How does DNA evolve Nucleotide substitutions 1 ATGCCCCAACTAAATACTACCGTATGGCCCACCATAATTACCCCCATACT
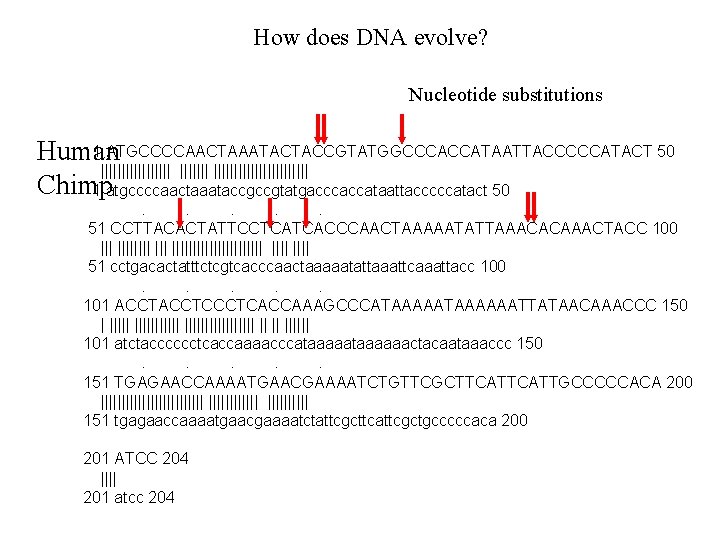
How does DNA evolve? Nucleotide substitutions 1 ATGCCCCAACTAAATACTACCGTATGGCCCACCATAATTACCCCCATACT 50 Human ||||||||||||||| Chimp 1 atgccccaactaaataccgccgtatgacccaccataattacccccatact 50. . . 51 CCTTACACTATTCCTCATCACCCAACTAAAAATATTAAACACAAACTACC 100 ||| ||||||||||||||| 51 cctgacactatttctcgtcacccaactaaaaatattaaattcaaattacc 100. . . 101 ACCTCCCTCACCAAAGCCCATAAAAAATTATAACAAACCC 150 | ||||||||||| || || |||||| 101 atctacccccctcaccaaaacccataaaaaactacaataaaccc 150. . . 151 TGAGAACCAAAATGAACGAAAATCTGTTCGCTTCATTGCCCCCACA 200 ||||||||||||| ||||| 151 tgagaaccaaaatgaacgaaaatctattcgcttcattcgctgcccccaca 200 201 ATCC 204 |||| 201 atcc 204
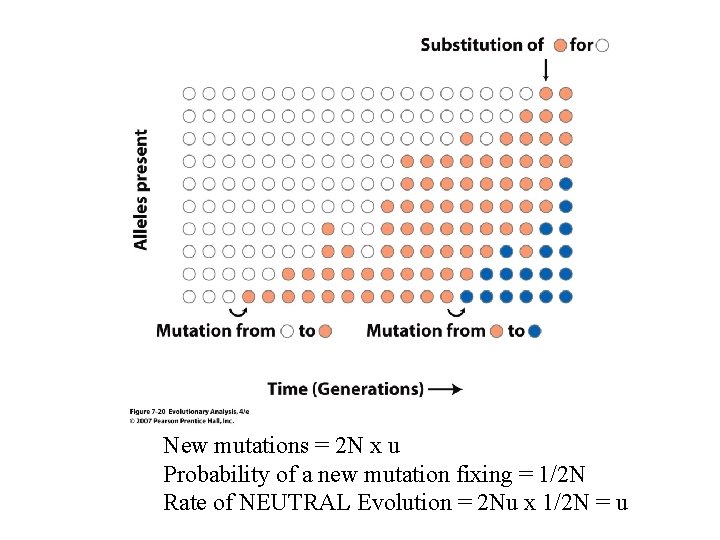
New mutations = 2 N x u Probability of a new mutation fixing = 1/2 N Rate of NEUTRAL Evolution = 2 Nu x 1/2 N = u
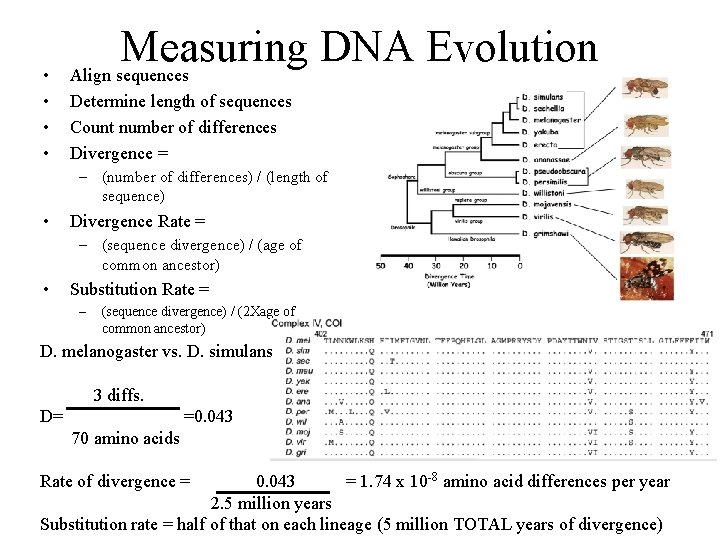
• • Measuring DNA Evolution Align sequences Determine length of sequences Count number of differences Divergence = – (number of differences) / (length of sequence) • Divergence Rate = – (sequence divergence) / (age of common ancestor) • Substitution Rate = – (sequence divergence) / (2 Xage of common ancestor) D. melanogaster vs. D. simulans 3 diffs. D= =0. 043 70 amino acids 0. 043 = 1. 74 x 10 -8 amino acid differences per year 2. 5 million years Substitution rate = half of that on each lineage (5 million TOTAL years of divergence) Rate of divergence =
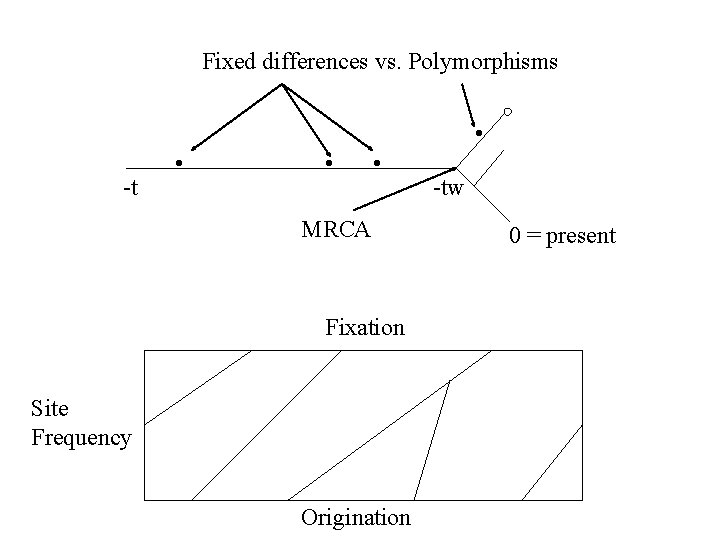
Fixed differences vs. Polymorphisms • -t • • • MRCA Fixation Site Frequency Origination -tw 0 = present
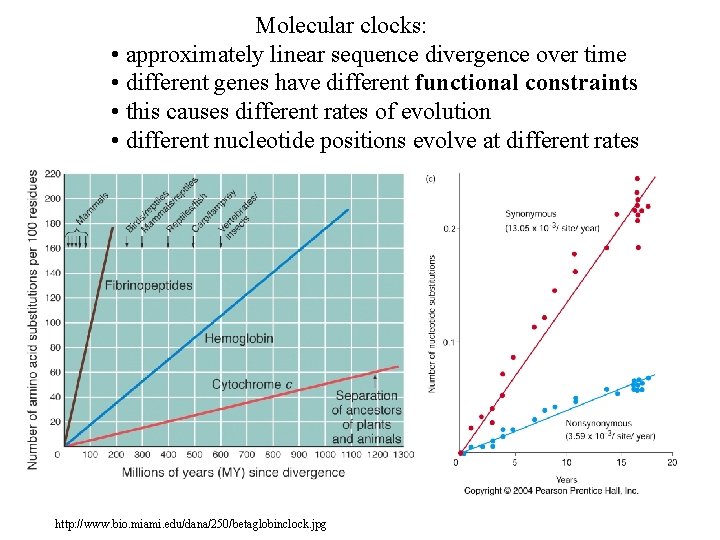
Molecular clocks: • approximately linear sequence divergence over time • different genes have different functional constraints • this causes different rates of evolution • different nucleotide positions evolve at different rates http: //www. bio. miami. edu/dana/250/betaglobinclock. jpg
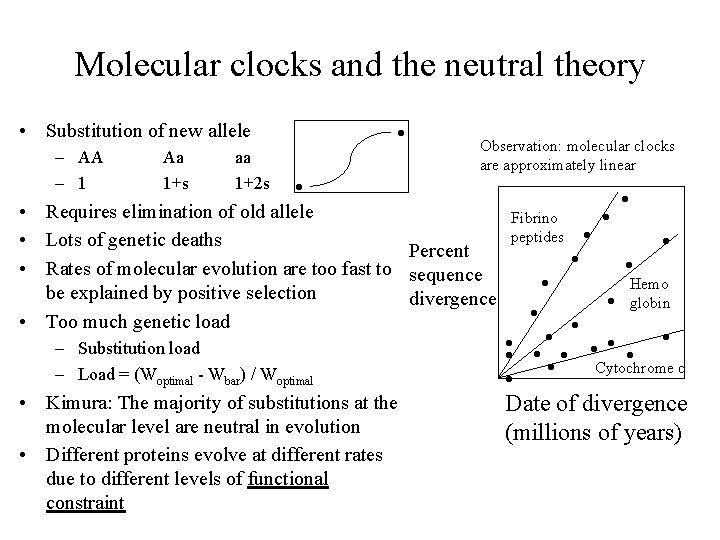
Molecular clocks and the neutral theory • Substitution of new allele – AA – 1 Aa 1+s aa 1+2 s • • Observation: molecular clocks are approximately linear • Requires elimination of old allele • Lots of genetic deaths Percent • Rates of molecular evolution are too fast to sequence be explained by positive selection divergence • Too much genetic load – Substitution load – Load = (Woptimal - Wbar) / Woptimal • Kimura: The majority of substitutions at the molecular level are neutral in evolution • Different proteins evolve at different rates due to different levels of functional constraint Fibrino peptides • • • Hemo globin • • • Cytochrome c • • Date of divergence (millions of years)
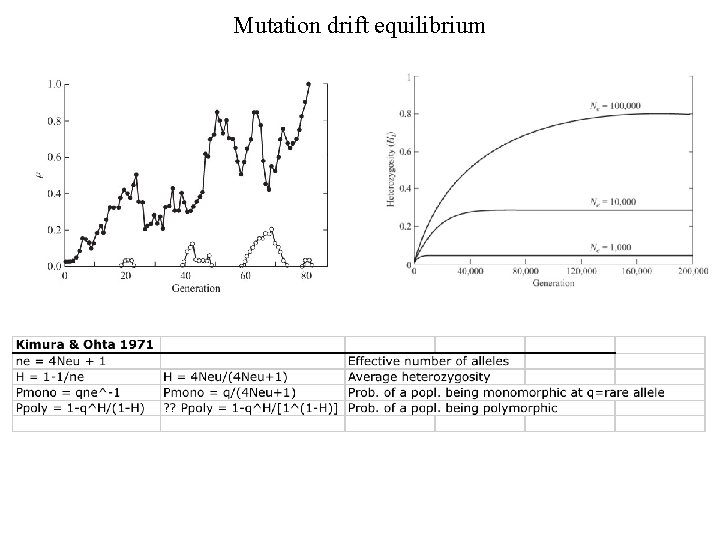
Mutation drift equilibrium

Living fossils slow rates of morphological change • Coelocanth – Only living crossopterygian – Thought to be extinct; caught in a net off Africa • Horseshoe crab • No obvious morphological change for ~150 million years – Birds evolved – Dinosaurs rose and went extinct – Mammals , land plants diversified • Horseshoe crab stasis due to life in relatively stable near-shore environment? http: //www. aqua. org/images/feb/horseshoebig. jpeg
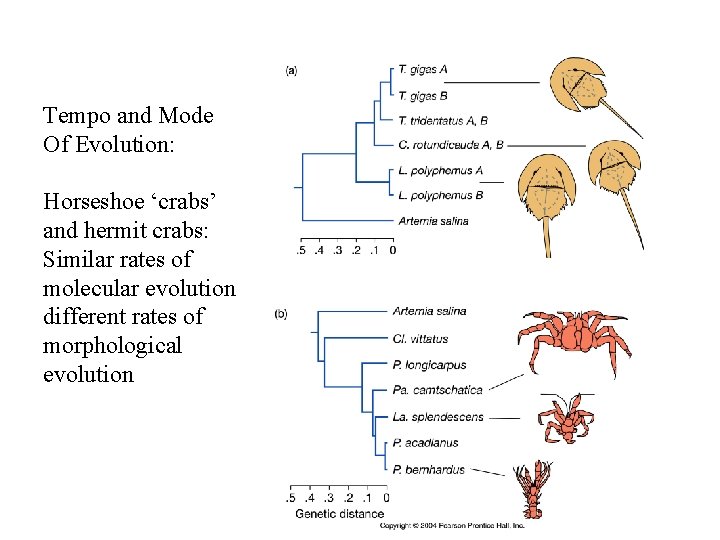
Tempo and Mode Of Evolution: Horseshoe ‘crabs’ and hermit crabs: Similar rates of molecular evolution different rates of morphological evolution
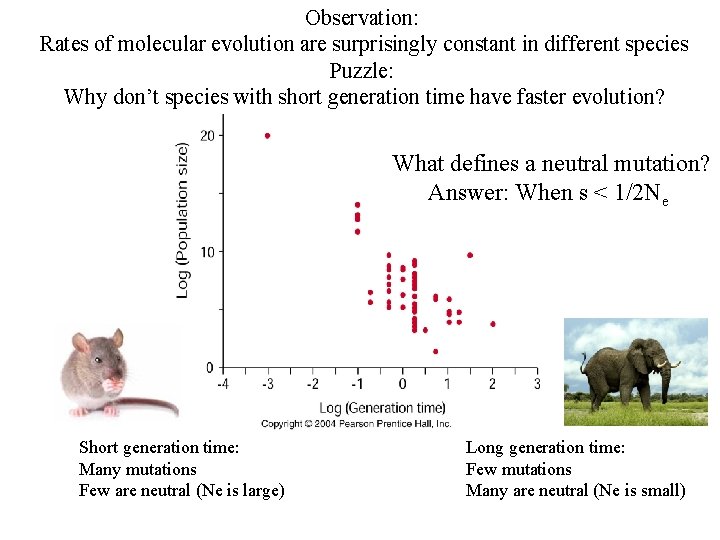
Observation: Rates of molecular evolution are surprisingly constant in different species Puzzle: Why don’t species with short generation time have faster evolution? What defines a neutral mutation? Answer: When s < 1/2 Ne Short generation time: Many mutations Few are neutral (Ne is large) Long generation time: Few mutations Many are neutral (Ne is small)

Protein polymorphism and the neutral theory • • Observation from allozyme data: About 1/4 of loci are polymorphic well – About 3000 loci in fruitflies • • Heterozygosity ~ 0. 3 on average Is variation maintained by selection? – AA – 1 -s • Aa 1 bands SS SF FF SS SF SF aa 1 -t Monomorphic enzyme This requires lots of genetic deaths Dimeric enzyme – Homozygotes eliminated • Genetic load – Segregation load – Load = (Woptimal - Wbar) / Woptimal – For 3000 loci with 10% selection (s and t = 5% each) this load could be huge – Too much genetic death to allow balancing selection • No problem if alleles are neutral – No genetic load with neutrality Geno types SS SF FF SF Kimura and Ohta: Protein polymorphism as a phase Of neutral evolution

- Slides: 13