Horton Moran Scrimgeour Perry Rawn Principles of Biochemistry

Horton • Moran • Scrimgeour • Perry • Rawn Principles of Biochemistry Fourth Edition Chapter 22 Protein Synthesis Copyright © 2006 Pearson Prentice Hall, Inc.
Chapter 22 - Protein Synthesis The ribosome, a complex of RNA and protein, is the site where genetic information is translated into protein

22. 1 The Genetic Code • Codons - three letter genetic code (nonoverlapping) • t. RNA - adapters between m. RNA and proteins • Reading frame - each potential starting point for interpreting the 3 letter code

Fig 22. 1 Overlapping vs nonoverlapping reading of the three-letter code
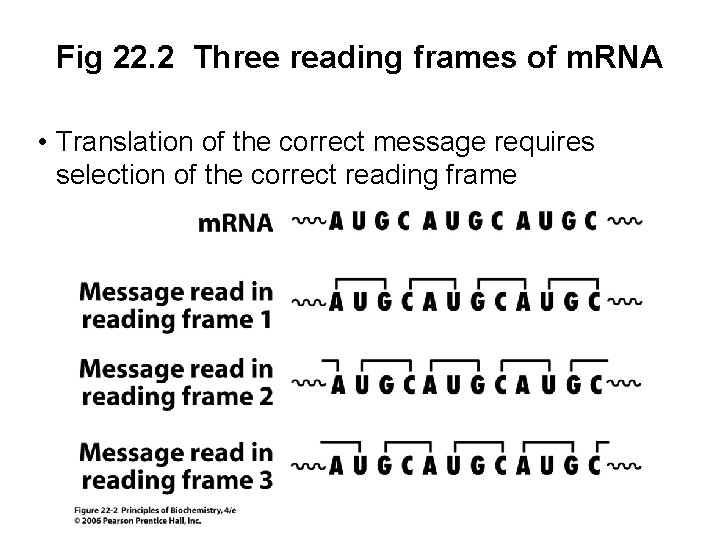
Fig 22. 2 Three reading frames of m. RNA • Translation of the correct message requires selection of the correct reading frame
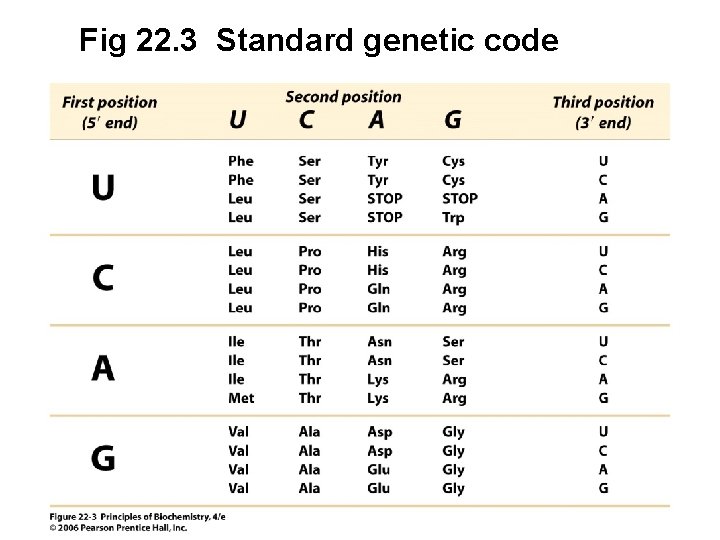
Fig 22. 3 Standard genetic code

Features of the Genetic Code 1. The genetic code is unambiguous. In any organism each codon corresponds to only one amino acid. 2. There are multiple codons for most amino acids (code is degenerate), and synonymous codons specify the same amino acid 3. The first two nucleotides of a codon are often enough to specify a given amino acid

Features of the Code (continued) 4. Codons with similar sequences specify similar amino acids 5. Only 61 of the 64 codons specify amino acids • Termination (stop codons): UAA, UGA, UAG • Initiation codon - Methionine codon (AUG) also specifies initiation site for protein synthesis

22. 2 Transfer RNA A. Three-Dimensional Structure of t. RNA – Transfer RNA molecules are the interpreters of the genetic code – Every cell must contain at least 20 t. RNA (one for every amino acid) – Each t. RNA must recognize at least one codon – t. RNAs have a “cloverleaf” type secondary structure with several loops or arms

Cloverleaf Secondary Structure of t. RNA • Figure 22. 4 (next slide) • Watson-Crick base pairing (dashed lines) • t. RNA has an acceptor stem and four arms • Conserved bases (gray)
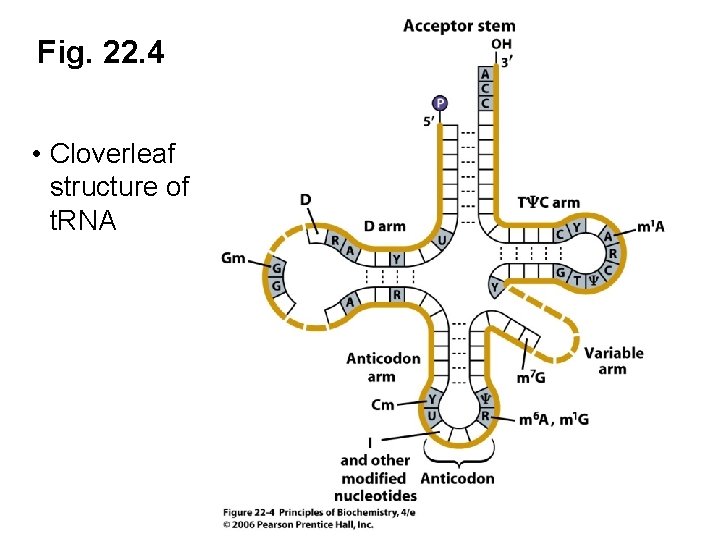
Fig. 22. 4 • Cloverleaf structure of t. RNA
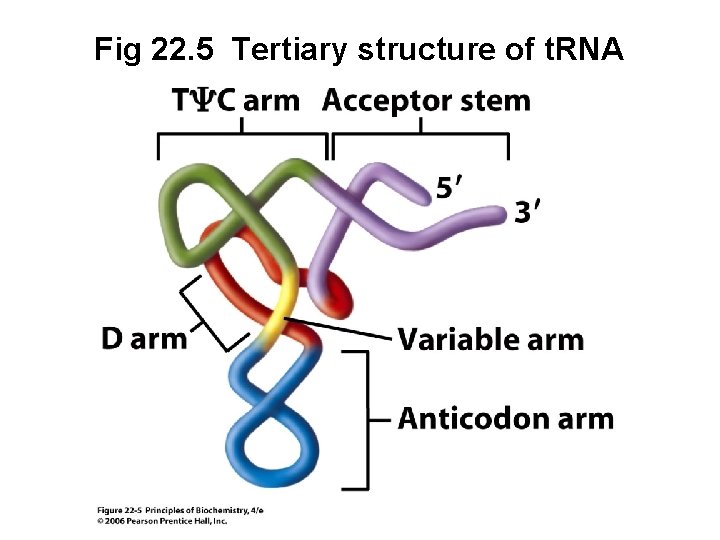
Fig 22. 5 Tertiary structure of t. RNA

t. RNA Arms • Acceptor stem - amino acid becomes covalently attached to t. RNA at the 3’ end of this stem • Anticodon arm - contains the anticodon, a three-base sequence that binds to a complementary codon in m. RNA (continued next slide)

t. RNA Arms (continued) • Ty. C arm - contains thymidylate (T) and pseudouridylate (y) followed by C • D arm - contains dihydrouridylate (D) • Variable arm - ranges from 3 -21 nucleotides
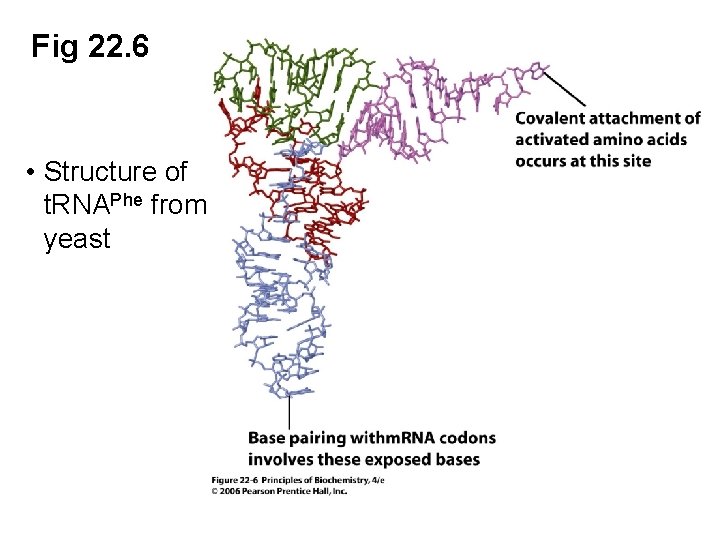
Fig 22. 6 • Structure of t. RNAPhe from yeast

Animations • Structure of t. RNA

B. t. RNA Anticodons Base-Pair with m. RNA Codons • t. RNA molecules are named for the amino acid that they carry (e. g. t. RNAPhe) • Base pairing between codon and anticodon is governed by rules of Watson-Crick (A-U, G-C) • However, the 5’ anticodon position has some flexibility in base pairing (the “wobble” position)
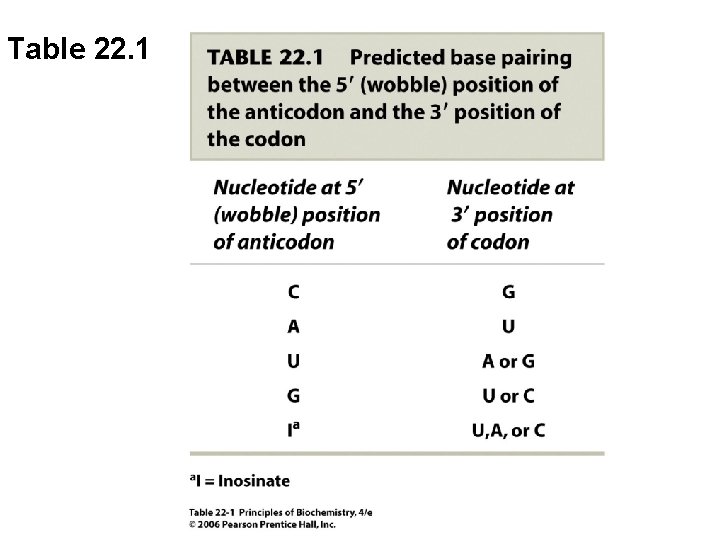
Table 22. 1
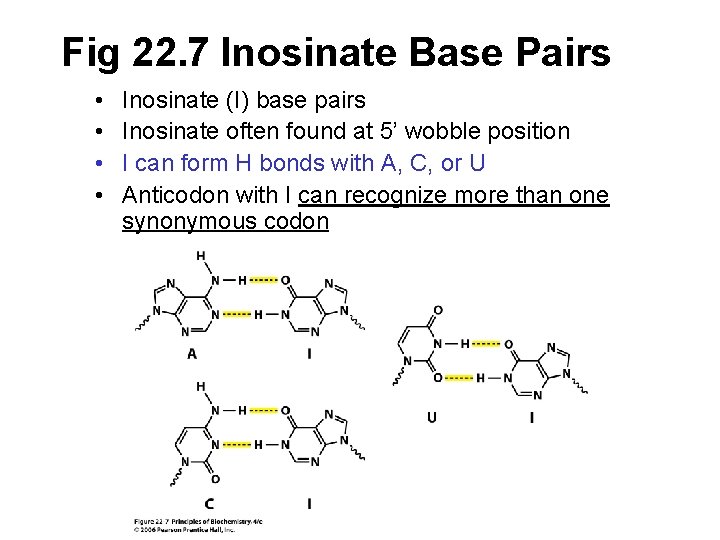
Fig 22. 7 Inosinate Base Pairs • • Inosinate (I) base pairs Inosinate often found at 5’ wobble position I can form H bonds with A, C, or U Anticodon with I can recognize more than one synonymous codon

Codon-Anticodon Recognition • Wobble allows some t. RNA molecules to recognize more than one codon • Isoacceptor t. RNA molecules - different t. RNA molecules that bind the same amino acids • Isoacceptor t. RNAs identified by Roman numerals or codons: t. RNAIAla, t. RNAIIAla or t. RNAGCGAla • Bacteria have 30 -60 different t. RNAs, eukaryotes have up to 80 different t. RNAs
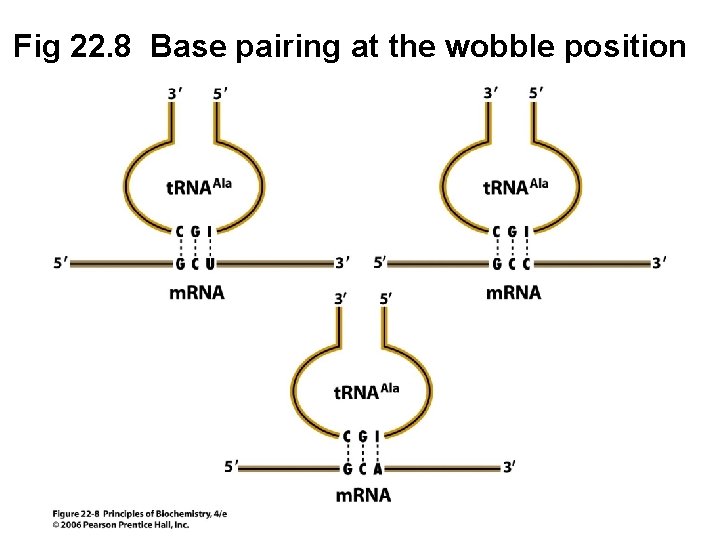
Fig 22. 8 Base pairing at the wobble position

22. 3 Aminoacyl-t. RNA Synthetases • Aminoacyl-t. RNA - amino acids are covalently attached to the 3’ end of each t. RNA molecule (named as: alanyl-t. RNAAla) • Aminoacyl-t. RNA synthetases catalyze reactions • Most species have at least 20 different aminoacyl-t. RNA synthetases (1 per amino acid) • Each synthetase specific for a particular amino acid, but may recognize isoacceptor t. RNAs

A. The Aminoacyl-t. RNA Synthetase Reaction • Aminoacyl-t. RNAs are high-energy molecules (the amino acid has been “activated”) • The activation of an amino acid by aminoacylt. RNA synthetase requires ATP Amino acid + t. RNA + ATP Aminoacyl-t. RNA + AMP + PPi
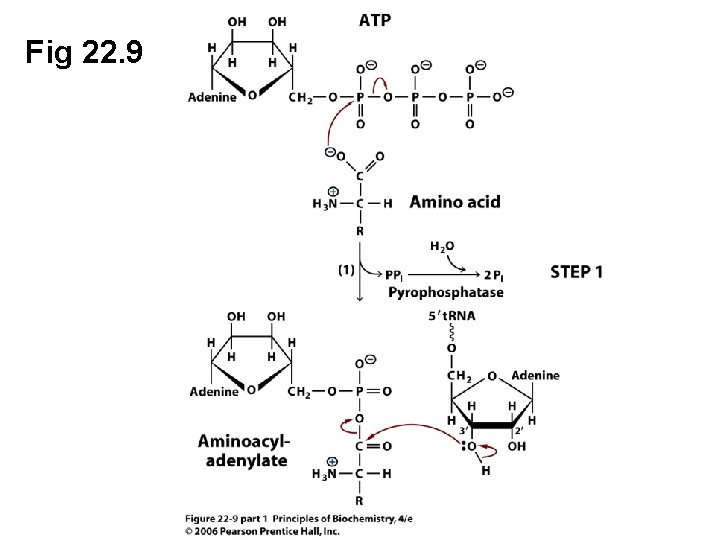
Fig 22. 9
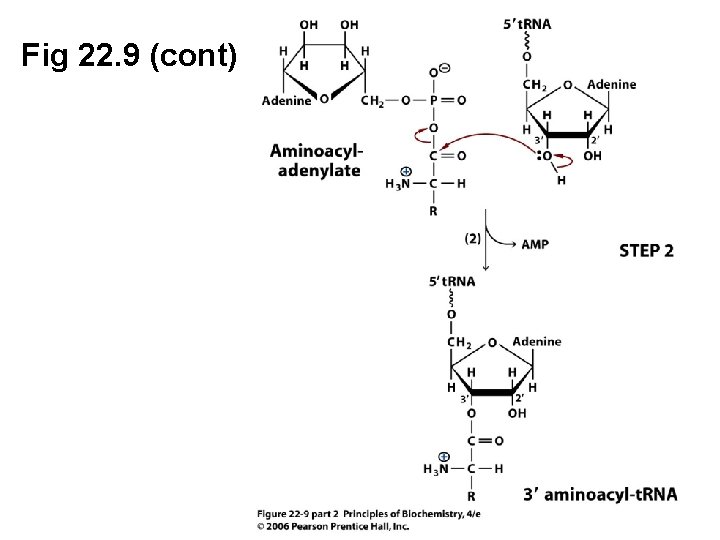
Fig 22. 9 (cont)

B. Specificity of Aminoacyl-t. RNA Synthetase • Attachment of the correct amino acid to the corresponding t. RNA is a critical step • Synthetase binds ATP and the correct amino acid (based on size, charge, hydrophobicity) • Synthetase then selectively binds specific t. RNA molecule based on structural features • Synthetase may recognize the anticodon as well as the acceptor stem
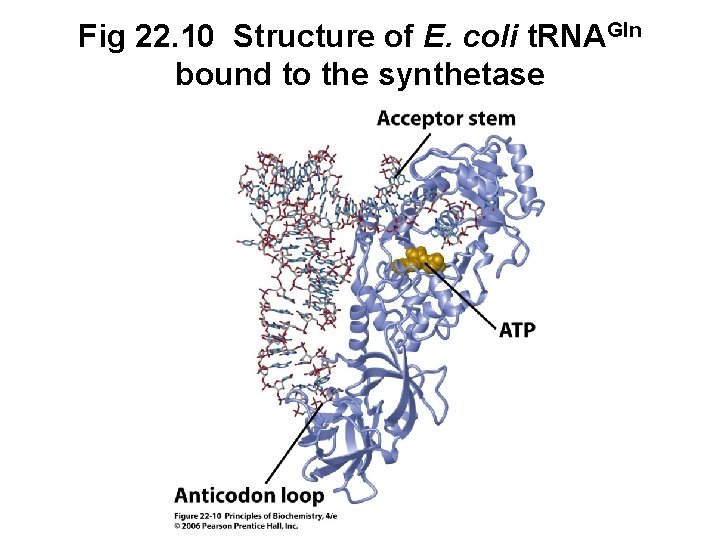
Fig 22. 10 Structure of E. coli t. RNAGln bound to the synthetase

C. Proofreading Activity of Aminoacylt. RNA Synthetases • Some aa-t. RNA synthetases can proofread • Isoleucyl-t. RNA synthetase may bind valine instead of isoleucine and incorporate it into valyl-adenylate • The valyl-adenylate is usually then hydrolyzed to valine and AMP so that valyl-t. RNAIle does not form
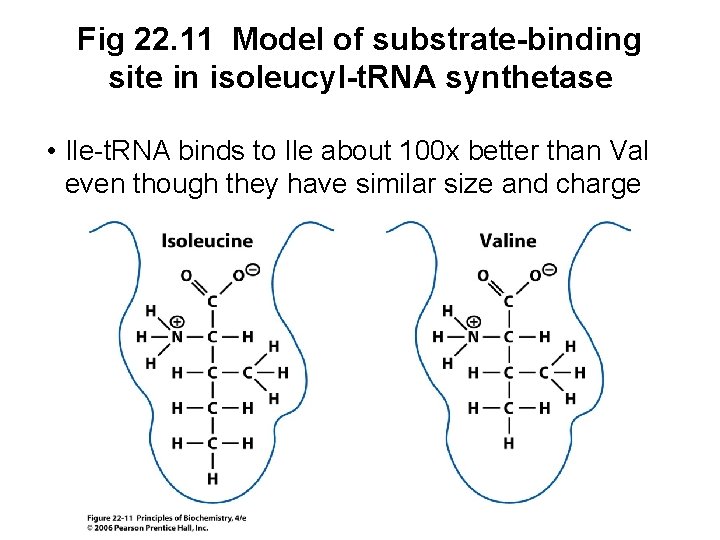
Fig 22. 11 Model of substrate-binding site in isoleucyl-t. RNA synthetase • Ile-t. RNA binds to Ile about 100 x better than Val even though they have similar size and charge

22. 4 Ribosomes • Protein synthesis is carried out by a complex composed of the ribosome, accessory protein factors, m. RNA and charged t. RNA molecules • Initiation complex assembles at first m. RNA codon, and disassembles at termination step • Ribosome complex moves 5’ m. RNA • Polypeptide is synthesized in N 3’ along template C direction

A. Ribosomes Are Composed of Both r. RNA and Protein • All ribosomes contain two subunits of unequal size • E. coli: 70 S composed of a 30 S and a 50 S • Eukaryotes: 80 S composed of a 40 S and a 60 S
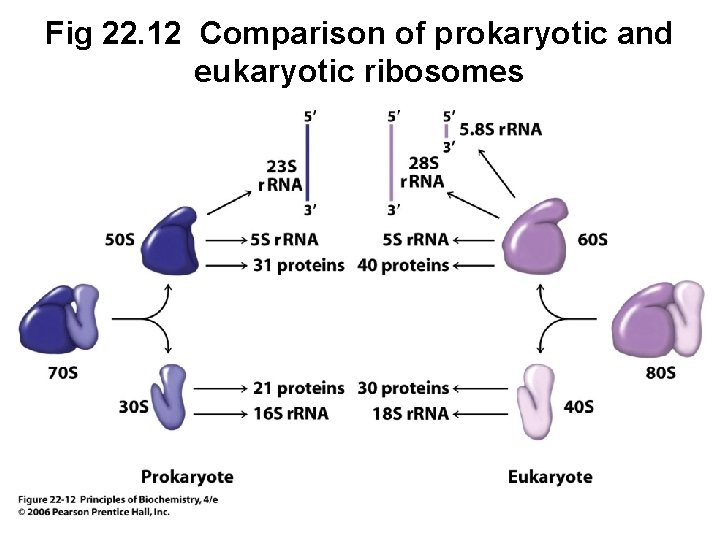
Fig 22. 12 Comparison of prokaryotic and eukaryotic ribosomes
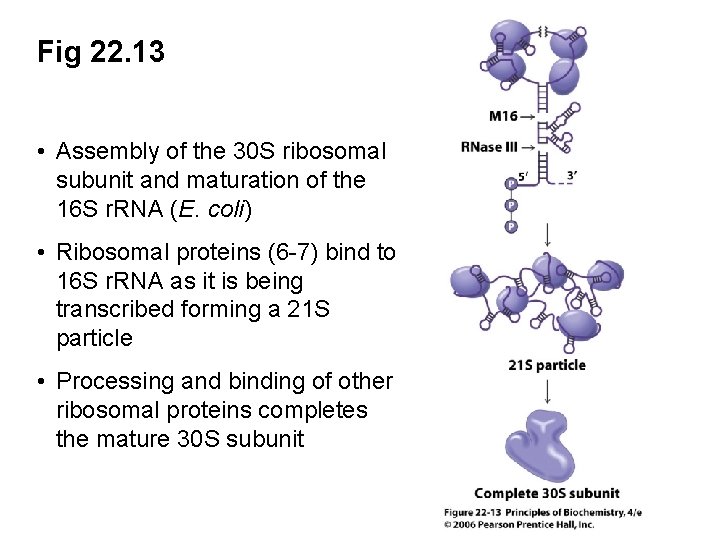
Fig 22. 13 • Assembly of the 30 S ribosomal subunit and maturation of the 16 S r. RNA (E. coli) • Ribosomal proteins (6 -7) bind to 16 S r. RNA as it is being transcribed forming a 21 S particle • Processing and binding of other ribosomal proteins completes the mature 30 S subunit
Fig 22. 14 Structure of the 30 S ribosomal subunit (T. thermophilus)

B. Ribosomes Contain Two Aminoacyl -t. RNA Binding Sites • Ribosome must align two charged t. RNA molecules so that anticodons interact with correct codons of m. RNA • Aminoacylated ends of the t. RNAs are positioned at the site of peptide bond formation • Ribosome must hold both m. RNA and growing polypeptide chain
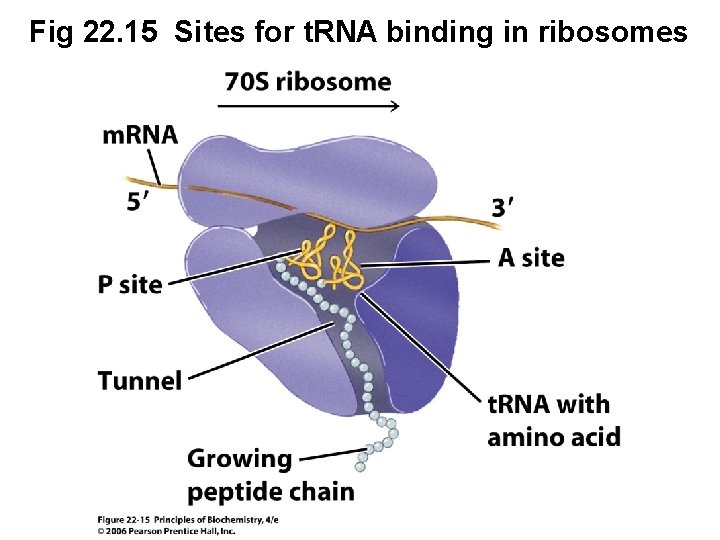
Fig 22. 15 Sites for t. RNA binding in ribosomes

22. 5 Initiation of Translation • The translation complex is assembled at the beginning of the m. RNA coding sequence • Complex consists of: Ribosomal subunits m. RNA template to be translated Initiator t. RNA molecule Protein initiation factors
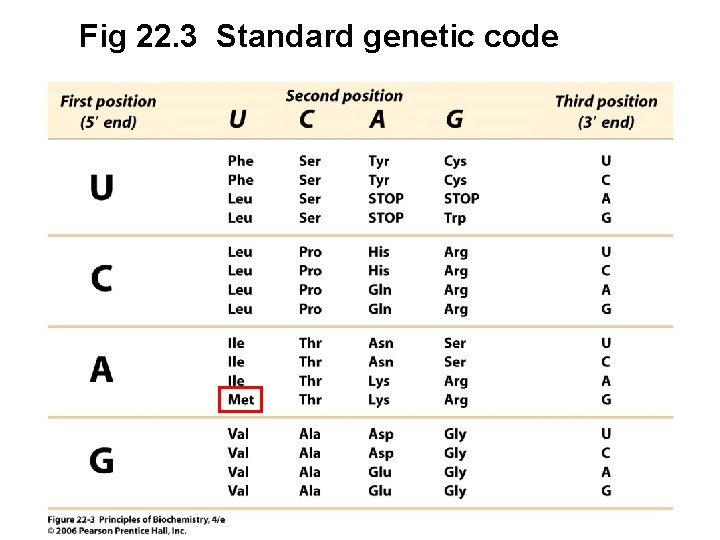
Fig 22. 3 Standard genetic code

A. Initiator t. RNA • First codon translated is usually AUG • Each cell contains at least two methionyl-t. RNAMet molecules which recognize AUG • The initiator t. RNA recognizes initiation codons • Second t. RNAMet recognizes only internal AUG • Bacteria: N-formylmethionyl-t. RNAf. Met • Eukaryotes: methionyl-t. RNAi. Met
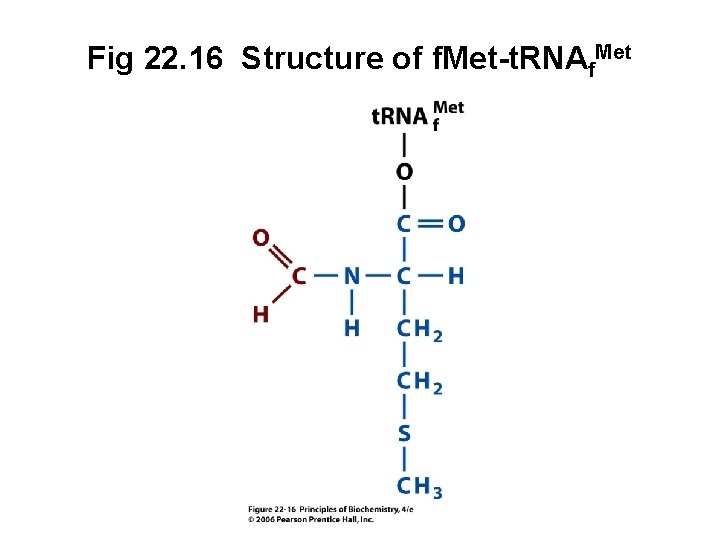
Fig 22. 16 Structure of f. Met-t. RNAf. Met

B. Initiation Complexes Assemble Only at Initiation Codons • Ribosome must recognize protein synthesis start • In prokaryotes, the 30 S ribosome binds to a region of the m. RNA (Shine-Dalgarno sequence) upstream of the initiation sequence • S-D sequence also binds to a complementary base sequence at the 3’ end of the 16 S r. RNA • Double-stranded RNA structure binds m. RNA to the ribosome
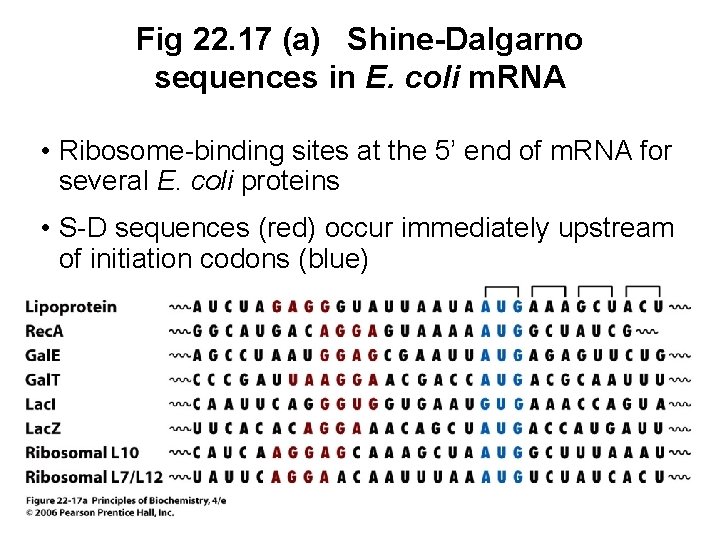
Fig 22. 17 (a) Shine-Dalgarno sequences in E. coli m. RNA • Ribosome-binding sites at the 5’ end of m. RNA for several E. coli proteins • S-D sequences (red) occur immediately upstream of initiation codons (blue)
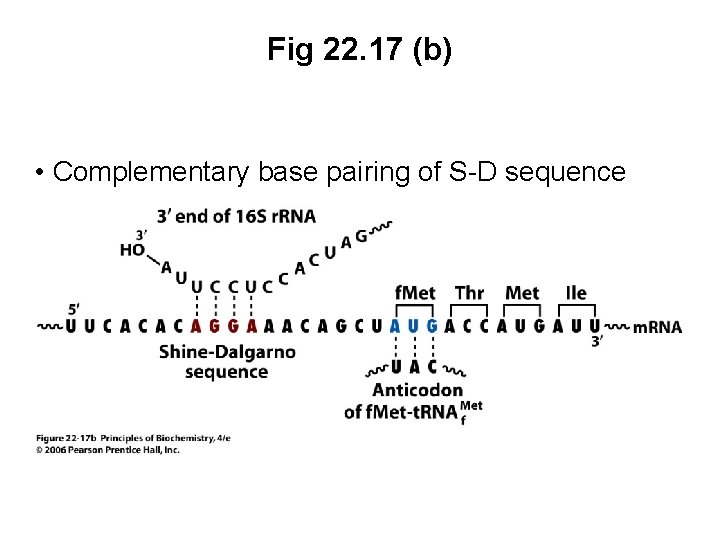
Fig 22. 17 (b) • Complementary base pairing of S-D sequence

C. Initiation Factors Help Form Initiation Complex • Initiation factors are required to form a complex • Prokaryote factors: IF-1, IF-2, IF-3 • Eukaryote factors: e. IFs (8 or more factors)
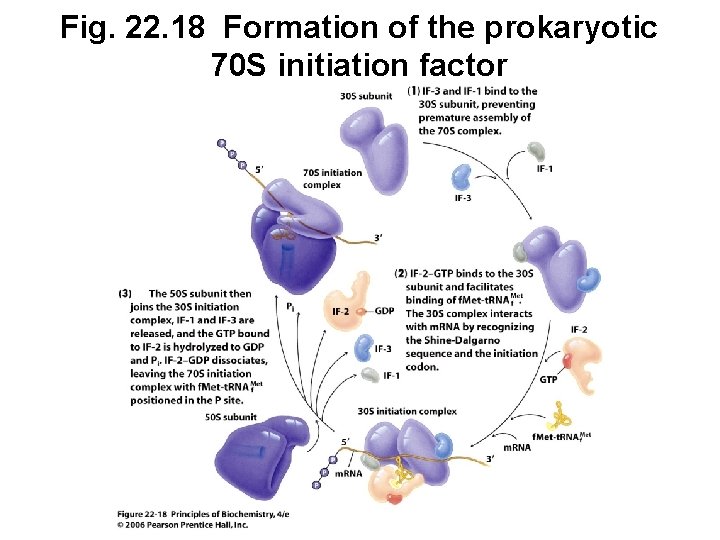
Fig. 22. 18 Formation of the prokaryotic 70 S initiation factor
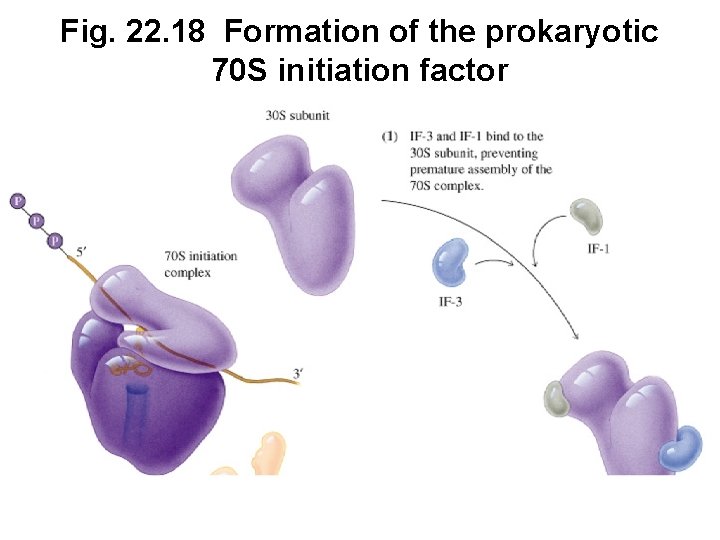
Fig. 22. 18 Formation of the prokaryotic 70 S initiation factor
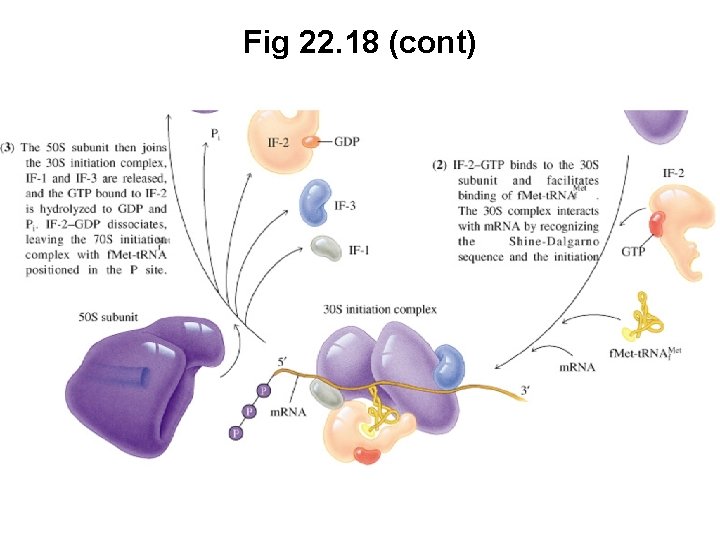
Fig 22. 18 (cont)

D. Translation Initiation in Eukaryotes • Eukaryotic initiation factor 4 (e. IF-4), (or cap binding protein, CBP) binds to the (5’ end) 7 methylguanylate cap of eukaryotic m. RNA • A preinitiation complex forms (40 S ribosome, aminoacylated initiator t. RNA, other factors) and searches the m. RNA 5’ 3’ for an initiator codon • The Met-t. RNAi. Met binds to AUG, and the 60 S ribosomal subunit binds to complete the complex

22. 6 Chain Elongation is a Three-Step Microcycle • The initiator t. RNA is in the P site • Site A is ready to receive an aminoacyl-t. RNA • Elongation is a three-step cycle: (1) Positioning the correct aa-t. RNA in site A (2) Formation of a peptide bond (3) Shifting m. RNA by one codon
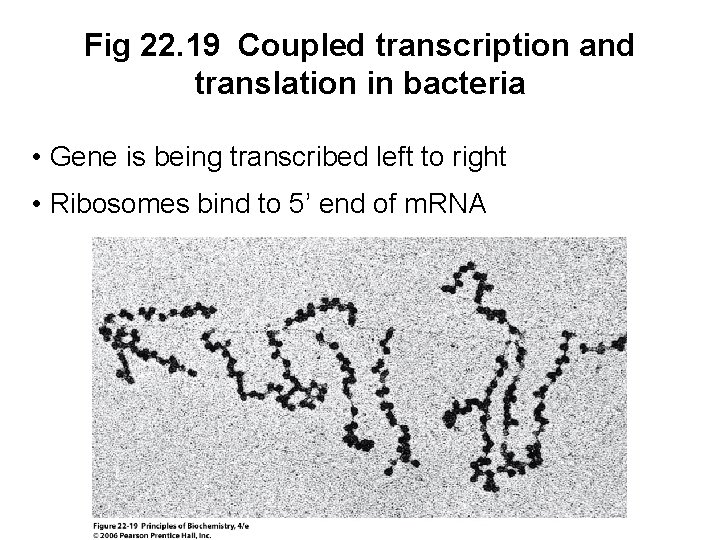
Fig 22. 19 Coupled transcription and translation in bacteria • Gene is being transcribed left to right • Ribosomes bind to 5’ end of m. RNA

A. Elongation Factors Dock an Aminoacyl-t. RNA in the A Site • Bacterial elongation factor EF-Tu helps the correct aa-t. RNA insert into site A • An EF-Tu-GTP complex binds to all aa-t. RNA molecules except f. Met-t. RNAf. Met (initiator) • A ternary complex of EF-Tu-GTP-aa-t. RNA binds in the ribosomal A site • If the anticodon of the aa-t. RNA correctly base pairs with the m. RNA codon, complex is stabilized
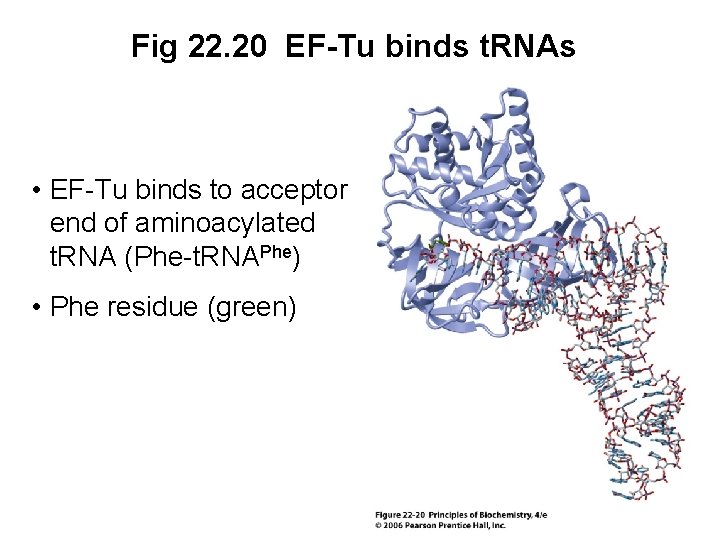
Fig 22. 20 EF-Tu binds t. RNAs • EF-Tu binds to acceptor end of aminoacylated t. RNA (Phe-t. RNAPhe) • Phe residue (green)
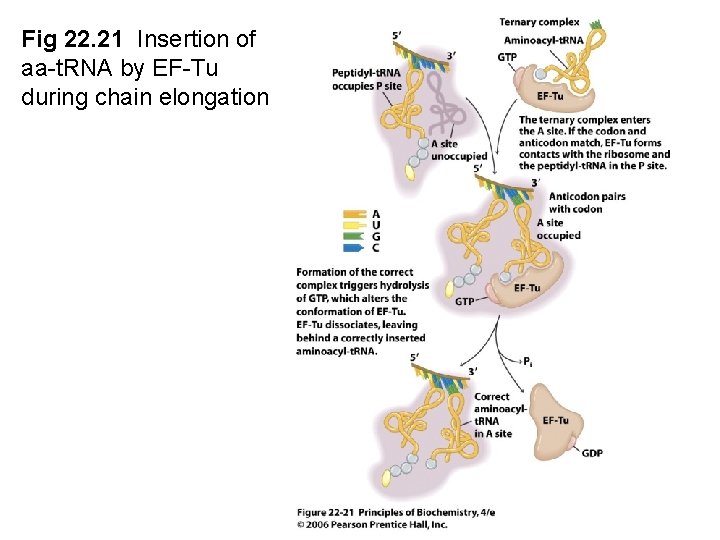
Fig 22. 21 Insertion of aa-t. RNA by EF-Tu during chain elongation
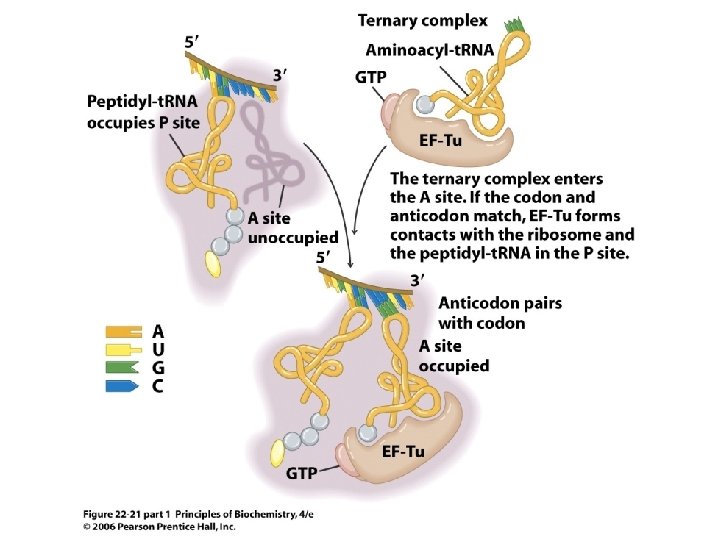
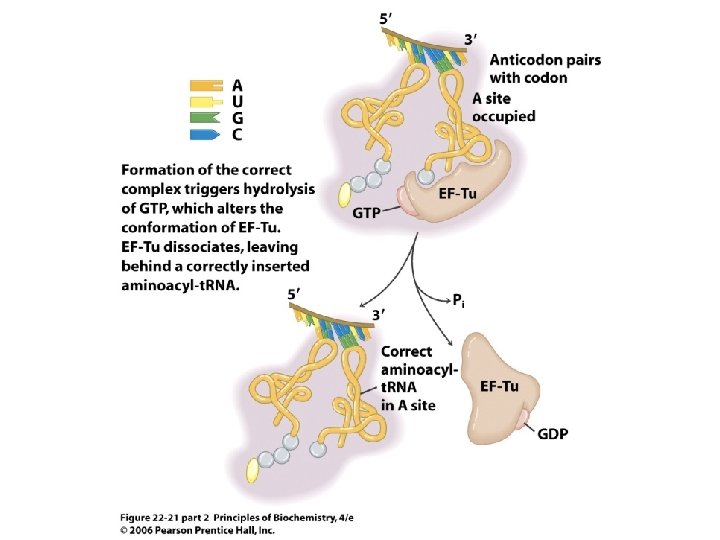
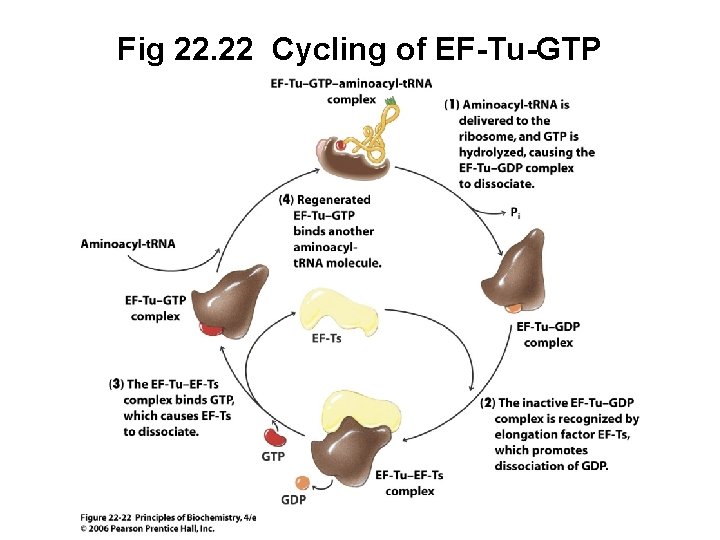
Fig 22. 22 Cycling of EF-Tu-GTP
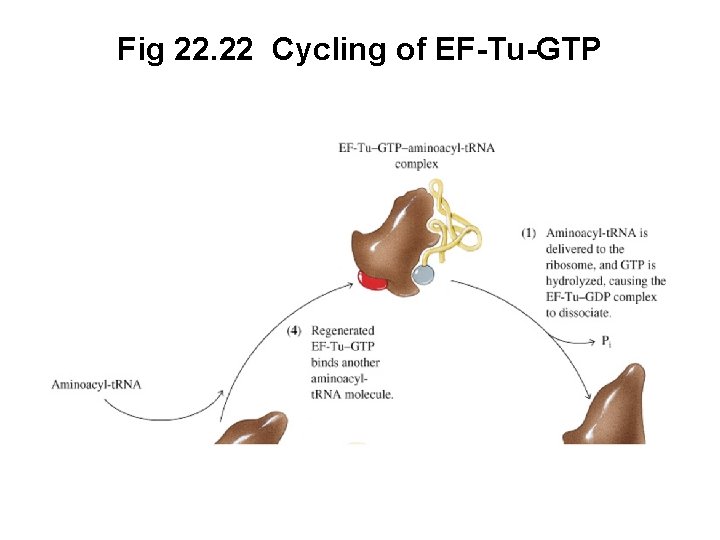
Fig 22. 22 Cycling of EF-Tu-GTP
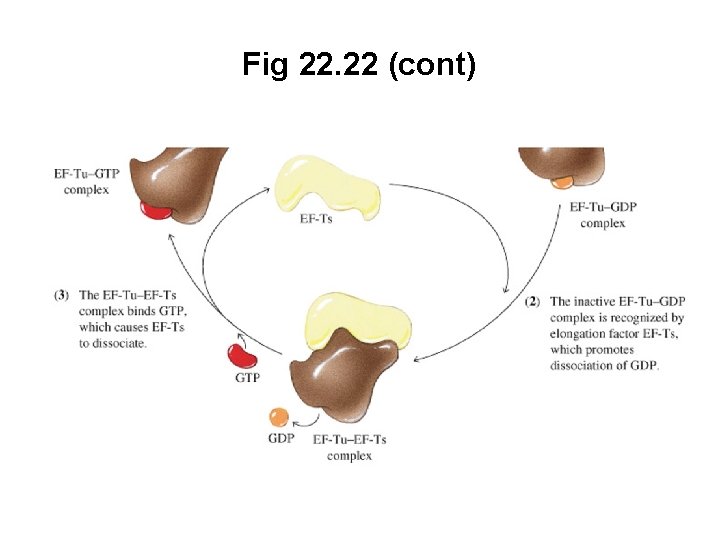
Fig 22. 22 (cont)

B. Peptidyl Transferase Catalyzes Peptide Bond Formation • Peptidyl transferase activity is contained within the large ribosomal subunit • Substrate binding site in 23 S r. RNA and 50 S ribosomal proteins • Catalytic activity from 23 S r. RNA (an RNA-catalyzed reaction)
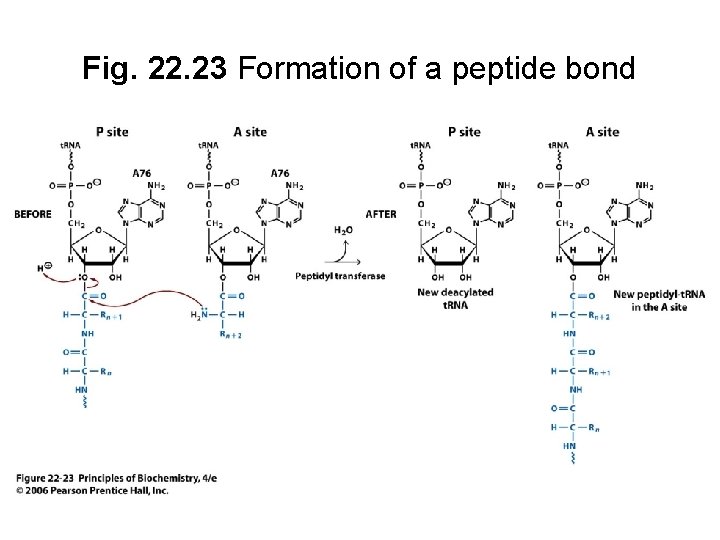
Fig. 22. 23 Formation of a peptide bond

C. Translocation Moves the Ribosome by One Codon • Translocation step: the new peptidyl-t. RNA is moved from the A site to the P site, while the m. RNA shifts by one codon • The deaminoacylated t. RNA has shifted from the P site to the E site (exit site) • Binding of EF-G-GTP to the ribosome completes translocation of peptidyl-t. RNA
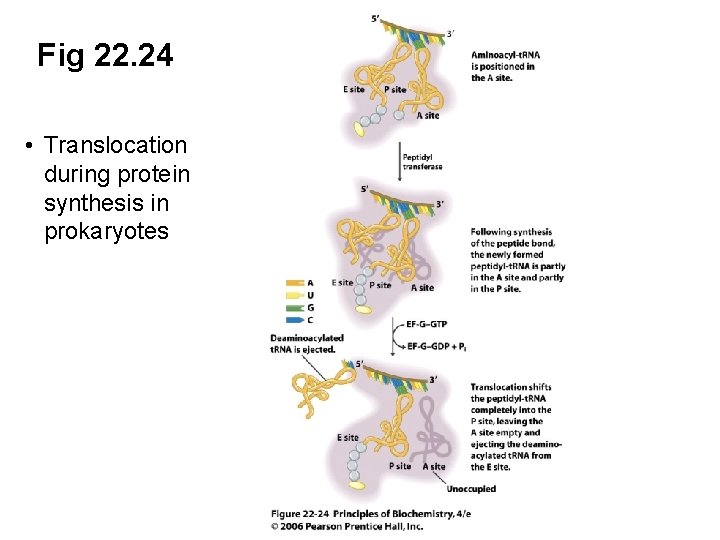
Fig 22. 24 • Translocation during protein synthesis in prokaryotes
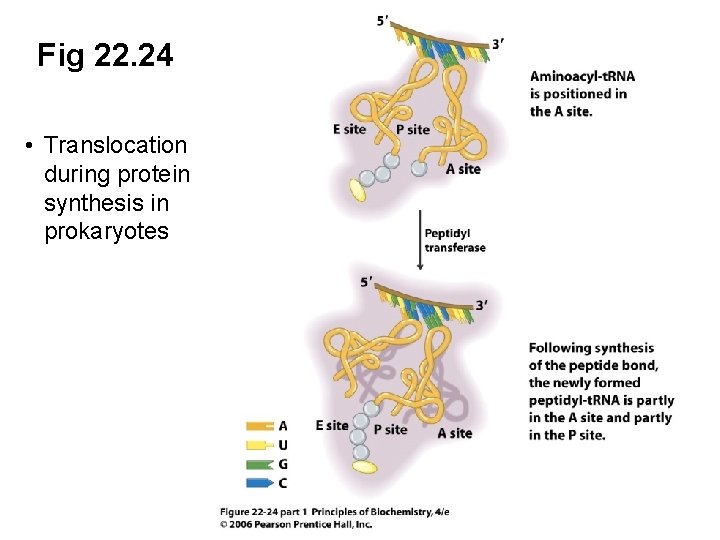
Fig 22. 24 • Translocation during protein synthesis in prokaryotes
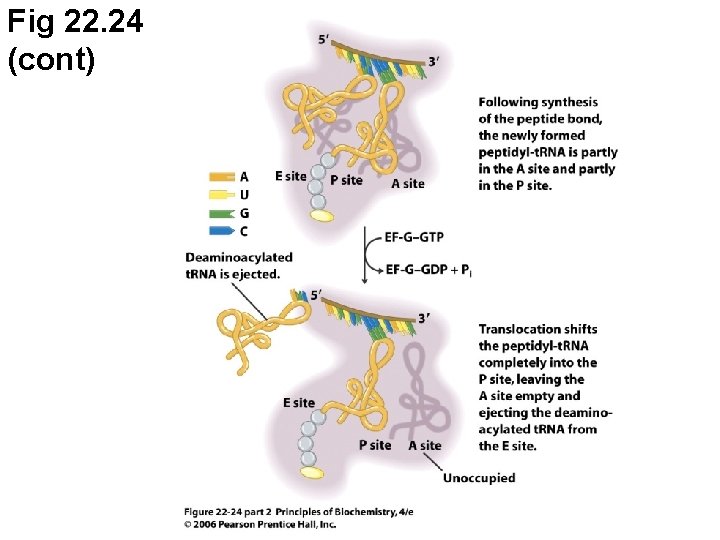
Fig 22. 24 (cont)

Formation of the Peptide Chain • Growing peptide chain extends from the peptidyl-t. RNA (P site) through a tunnel in the 50 S subunit • Newly synthesized polypeptide does not begin to fold until it emerges from the tunnel • Elongation in eukaryotes is similar to E. coli: EF-1 a - docks the aa-t. RNA into A site EF-1 b - recycles EF-1 a EF-2 - carries out translocation

Animations • t. RNA binding ribosomes

22. 7 Termination of Translation • E. coli release factors: RF-1, RF-2, RF-3 • Translocation positions one of three termination codons in A site: UGA, UAG, UAA • No t. RNA molecules recognize these codons and protein synthesis stalls • One of the release factors binds and causes hydrolysis of the peptidyl-t. RNA to release the polypeptide chain

Animations • Protein synthesis

22. 8 Protein Synthesis is Energetically Expensive • Four phosphoanhydride bonds are cleaved for each amino acid added to a polypeptide chain Amino acid activation: Two ~P bonds ATP AMP + 2 Pi Chain elongation: Two ~P bonds 2 GTP 2 GDP + 2 Pi

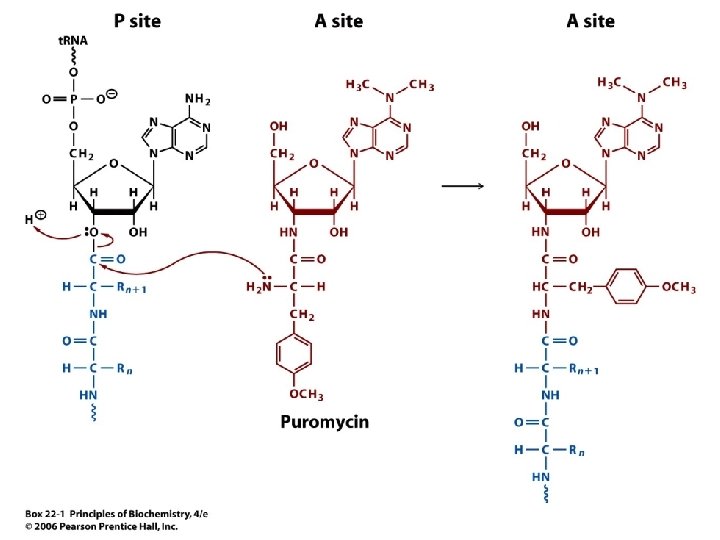
Box 22. 1 Some Antibiotics Inhibit Protein Synthesis • Some antibiotics prevent bacterial growth by inhibiting the formation of peptide bonds • Puromycin (next slide) resembles the 3’ end of an aminoacyl-t. RNA, and can enter the A site of a ribosome • The peptidyl-puromycin formed is bound weakly in the A site and dissociates terminating protein synthesis

22. 9 Regulation of Protein Synthesis A. Ribosomal Protein Synthesis Is Coupled to Ribosome Assembly in E. coli – Synthesis of ribosomal proteins is tightly regulated at the level of translation – Ribosomal protein genes encode one ribosomal protein that inhibits translation of its own polycistrionic m. RNA by binding near the initiation codon of the m. RNA
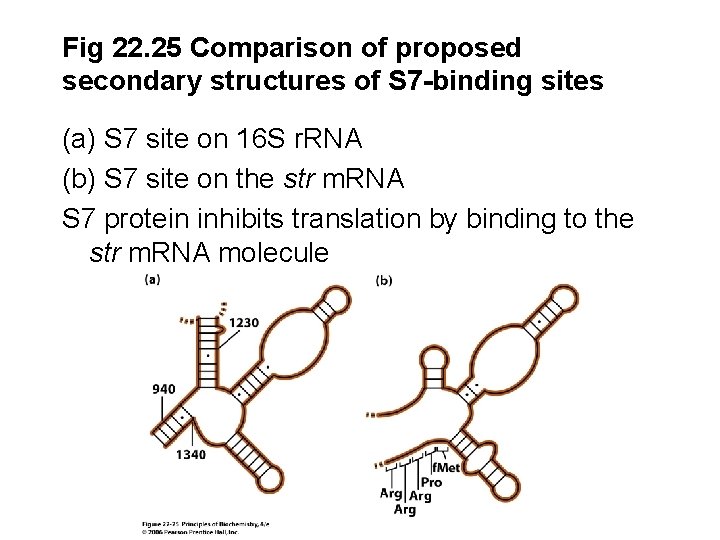
Fig 22. 25 Comparison of proposed secondary structures of S 7 -binding sites (a) S 7 site on 16 S r. RNA (b) S 7 site on the str m. RNA S 7 protein inhibits translation by binding to the str m. RNA molecule

B. Globin Synthesis Depends on Heme Availability • Hemoglobin synthesis requires globin chains and heme in stoichiometric amounts • Globin synthesis is controlled by regulation of translation initiation • Heme-controlled inhibitor (HCI) phosphorylates factor e. IF-2 which then cannot participate in translation initiation • High heme levels interfere with HCI so that globin synthesis proceeds
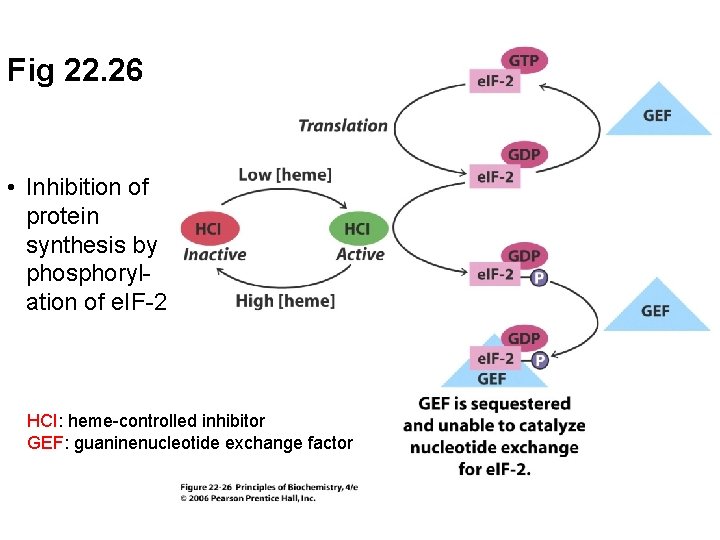
Fig 22. 26 • Inhibition of protein synthesis by phosphorylation of e. IF-2 HCI: heme-controlled inhibitor GEF: guaninenucleotide exchange factor

C. The E. coli trp Operon Is Regulated by Repression and Attenuation • The trp operon in E. coli encodes the proteins necessary for tryptophan biosynthesis • Because tryptophan is a negative regulator of its own biosynthesis, synthesis can be repressed when exogenous Trp is available • Tryptophan is a corepressor of the trp. O operator (Figure 22. 27, next slide)
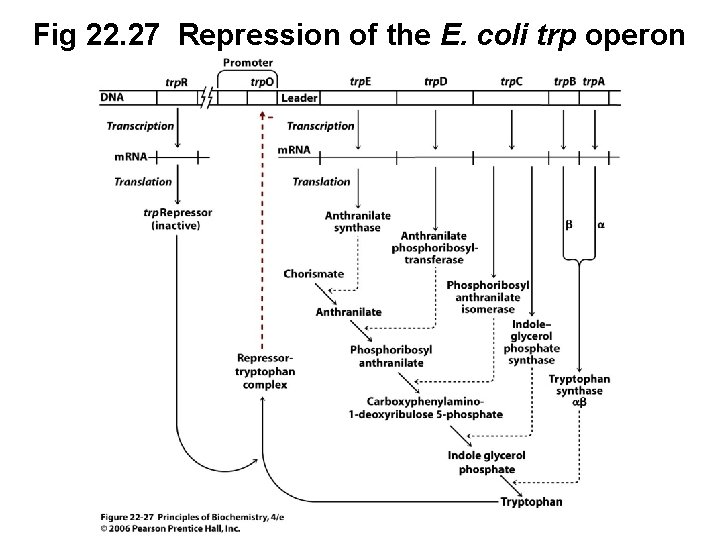
Fig 22. 27 Repression of the E. coli trp operon (continued next slide)
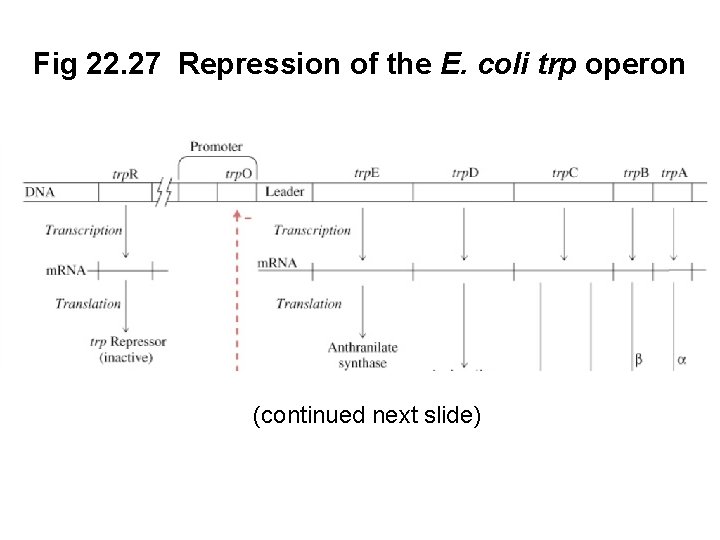
Fig 22. 27 Repression of the E. coli trp operon (continued next slide)
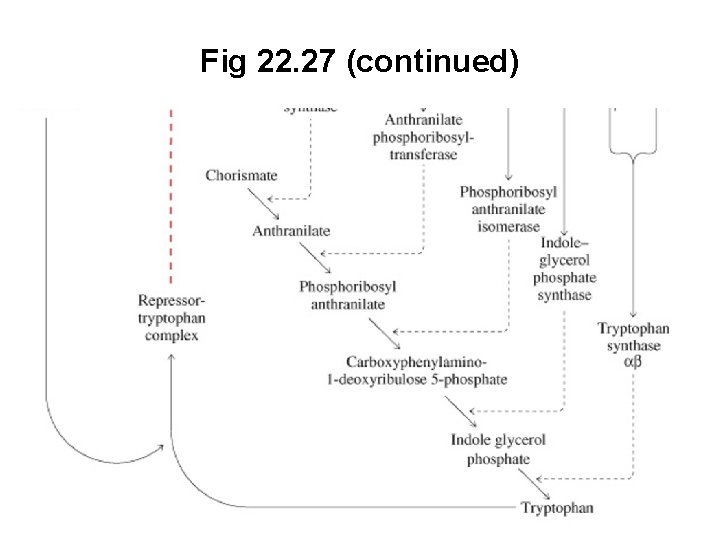
Fig 22. 27 (continued)

Attenuation in E. coli • A second mechanism for regulation of the E. coli trp operon depends on translation • Determines whether transcription of the operon proceeds or terminates prematurely • GC-rich regions in the m. RNA trp leader region can base pair to form two alternative hairpin structures which affect transcription
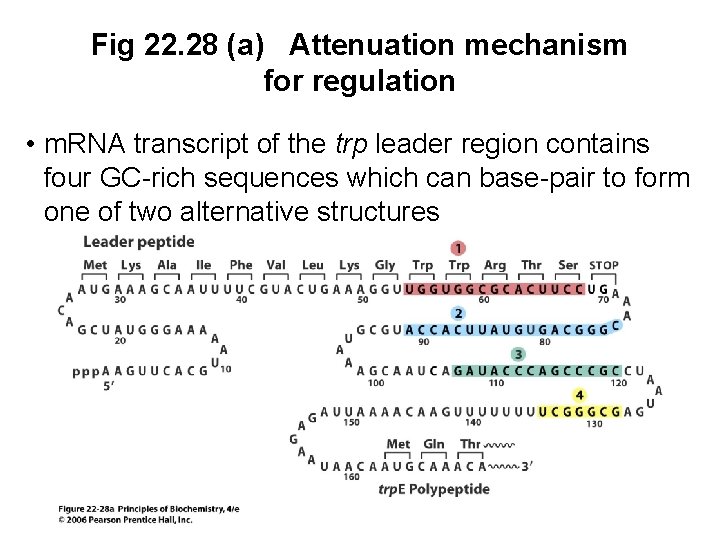
Fig 22. 28 (a) Attenuation mechanism for regulation • m. RNA transcript of the trp leader region contains four GC-rich sequences which can base-pair to form one of two alternative structures
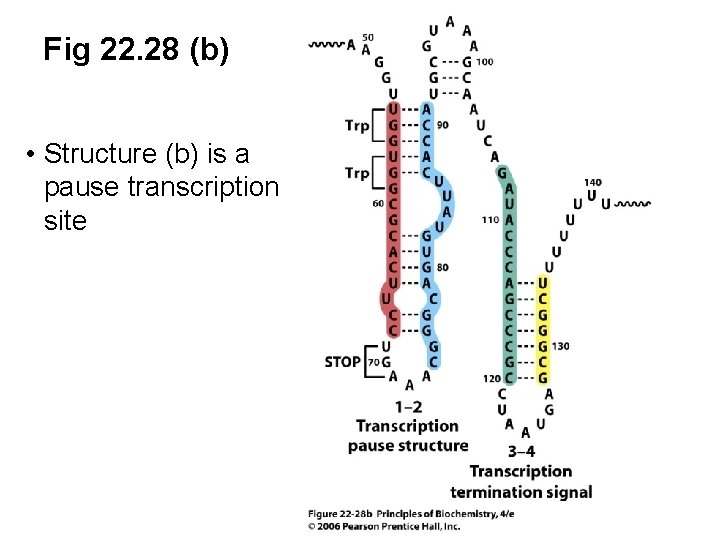
Fig 22. 28 (b) • Structure (b) is a pause transcription site
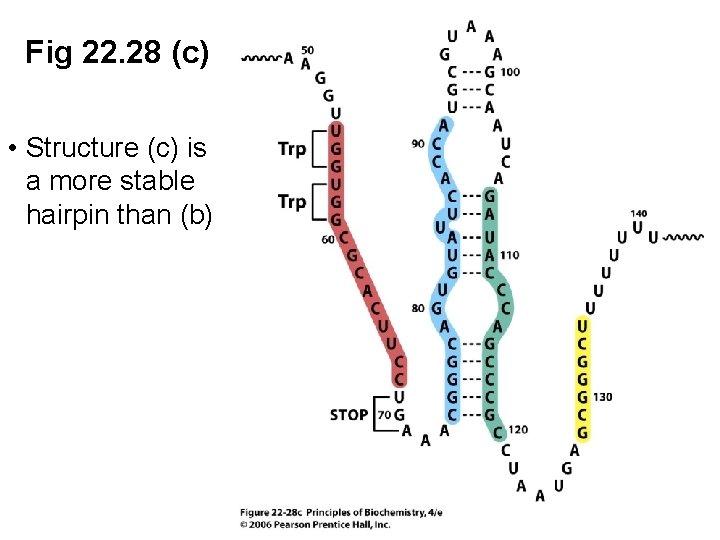
Fig 22. 28 (c) • Structure (c) is a more stable hairpin than (b)
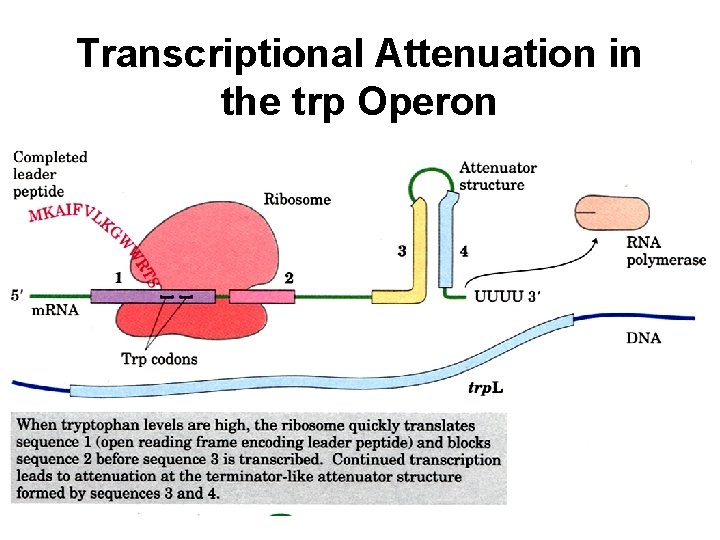
Transcriptional Attenuation in the trp Operon
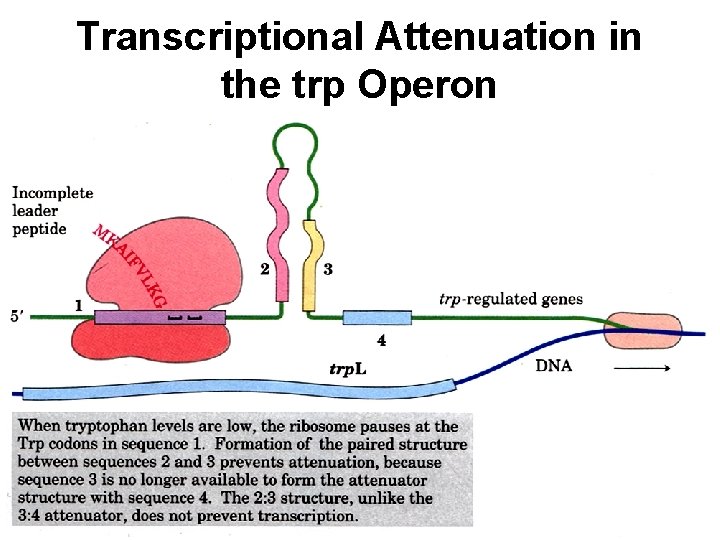
Transcriptional Attenuation in the trp Operon

22. 10 Posttranslational Processing • Posttranslational modifications can occur either before the polypeptide chain is complete (cotranslational) or after (posttranslational) ─ deformylation of N-terminal residue (prok) ─ removal of N-terminal methionine residue ─ formation of disulfide bonds ─ cleavage by proteinases ─ phosphorylation or acetylation
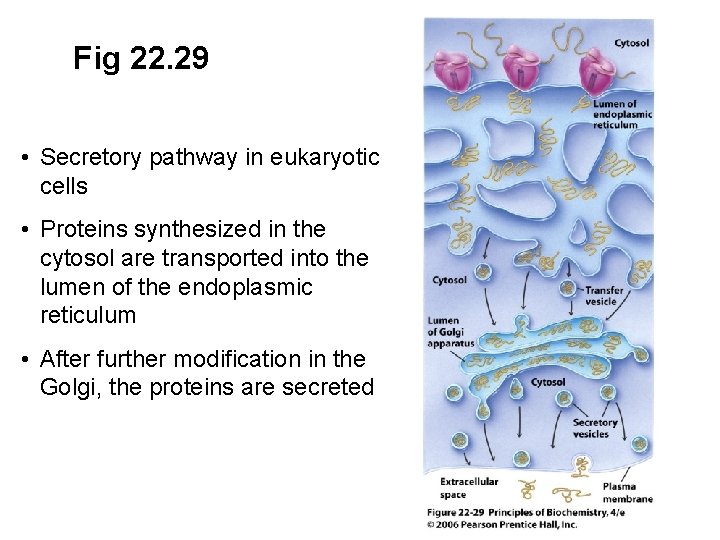
Fig 22. 29 • Secretory pathway in eukaryotic cells • Proteins synthesized in the cytosol are transported into the lumen of the endoplasmic reticulum • After further modification in the Golgi, the proteins are secreted
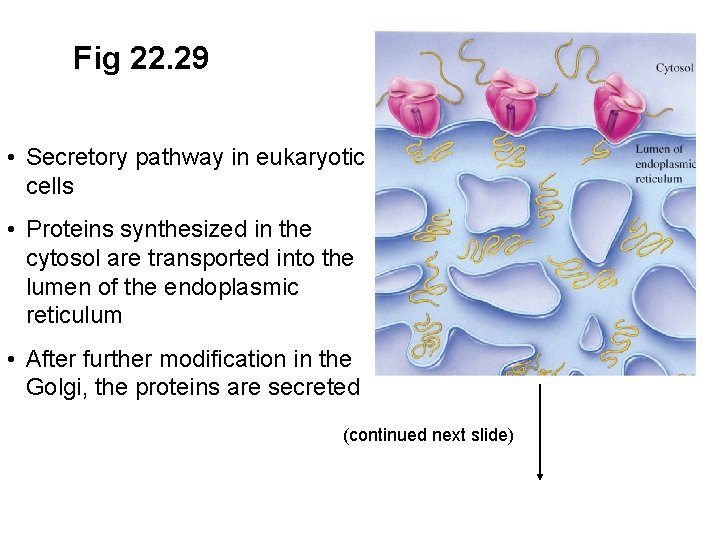
Fig 22. 29 • Secretory pathway in eukaryotic cells • Proteins synthesized in the cytosol are transported into the lumen of the endoplasmic reticulum • After further modification in the Golgi, the proteins are secreted (continued next slide)
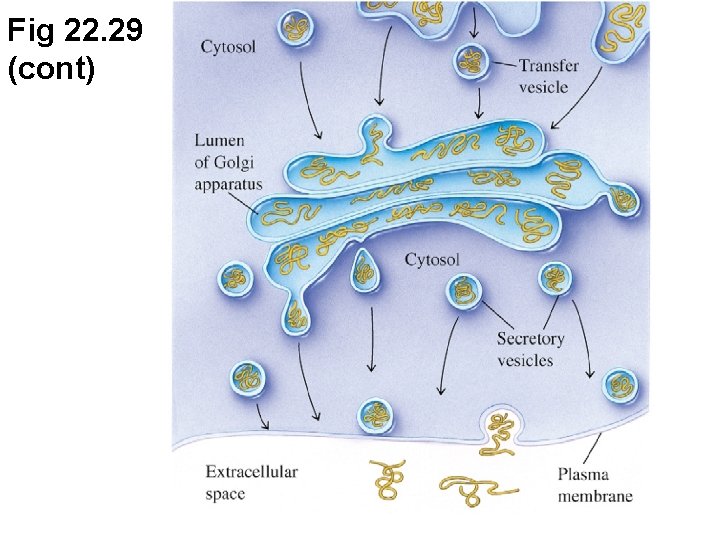
Fig 22. 29 (cont)

Fig 22. 30 Secretory Vesicles in a maize rootcap cell

A. The Signal Hypothesis • Secreted proteins are synthesized by ribosomes on the surface of the endoplasmic reticulum • A signal peptide is present on the N-terminus that signals the protein to cross a membrane • Signal peptides are 16 -30 residues long, and include 4 -15 hydrophobic residues

Fig 22. 31 Signal peptides from secreted proteins • Hydrophobic residues in blue, arrows mark sites where signal peptide is cleaved from the precursor
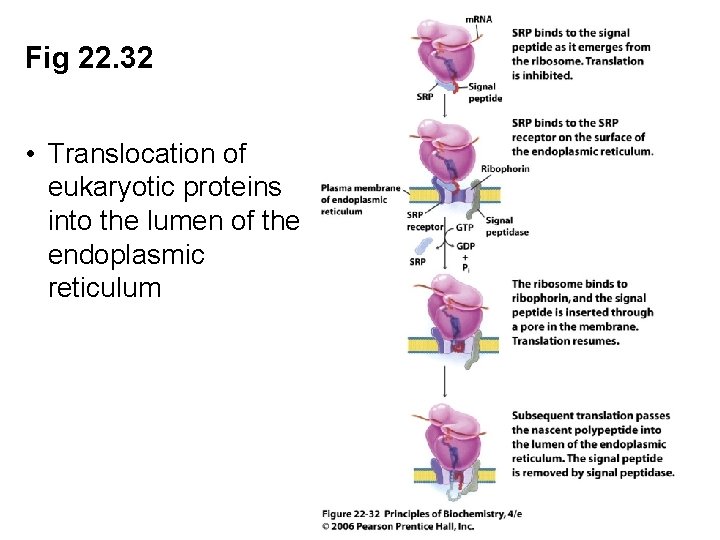
Fig 22. 32 • Translocation of eukaryotic proteins into the lumen of the endoplasmic reticulum
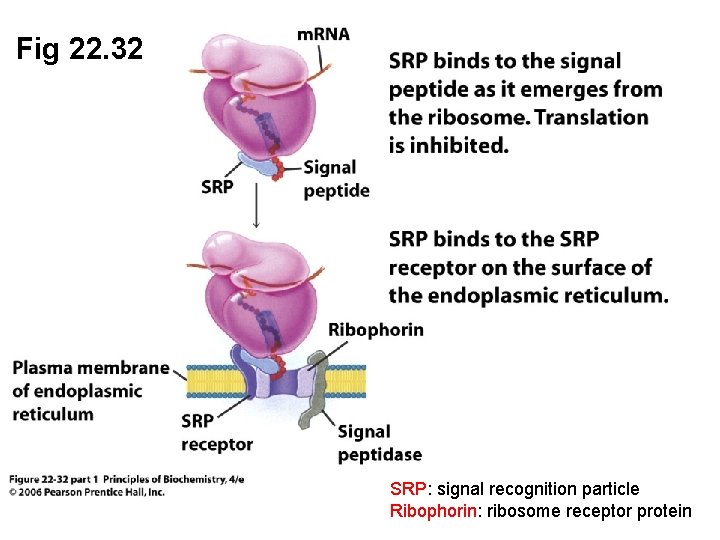
Fig 22. 32 SRP: signal recognition particle Ribophorin: ribosome receptor protein
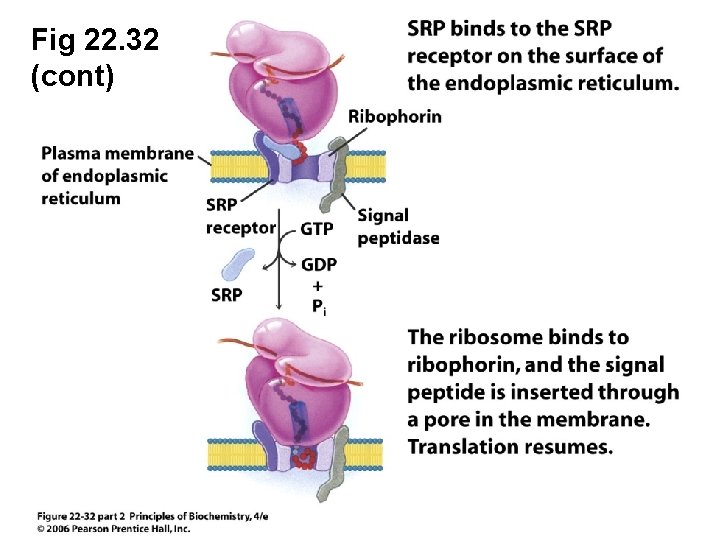
Fig 22. 32 (cont)
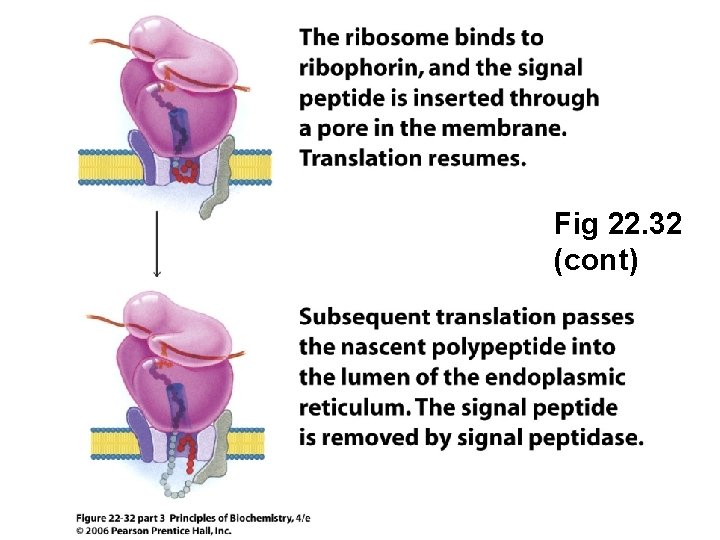
Fig 22. 32 (cont)

B. Glycosylation of proteins • Many integral membrane and secretory proteins contain covalently bound oligosaccharide chains • Carbohydrate may be from 1% to 80% of the mass of the glycoprotein • A common glycosylation reaction is the covalent attachment of a complex oligosaccharide to the side chain of an asparagine residue
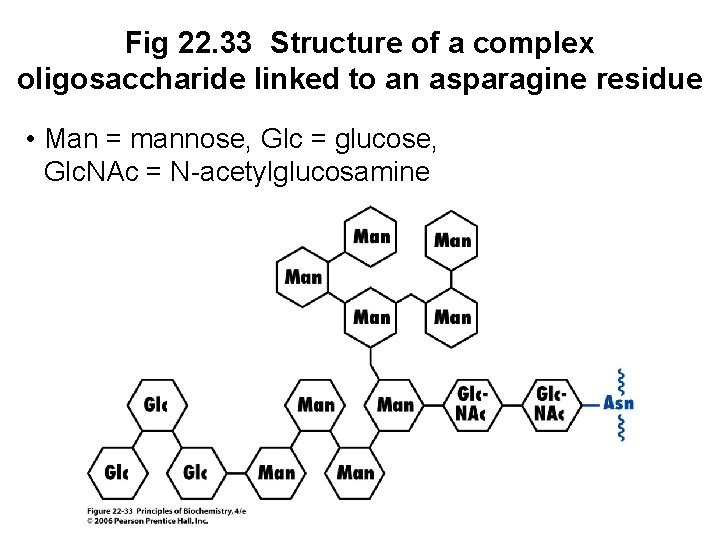
Fig 22. 33 Structure of a complex oligosaccharide linked to an asparagine residue • Man = mannose, Glc = glucose, Glc. NAc = N-acetylglucosamine
- Slides: 98