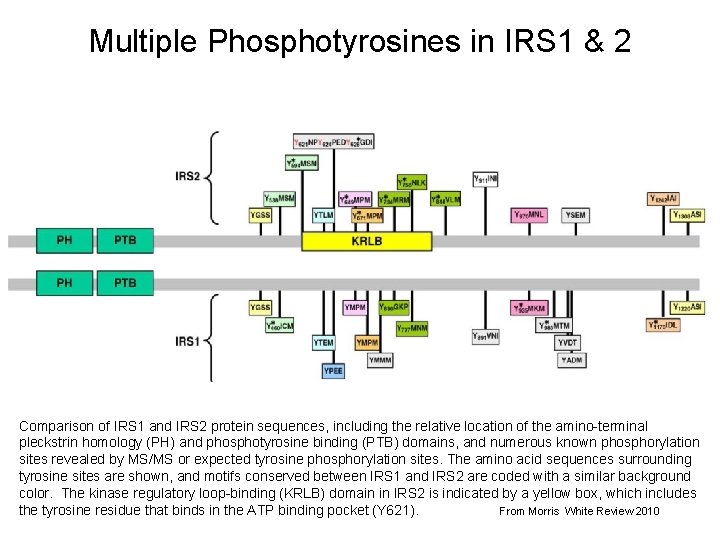
Multiple Phosphotyrosines in IRS 1 & 2 Comparison of IRS 1 and IRS 2 protein sequences, including the relative location of the amino-terminal pleckstrin homology (PH) and phosphotyrosine binding (PTB) domains, and numerous known phosphorylation sites revealed by MS/MS or expected tyrosine phosphorylation sites. The amino acid sequences surrounding tyrosine sites are shown, and motifs conserved between IRS 1 and IRS 2 are coded with a similar background color. The kinase regulatory loop-binding (KRLB) domain in IRS 2 is indicated by a yellow box, which includes 6 the tyrosine residue that binds in the ATP binding pocket (Y 621). From Morris White Review 2010

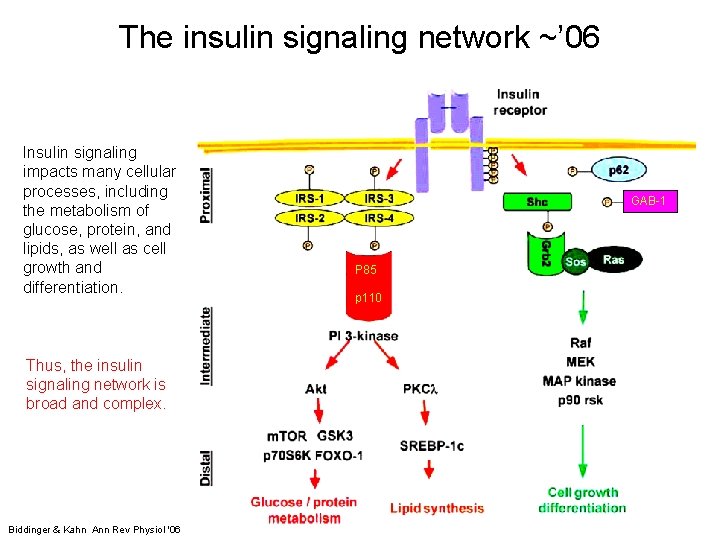
The insulin signaling network ~’ 06 Insulin signaling impacts many cellular processes, including the metabolism of glucose, protein, and lipids, as well as cell growth and differentiation. GAB-1 P 85 p 110 Thus, the insulin signaling network is broad and complex. 7 Biddinger & Kahn Ann Rev Physiol '06

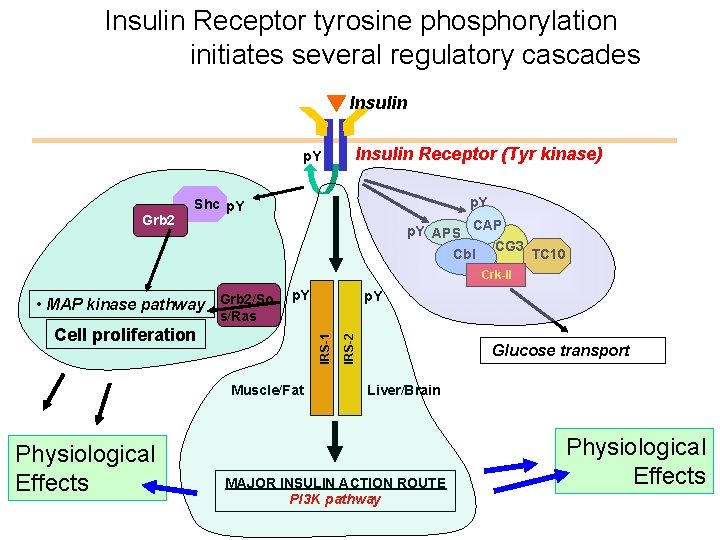
Insulin Receptor tyrosine phosphorylation initiates several regulatory cascades Insulin p. Y Grb 2 Insulin Receptor (Tyr kinase) p. Y Shc p. Y APS CAP CG 3 TC 10 Cbl Crk-II • MAP kinase pathway Grb 2/So p. Y IRS-1 Cell proliferation Muscle/Fat Physiological Effects IRS-2 s/Ras Glucose transport Liver/Brain MAJOR INSULIN ACTION ROUTE PI 3 K pathway Physiological Effects

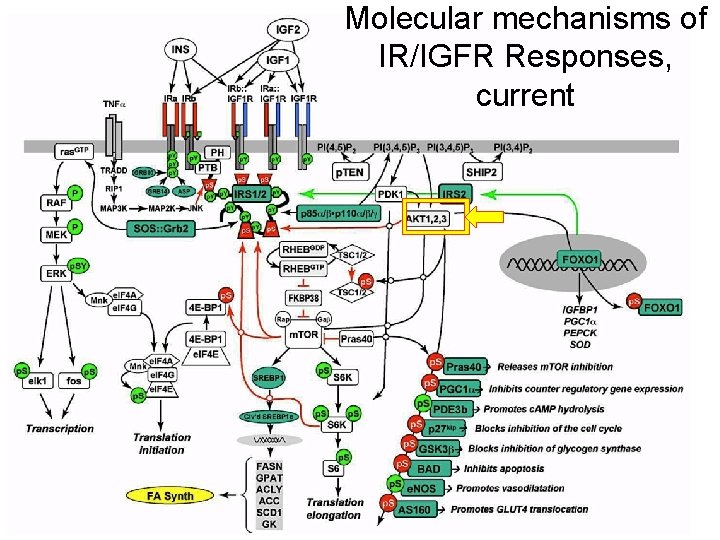
Molecular mechanisms of IR/IGFR Responses, current 9

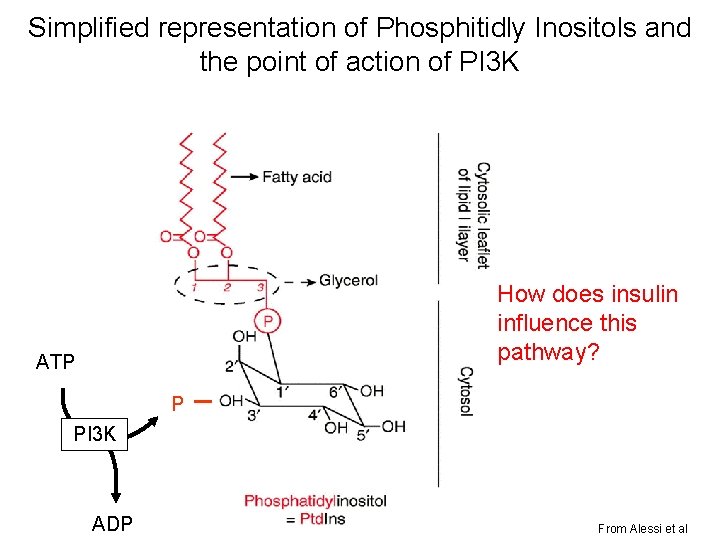
Simplified representation of Phosphitidly Inositols and the point of action of PI 3 K How does insulin influence this pathway? ATP P PI 3 K ADP From Alessi et al

IRS can bring PI 3 K to membrane; but how does this lead to activation of Akt/PKB?

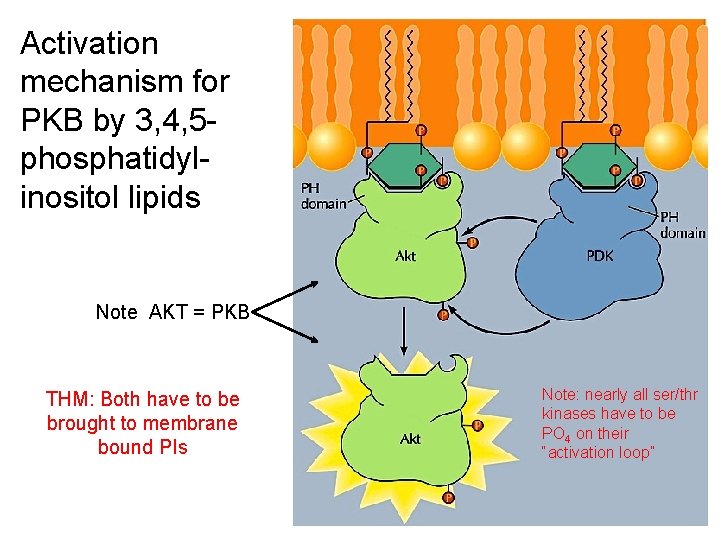
Activation mechanism for PKB by 3, 4, 5 phosphatidylinositol lipids Note AKT = PKB THM: Both have to be brought to membrane bound PIs Note: nearly all ser/thr kinases have to be PO 4 on their “activation loop”

What are some of the important pathways that are regulated by Insulin? Example of glucose transport in a cell type specific manner

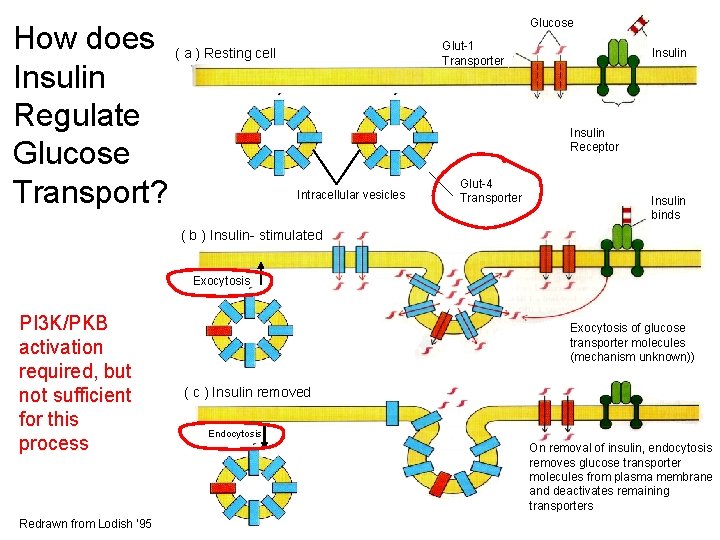
How does ( a ) Resting cell Insulin Regulate Glucose Transport? Glucose Glut-1 Transporter Insulin Receptor Intracellular vesicles Glut-4 Transporter Insulin binds ( b ) Insulin- stimulated Exocytosis PI 3 K/PKB activation required, but not sufficient for this process Redrawn from Lodish ‘ 95 Exocytosis of glucose transporter molecules (mechanism unknown)) ( c ) Insulin removed Endocytosis On removal of insulin, endocytosis removes glucose transporter molecules from plasma membrane and deactivates remaining transporters

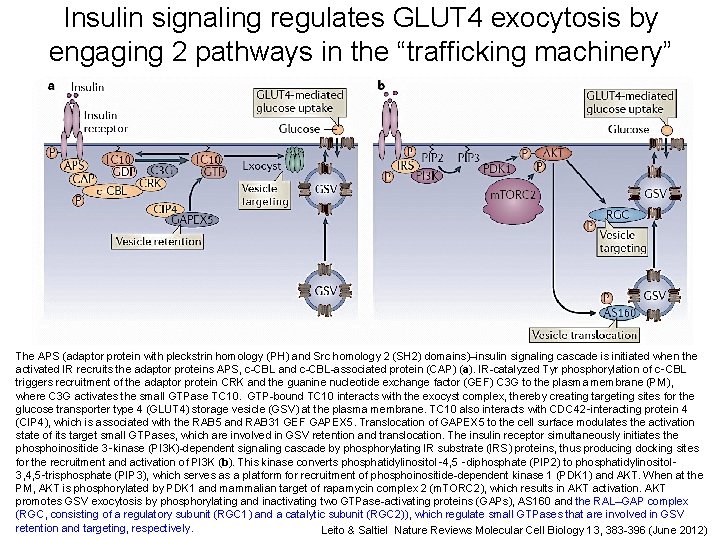
Insulin signaling regulates GLUT 4 exocytosis by engaging 2 pathways in the “trafficking machinery” The APS (adaptor protein with pleckstrin homology (PH) and Src homology 2 (SH 2) domains)–insulin signaling cascade is initiated when the activated IR recruits the adaptor proteins APS, c-CBL and c-CBL-associated protein (CAP) (a). IR-catalyzed Tyr phosphorylation of c‑CBL triggers recruitment of the adaptor protein CRK and the guanine nucleotide exchange factor (GEF) C 3 G to the plasma membrane (PM), where C 3 G activates the small GTPase TC 10. GTP-bound TC 10 interacts with the exocyst complex, thereby creating targeting sites for the glucose transporter type 4 (GLUT 4) storage vesicle (GSV) at the plasma membrane. TC 10 also interacts with CDC 42‑interacting protein 4 (CIP 4), which is associated with the RAB 5 and RAB 31 GEF GAPEX 5. Translocation of GAPEX 5 to the cell surface modulates the activation state of its target small GTPases, which are involved in GSV retention and translocation. The insulin receptor simultaneously initiates the phosphoinositide 3‑kinase (PI 3 K)-dependent signaling cascade by phosphorylating IR substrate (IRS) proteins, thus producing docking sites for the recruitment and activation of PI 3 K (b). This kinase converts phosphatidylinositol‑ 4, 5 ‑diphosphate (PIP 2) to phosphatidylinositol‑ 3, 4, 5‑trisphosphate (PIP 3), which serves as a platform for recruitment of phosphoinositide-dependent kinase 1 (PDK 1) and AKT. When at the PM, AKT is phosphorylated by PDK 1 and mammalian target of rapamycin complex 2 (m. TORC 2), which results in AKT activation. AKT promotes GSV exocytosis by phosphorylating and inactivating two GTPase-activating proteins (GAPs), AS 160 and the RAL–GAP 15 complex (RGC, consisting of a regulatory subunit (RGC 1) and a catalytic subunit (RGC 2)), which regulate small GTPases that are involved in GSV retention and targeting, respectively. Leito & Saltiel Nature Reviews Molecular Cell Biology 13, 383 -396 (June 2012)

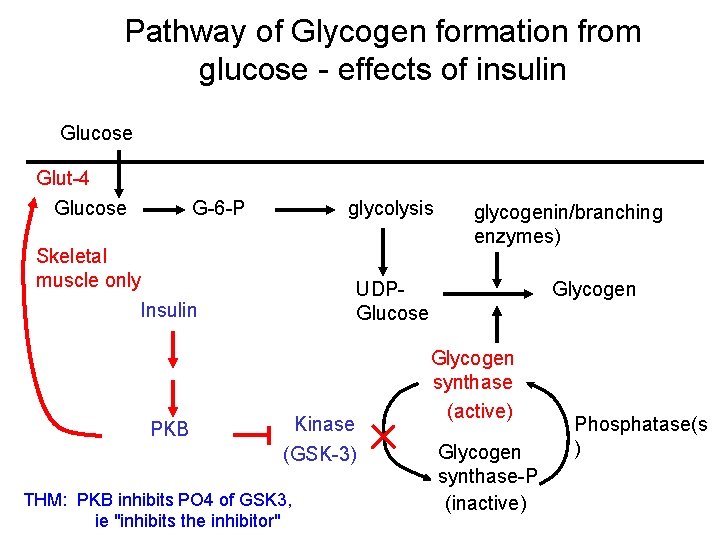
Pathway of Glycogen formation from glucose - effects of insulin Glucose Glut-4 Glucose G-6 -P glycolysis Skeletal muscle only UDPGlucose Insulin PKB glycogenin/branching enzymes) Kinase (GSK-3) THM: PKB inhibits PO 4 of GSK 3, ie "inhibits the inhibitor" Glycogen synthase (active) Glycogen synthase-P (inactive) Phosphatase(s )

Unique substrate specificity of GSK 3 THM: often a priming phosphorylation – Why is GSK a substrate? Nature Reviews Molecular Cell Biology 2; 769 -776

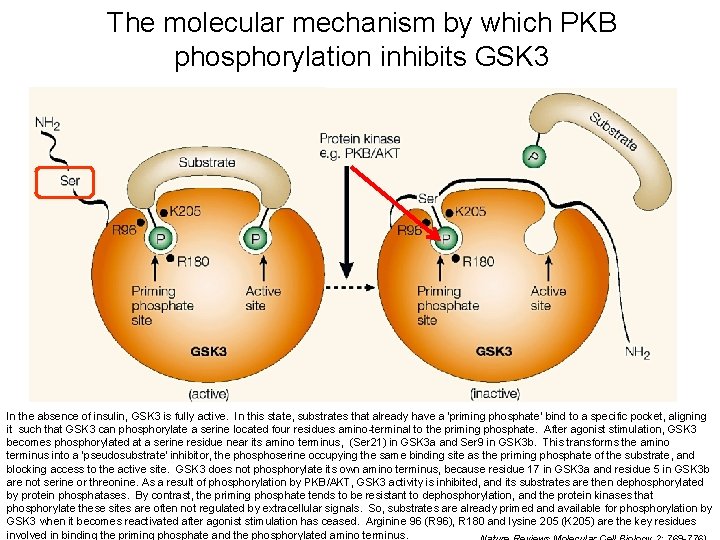
Does this also give us a hint how phosphorylation of GSK 3 by PKB inhibits activity? Yes, PO 4 interferes with substrate binding

The molecular mechanism by which PKB phosphorylation inhibits GSK 3 In the absence of insulin, GSK 3 is fully active. In this state, substrates that already have a 'priming phosphate' bind to a specific pocket, aligning it such that GSK 3 can phosphorylate a serine located four residues amino-terminal to the priming phosphate. After agonist stimulation, GSK 3 becomes phosphorylated at a serine residue near its amino terminus, (Ser 21) in GSK 3 a and Ser 9 in GSK 3 b. This transforms the amino terminus into a 'pseudosubstrate' inhibitor, the phoserine occupying the same binding site as the priming phosphate of the substrate, and blocking access to the active site. GSK 3 does not phosphorylate its own amino terminus, because residue 17 in GSK 3 a and residue 5 in GSK 3 b are not serine or threonine. As a result of phosphorylation by PKB/AKT, GSK 3 activity is inhibited, and its substrates are then dephosphorylated by protein phosphatases. By contrast, the priming phosphate tends to be resistant to dephosphorylation, and the protein kinases that 19 phosphorylate these sites are often not regulated by extracellular signals. So, substrates are already primed and available for phosphorylation by GSK 3 when it becomes reactivated after agonist stimulation has ceased. Arginine 96 (R 96), R 180 and lysine 205 (K 205) are the key residues involved in binding the priming phosphate and the phosphorylated amino terminus.

How do we determine the relative importance of the various pathway components? Conditional and Tissue-specific Mammalian gene disruption

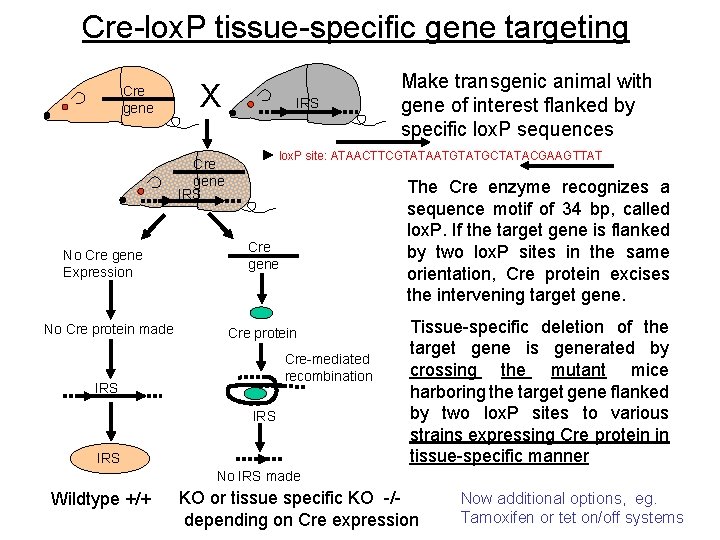
Cre-lox. P tissue-specific gene targeting Cre gene X IRS lox. P site: ATAACTTCGTATAATGTATGCTATACGAAGTTAT Cre gene IRS No Cre gene Expression No Cre protein made Make transgenic animal with gene of interest flanked by specific lox. P sequences The Cre enzyme recognizes a sequence motif of 34 bp, called lox. P. If the target gene is flanked by two lox. P sites in the same orientation, Cre protein excises the intervening target gene. Cre gene Cre protein Cre-mediated recombination IRS IRS Tissue-specific deletion of the target gene is generated by crossing the mutant mice harboring the target gene flanked by two lox. P sites to various strains expressing Cre protein in tissue-specific manner No IRS made Wildtype +/+ KO or tissue specific KO -/depending on Cre expression Now additional options, eg. 21 Tamoxifen or tet on/off systems

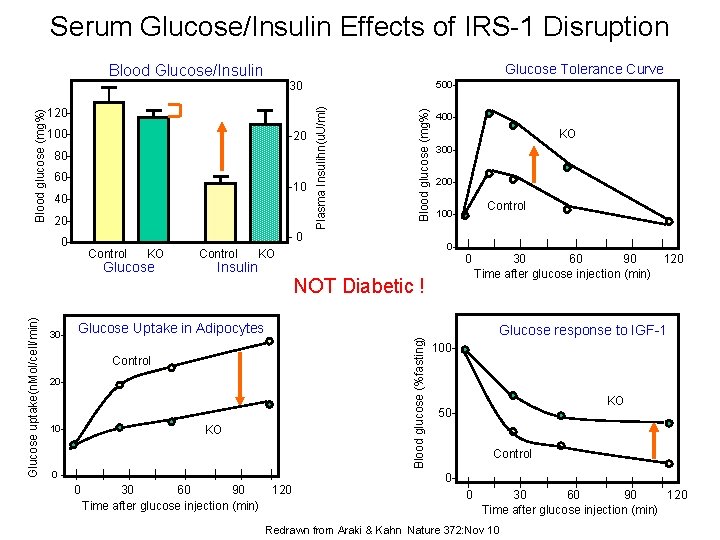
Serum Glucose/Insulin Effects of IRS-1 Disruption Glucose Tolerance Curve 500 - 120100 - -20 8060 - -10 4020 - Blood glucose (mg%) 30 Plasma Insulihn(u. U/ml) Blood glucose (mg%) Blood Glucose/Insulin -0 0 - Control KO Glucose Control 400 - KO 300200 - 0 - KO Control 100 - Insulin | 0 30 - Control 20 - 10 - 0 - | | | 30 60 90 120 Time after glucose injection (min) Glucose Uptake in Adipocytes KO | 0 | | | | 30 60 90 120 Time after glucose injection (min) Blood glucose (%fasting) Glucose uptake(n. Mol/cell/min) NOT Diabetic ! | Glucose response to IGF-1 100 - KO 50 Control 0 - | 0 | | | | 30 60 90 120 Time after glucose injection (min) Redrawn from Araki & Kahn Nature 372: Nov 10

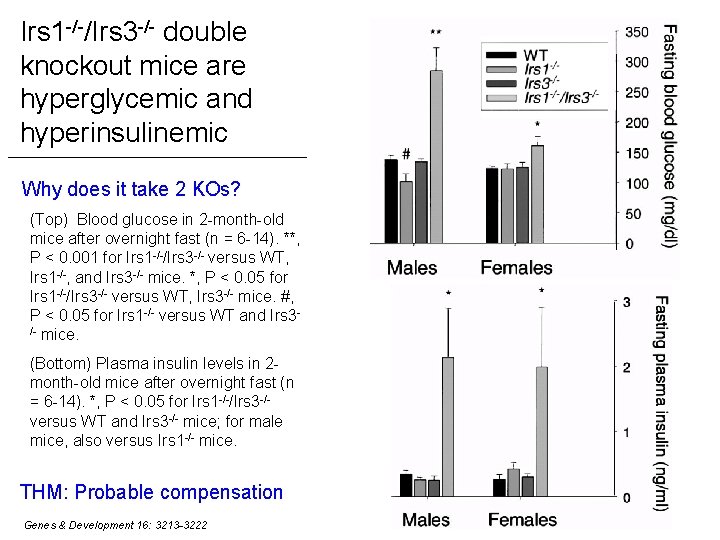
Irs 1 -/-/Irs 3 -/- double knockout mice are hyperglycemic and hyperinsulinemic Why does it take 2 KOs? (Top) Blood glucose in 2 -month-old mice after overnight fast (n = 6 -14). **, P < 0. 001 for Irs 1 -/-/Irs 3 -/- versus WT, Irs 1 -/-, and Irs 3 -/- mice. *, P < 0. 05 for Irs 1 -/-/Irs 3 -/- versus WT, Irs 3 -/- mice. #, P < 0. 05 for Irs 1 -/- versus WT and Irs 3/- mice. (Bottom) Plasma insulin levels in 2 month-old mice after overnight fast (n = 6 -14). *, P < 0. 05 for Irs 1 -/-/Irs 3 -/versus WT and Irs 3 -/- mice; for male mice, also versus Irs 1 -/- mice. THM: Probable compensation Genes & Development 16: 3213 -3222 23

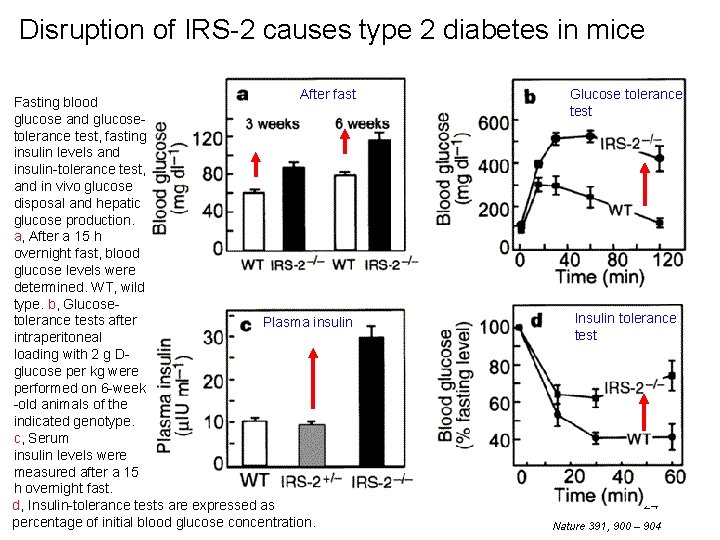
Disruption of IRS-2 causes type 2 diabetes in mice After fast Fasting blood glucose and glucosetolerance test, fasting insulin levels and insulin-tolerance test, and in vivo glucose disposal and hepatic glucose production. a, After a 15 h overnight fast, blood glucose levels were determined. WT, wild type. b, Glucosetolerance tests after Plasma insulin intraperitoneal loading with 2 g Dglucose per kg were performed on 6 -week -old animals of the indicated genotype. c, Serum insulin levels were measured after a 15 h overnight fast. d, Insulin-tolerance tests are expressed as percentage of initial blood glucose concentration. Glucose tolerance test Insulin tolerance test 24 Nature 391, 900 – 904

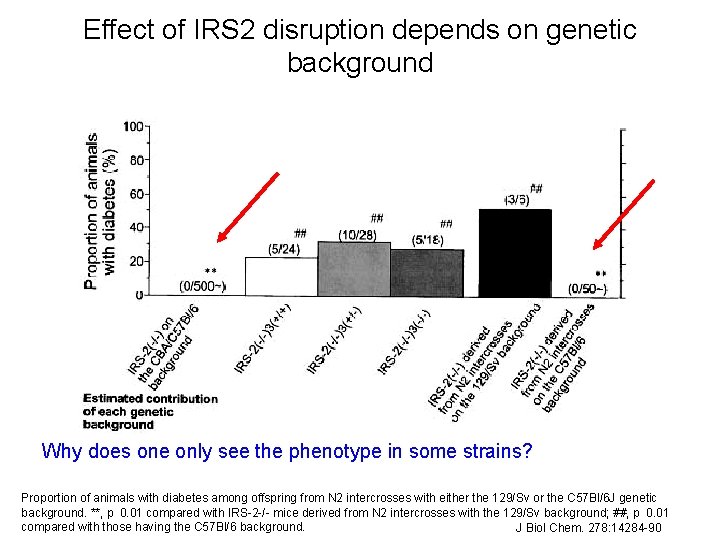
Effect of IRS 2 disruption depends on genetic background Why does one only see the phenotype in some strains? Proportion of animals with diabetes among offspring from N 2 intercrosses with either the 129/Sv or the C 57 Bl/6 J genetic 25 background. **, p 0. 01 compared with IRS-2 -/- mice derived from N 2 intercrosses with the 129/Sv background; ##, p 0. 01 compared with those having the C 57 Bl/6 background. J Biol Chem. 278: 14284 -90

What are phenotypes for tissue specific IR and IRS Knockouts? Are major roles of IRS proteins tissue specific? Why do tissue specific knockouts?

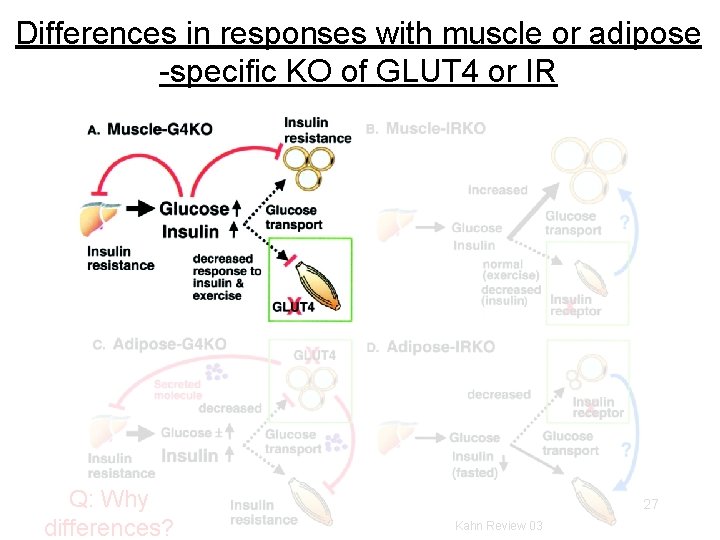
Differences in responses with muscle or adipose -specific KO of GLUT 4 or IR Q: Why differences? 27 Kahn Review 03

Advantages of tissue specific knockouts Tissue specific knockouts allow one to begin to assess in a quantitative manner the relative contribution of various parts of complex pathway to overall phenotype. Also allows many but not all developmental issues to be addressed

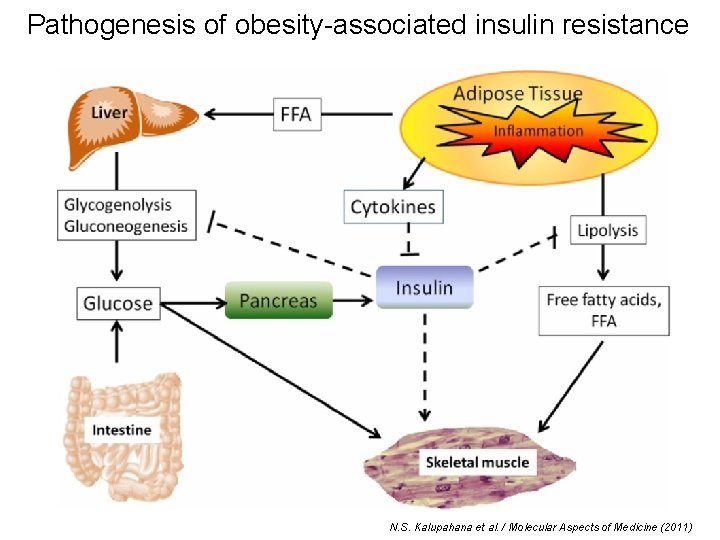
Pathogenesis of obesity-associated insulin resistance N. S. Kalupahana et al. / Molecular Aspects of Medicine (2011)

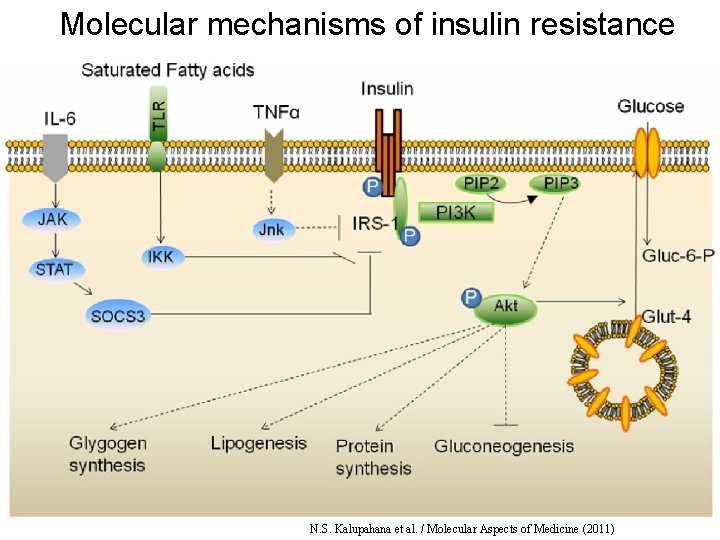
Molecular mechanisms of insulin resistance N. S. Kalupahana et al. / Molecular Aspects of Medicine (2011)

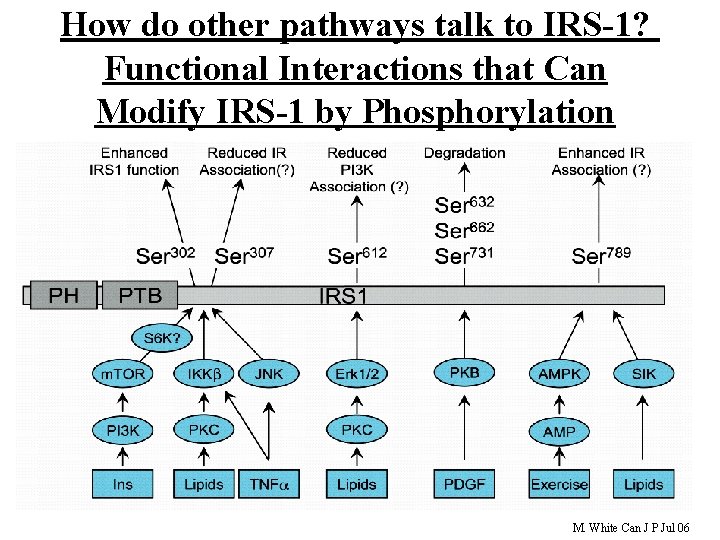
How do other pathways talk to IRS-1? Functional Interactions that Can Modify IRS-1 by Phosphorylation 31 M. White Can J P Jul 06

Beavo Take Home Question 2013 Please read the posted review by Kahn and the research paper by James. 1) For the James’ paper briefly outline your opinion about at least two strengths of the approach that they are using to help further refine the well-studied insulin signaling pathway. 2) Identify what you feel are at least two limitations of the approach that they use and briefly explain your reasoning for saying so. Your answer therefore should have two parts and each part will be graded with approximately equal weight. (One page total please).

‘nuff for now

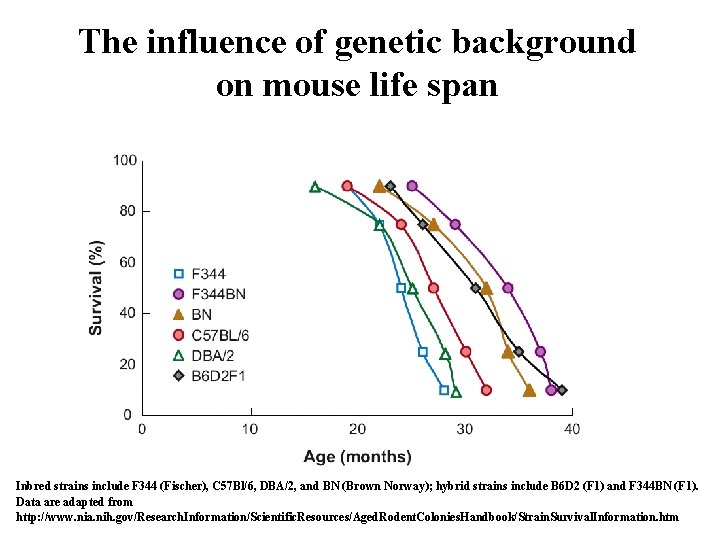
The influence of genetic background on mouse life span Inbred strains include F 344 (Fischer), C 57 Bl/6, DBA/2, and BN (Brown Norway); hybrid strains include B 6 D 2 (F 1) and F 344 BN (F 1). Data are adapted from 34 http: //www. nia. nih. gov/Research. Information/Scientific. Resources/Aged. Rodent. Colonies. Handbook/Strain. Survival. Information. htm

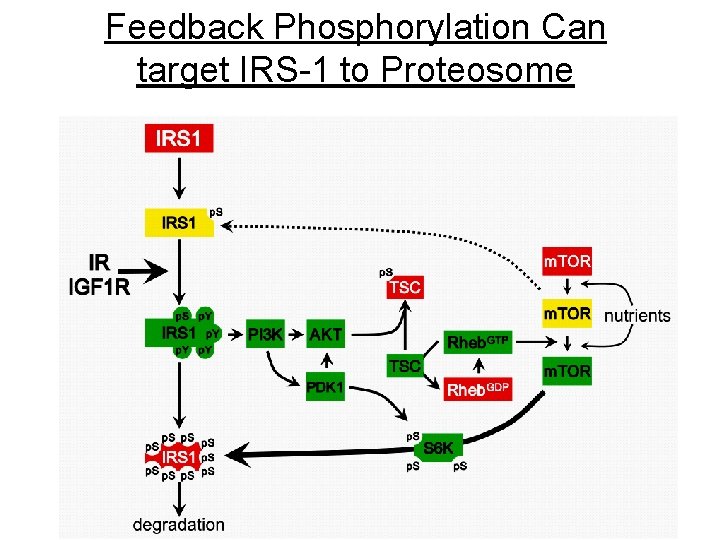
Feedback Phosphorylation Can target IRS-1 to Proteosome 35

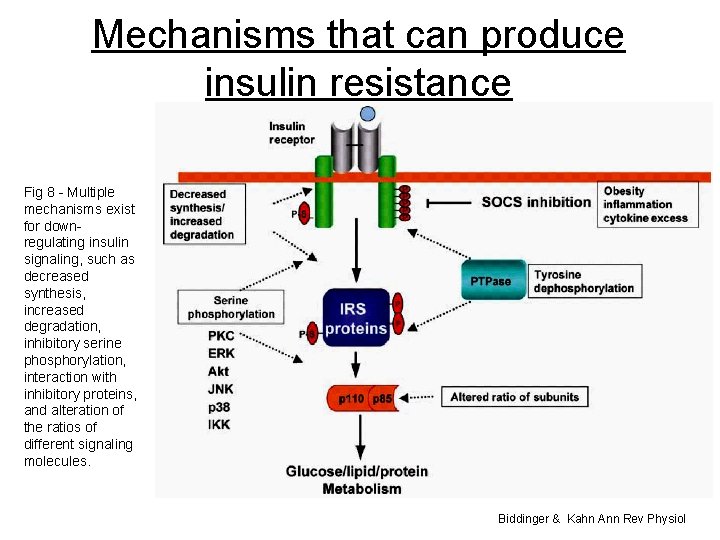
Mechanisms that can produce insulin resistance Fig 8 - Multiple mechanisms exist for downregulating insulin signaling, such as decreased synthesis, increased degradation, inhibitory serine phosphorylation, interaction with inhibitory proteins, and alteration of the ratios of different signaling molecules. Biddinger & Kahn Ann Rev Physiol

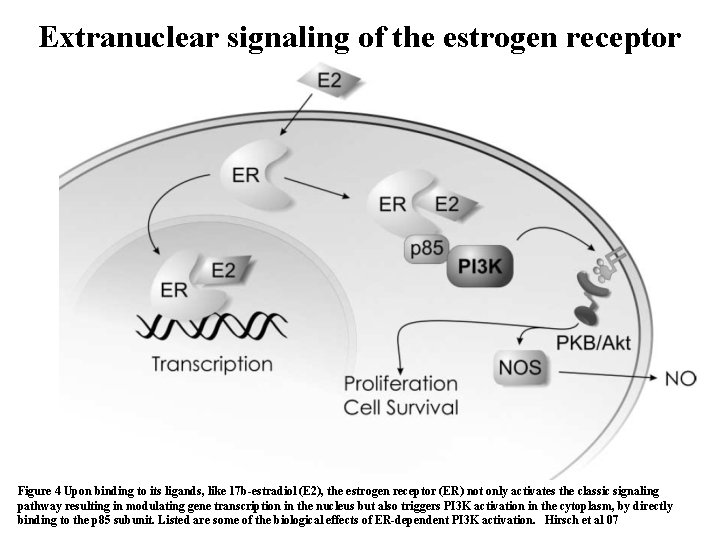
Extranuclear signaling of the estrogen receptor Figure 4 Upon binding to its ligands, like 17 b-estradiol (E 2), the estrogen receptor (ER) not only activates the classic signaling 37 pathway resulting in modulating gene transcription in the nucleus but also triggers PI 3 K activation in the cytoplasm, by directly binding to the p 85 subunit. Listed are some of the biological effects of ER-dependent PI 3 K activation. Hirsch et al 07

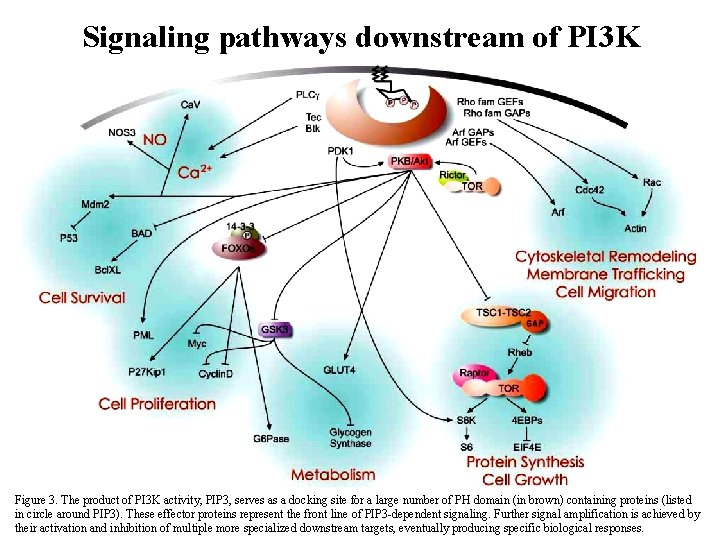
Signaling pathways downstream of PI 3 K Figure 3. The product of PI 3 K activity, PIP 3, serves as a docking site for a large number of PH domain (in brown) containing proteins 38 (listed in circle around PIP 3). These effector proteins represent the front line of PIP 3 -dependent signaling. Further signal amplification is achieved by their activation and inhibition of multiple more specialized downstream targets, eventually producing specific biological responses.

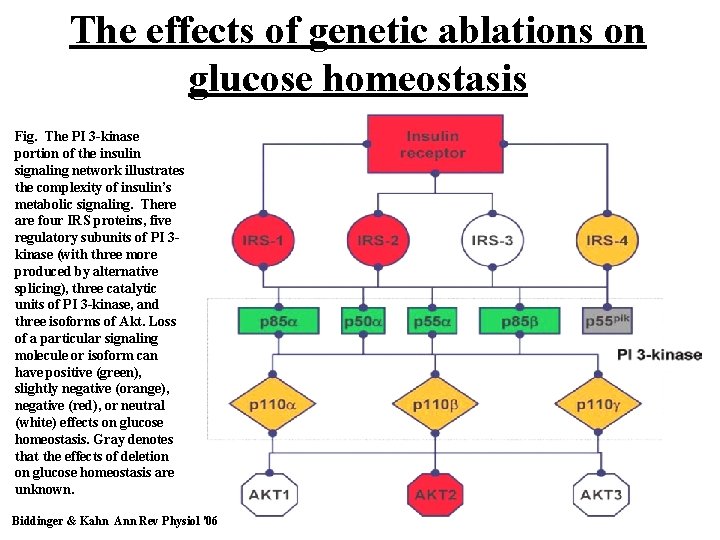
The effects of genetic ablations on glucose homeostasis Fig. The PI 3 -kinase portion of the insulin signaling network illustrates the complexity of insulin’s metabolic signaling. There are four IRS proteins, five regulatory subunits of PI 3 kinase (with three more produced by alternative splicing), three catalytic units of PI 3 -kinase, and three isoforms of Akt. Loss of a particular signaling molecule or isoform can have positive (green), slightly negative (orange), negative (red), or neutral (white) effects on glucose homeostasis. Gray denotes that the effects of deletion on glucose homeostasis are unknown. Biddinger & Kahn Ann Rev Physiol '06 39

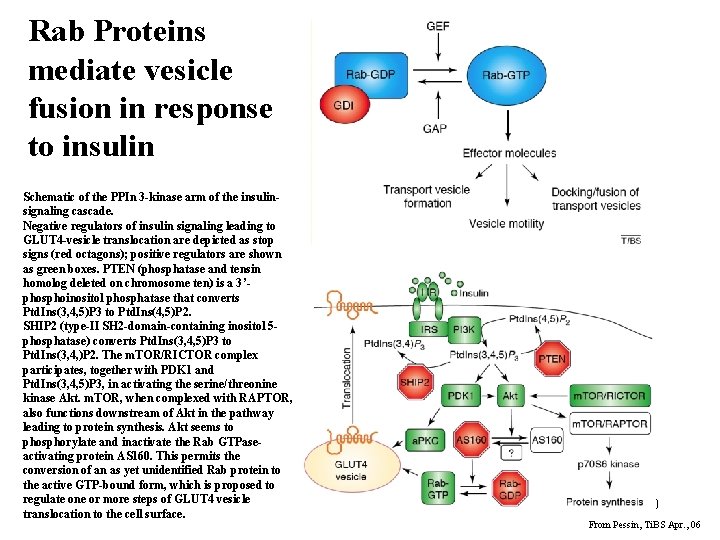
Rab Proteins mediate vesicle fusion in response to insulin Schematic of the PPIn 3 -kinase arm of the insulinsignaling cascade. Negative regulators of insulin signaling leading to GLUT 4 -vesicle translocation are depicted as stop signs (red octagons); positive regulators are shown as green boxes. PTEN (phosphatase and tensin homolog deleted on chromosome ten) is a 3’phosphoinositol phosphatase that converts Ptd. Ins(3, 4, 5)P 3 to Ptd. Ins(4, 5)P 2. SHIP 2 (type-II SH 2 -domain-containing inositol 5 phosphatase) converts Ptd. Ins(3, 4, 5)P 3 to Ptd. Ins(3, 4, )P 2. The m. TOR/RICTOR complex participates, together with PDK 1 and Ptd. Ins(3, 4, 5)P 3, in activating the serine/threonine kinase Akt. m. TOR, when complexed with RAPTOR, also functions downstream of Akt in the pathway leading to protein synthesis. Akt seems to phosphorylate and inactivate the Rab GTPaseactivating protein AS 160. This permits the conversion of an as yet unidentified Rab protein to the active GTP-bound form, which is proposed to regulate one or more steps of GLUT 4 vesicle translocation to the cell surface. 40 From Pessin, Ti. BS Apr. , 06

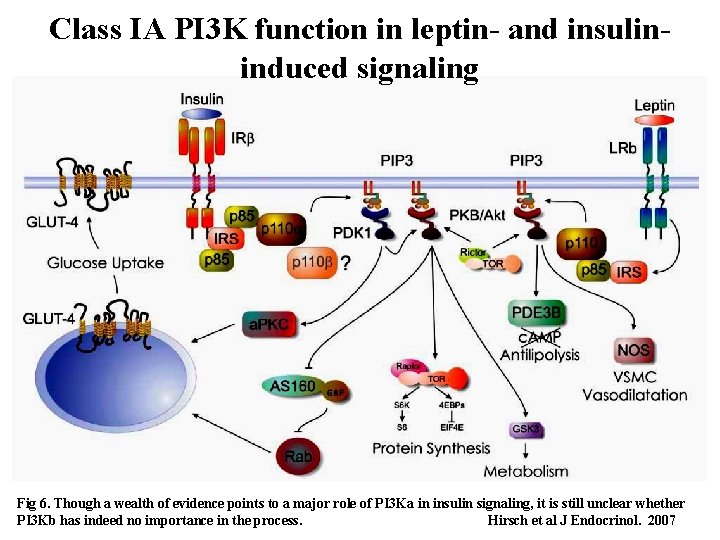
Class IA PI 3 K function in leptin- and insulininduced signaling Fig 6. Though a wealth of evidence points to a major role of PI 3 Ka in insulin signaling, it is still unclear whether 41 PI 3 Kb has indeed no importance in the process. Hirsch et al J Endocrinol. 2007

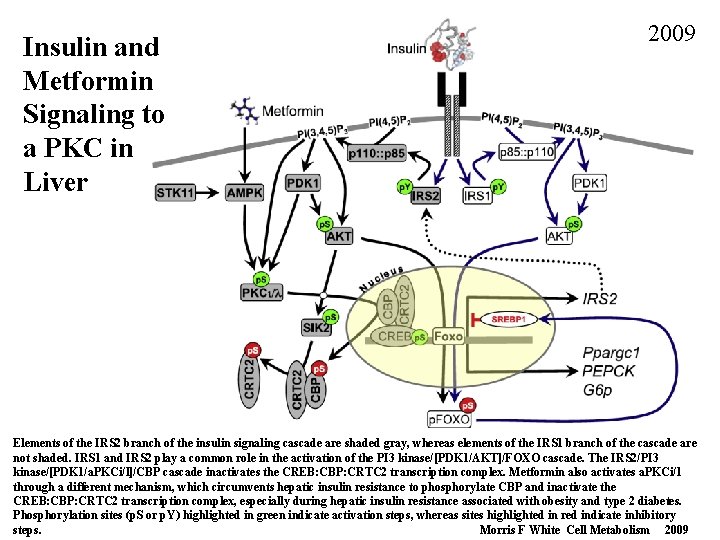
Insulin and Metformin Signaling to a PKC in Liver 2009 Elements of the IRS 2 branch of the insulin signaling cascade are shaded gray, whereas elements of the IRS 1 branch of the cascade are not shaded. IRS 1 and IRS 2 play a common role in the activation of the PI 3 kinase/[PDK 1/AKT]/FOXO cascade. The IRS 2/PI 3 kinase/[PDK 1/a. PKCi/l]/CBP cascade inactivates the CREB: CBP: CRTC 2 transcription complex. Metformin also activates a. PKCi/l through a different mechanism, which circumvents hepatic insulin resistance to phosphorylate CBP and inactivate the CREB: CBP: CRTC 2 transcription complex, especially during hepatic insulin resistance associated with obesity and type 2 diabetes. Phosphorylation sites (p. S or p. Y) highlighted in green indicate activation steps, whereas sites highlighted in red indicate inhibitory steps. Morris F White Cell Metabolism 2009

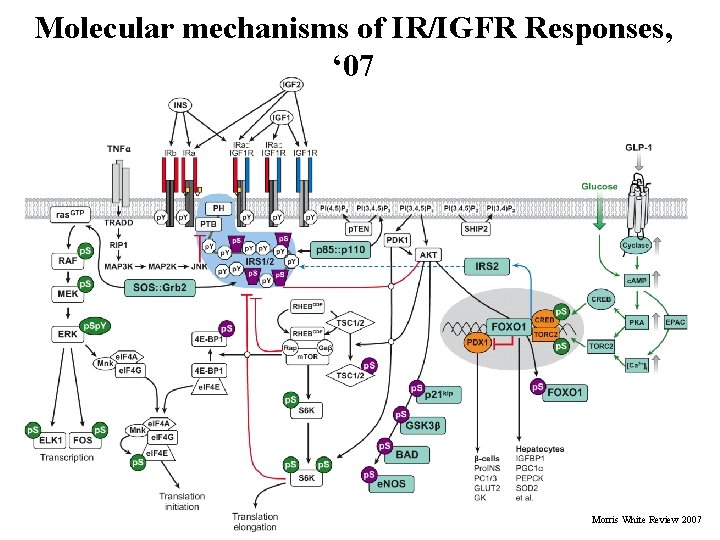
Molecular mechanisms of IR/IGFR Responses, ‘ 07 43 Morris White Review 2007

Legend for previous figure - The mammalian insulin-like signaling cascade, ‘ 07/’ 08 In mammals, the insulin/insulin-like growth factor (IGF) family consists of three hormones: insulin, insulin-like growth factor 1 (IGF 1), and insulin-like growth factor 2 (IGF 2). These peptide ligands bind as indicated to five distinct receptor isoforms that generate cytoplasmic signals: two insulin receptor isoforms, IRa and IRb (red ); the insulin like growth factor receptor, IGF 1 R (blue); and two hybrid receptors, IRa: : IGF 1 R and IRb: : IGF 1 R. IGF 2 also binds to the mannose-6 -phosphate receptor, which mediates the endocytosis and degradation of IGF 2 (not shown). Activation of the receptors promotes tyrosine phosphorylation of cellular proteins—IRS 1 and IRS 2 occur in nearly all cells, but other substrates exist in specific cells and tissues. The SOS/GRB 2 branch activates the ras→RAF→MEK→ERK 1/2 cascade. Activated ERK stimulates transcriptional activity by direct phosphorylation of ELK 1 and by phosphorylation of FOS through p 90 rsk (RSK). Activation of PI 3 -kinase during recruitment to IRS 1/2 produces PI(3, 4)P 2 and PI(3, 4, 5)P 3 (antagonized by the action of PTEN or SHIP 2), which recruit PDK 1 and AKT to the plasma membrane, where AKT is activated by PDK-mediated phosphorylation. AKT phosphorylates many substrates, including the cyclin-dependent kinase inhibitor p 21 kip, GSK 3β, BAD, e. NOS, and FOXO 1; phosphorylation of FOXO 1 inactivates this transcription factor and causes its sequestration in the cytosol, which alters the expression of many genes, including those encoding IGFBP 1, PGC 1α, PEPCK, and SOD 2. The m. TOR kinase is activated by RHEBGTP, which accumulates upon inhibition of the GAP activity of the TSC 1: : TSC 2 complex by AKT-mediated phosphorylation. The p 70 s 6 k is primed for activation by PDK 1 -mediated phosphorylation and ultimately activated by m. TOR-mediated phosphorylation. Insulin stimulates protein synthesis by altering the intrinsic activity or binding properties of key translation initiation and elongation factors (e. IFs and e. EFs, respectively), including e. IF 4 A and e. IF 4 G. In particular, phosphorylation of 4 E-BP 1 releases e. IF 4 E to form an active complex promoting translation initiation. In β-cells, GLP-1 and glucose promote Ca 2+ and c. AMP-mediated signaling that stimulates IRS 2 expression through CREB and TORC 2. Regarding phosphorylation depicted, green indicates stimulatory phosphorylation, and purple represents inhibitory phosphorylation. Abbreviations: AKT, protein kinase B; BAD, Bcl-2 -associated death promoter; c. AMP, cyclic-3 -5 -adenosine monophosphate; CREB, c. AMP response element (CRE) binding protein; cyclase, adenylylcyclase; e. NOS, endothelial nitric oxide synthase; EPAC, exchange protein activated by c. AMP; GAP, guanosine triphosphatase–associated protein; GLP 1, glucagon-like peptide 1; GRB 2, factor receptor binding protein 2; GSK 3β, glycogen synthase kinase 3β; IGFBP 1, IGF binding protein-1; INS, insulinoma; IRS 1, insulin receptor substrate 1; JNK, Jun NH 2 -terminal kinase; MAP 2 K, MAPK kinase; MAP 3 K, MAPK kinase; m. TOR, mammalian target of rapamycin; PDK, PI-dependent protein kinase; PDX 1: pancreatic and duodenal homeobox factor 1; PEPCK, phosphoenolpyruvate carboxykinase; PGC 1α, peroxisome proliferator–activated receptor (PPAR) gamma coactivator 1 alpha; PH, pleckstrin homology domain; PKA, protein kinase A; PTB, phosphotyrosine-binding domain; PTEN and SHIP 2, phospholipid phosphatases; RIP 1, Ser/Thr protein kinase receptor–interacting protein; SOD 2, superoxide dismutase 2; SOS, son 44 death of-sevenless; TORC 2, transducer of regulated CREB activity 2; TRADD: tumor necrosis factor receptor (TNFR)-associated domain; TSC, tuberous sclerosis complex

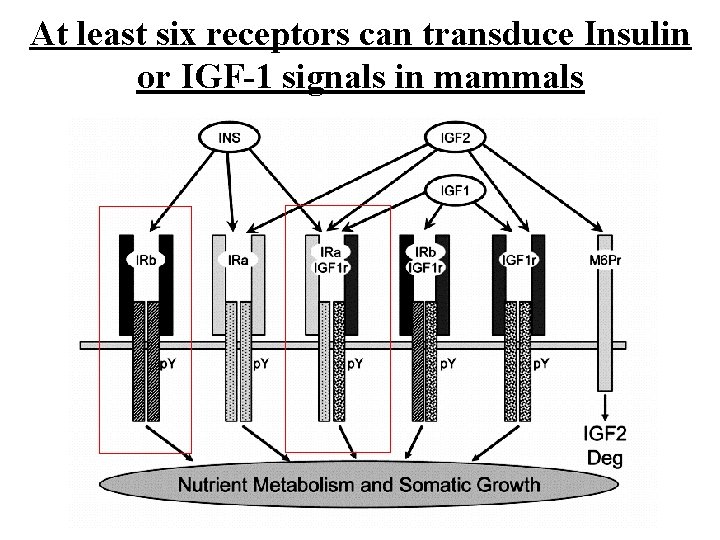
At least six receptors can transduce Insulin or IGF-1 signals in mammals 45