Horizontal transfer and pathogenicity in bacteria Jlia Vctor

Horizontal transfer and pathogenicity in bacteria Júlia Víctor Baldomà Genomics December 2018

Table of contents • Introduction • Mechanisms of transfer of DNA • MGE involved in horizontal gene transfer of virulence determinants • MGEs and horizontal gene transfer in the virulence of patogens: • E. coli • Salmonella • Streptococcus • Clostridium • Conclusions • Bibliography

• Introduction • Mechanisms of transfer of DNA • MGE involved in horizontal gene transfer of virulence determinants • MGEs and horizontal gene transfer in the virulence of patogens: • E. coli • Salmonella • Streptococcus • Clostridium • Conclusions • Bibliography
Introduction Bacterial genome frequently contain a significant amount of foreign DNA Originated in another organism Horizontal gene transfer (HGT) Mobile genetic elements (MGE) Impact in bacterial genomes: PATOGENICITY Although they have several mechanisms for resisting it they have a major role in bacterial evolution

• Introduction • Mechanisms of transfer of DNA • MGE involved in horizontal gene transfer of virulence determinants • MGEs and horizontal gene transfer in the virulence of patogens: • E. coli • Salmonella • Streptococcus • Clostridium • Conclusions • Bibliography
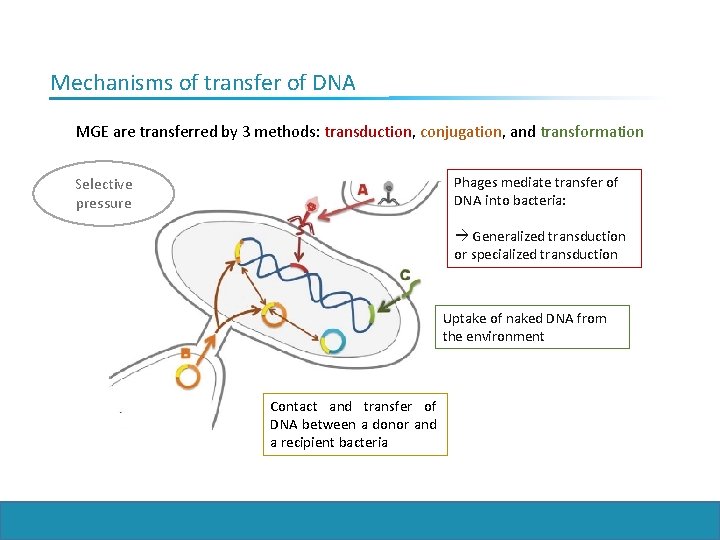
Mechanisms of transfer of DNA MGE are transferred by 3 methods: transduction, conjugation, and transformation Phages mediate transfer of DNA into bacteria: Selective pressure Generalized transduction or specialized transduction Uptake of naked DNA from the environment Contact and transfer of DNA between a donor and a recipient bacteria
• Introduction • Mechanisms of transfer of DNA • MGE involved in horizontal gene transfer of virulence determinants • MGEs and horizontal gene transfer in the virulence of patogens: • E. coli • Salmonella • Streptococcus • Clostridium • Conclusions • Bibliography

MGE involved in HGT of virulence determinants IS and transposons: IS: small segment of DNA, genes that allow to insert at numerous sites of DNA. Transposase enzyme + inverted repeat sequences. Preference for A-T rich area. Transposons: in addition to transposition function carry other genes Plasmids: Extrachromosomal DNA molecules Can carry or not genes that permit their transfer Not part of core genome Encode range of genes for a variety of nonessential functions Bacteriophages: Contain bacterial toxins that are incorporated into the bacterial chromosome Pathogenicity islands: Large segments of foreign DNA inserted into bacterial chromosome Flanked by direct repeats, usually contain mobility genes % of gens encoding virulence factors was higher than outside PAIs
• Introduction • Mechanisms of transfer of DNA • MGE involved in horizontal gene transfer of virulence determinants • MGEs and horizontal gene transfer in the virulence of patogens: • E. coli • Salmonella • Streptococcus • Clostridium • Conclusions • Bibliography
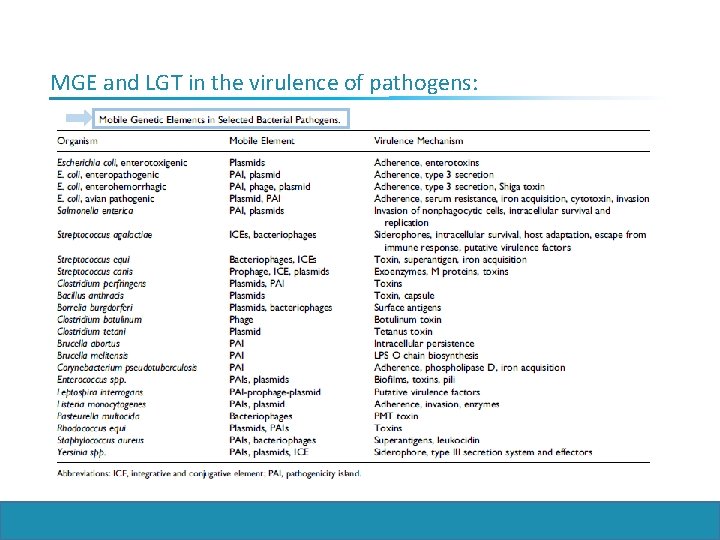
MGE and LGT in the virulence of pathogens:

MGE and HGT in the virulence of pathogens E. coli Pan genome (16000 genes) = core of essential genes (1700 genes) + accessory genome Genome size ranges from 4. 6 Mb to 5. 6 Mb Pathogenic E. coli O 157: H 7 strain vs. Non pathogenic E. coli K 12 strain Enterotoxigenic E. coli posses plasmids that encode the critical virulence elements E. coli O 157: H 7 strain Plasmids, phages, and PAIs transferred to non pathogenic E. coli have created a range of pathogens that cause a variety of diseases

MGE and HGT in the virulence of pathogens Facultative intracellular pathogen that can invade epithelial cells and macrophages Salmonella Variations in presence and sequence of MGEs contribute to differences among strains Core genome + accessory genome (PAIs, bacteriophages and plasmids) Share common ancestor with E. coli Acquisition of SPIs Systemic infection SPI-1: enables to invade the intestinal epithelium SPI-2: required for bacterial survival in the host (they survive in macrophages) Salmonella enterica

MGE and HGT in the virulence of pathogens Streptococcus Important group of human and animal pathogens HGT has an important role S. pyogenes Human specific pathogen Cause variety of diseases Acquision of virulence and colonization factors Several acquired genes are located in prophages S. aglactiae Numerous MGE: PAIs, prophages. . . Genes involved in metabolism, virulence factors. . . Streptococcus pyogenes S. equi subs. zooepidemicus and S. equi subs. equi Losses and gains Accumulation of ISs Acquisition of prophages S. canis Animal pathogen Multiple MGEs Integrated plasmid

MGE and HGT in the virulence of pathogens Clostridium Diseases caused by Clostridium Role of extrachromosomal elements Clostridium toxin genes located on plasmids Gene for tetanus toxin present in 74 -Kba plasmid Clostridium perfringens toxins carried on large conjugative plasmids Clostridium tetani Necrotic enteritis as another example of the role of MGEs in disease: Caused by a specific strain of C. perfringens Mobile virulent plasmids that share same conserved backbone for conjugative transfer

• Introduction • Mechanisms of transfer of DNA • MGE involved in horizontal gene transfer of virulence determinants • MGEs and horizontal gene transfer in the virulence of patogens: • E. coli • Salmonella • Streptococcus • Clostridium • Conclusions • Bibliography

Conclusions Study of HGT contributed to understanding pathogenesis, adaptability and evolution Coexistence of antimicrobial resistance genes and virulent genes on the same MGE: antimicrobial use may coselect for antimicrobial resistance and virulence factors Look changes over million of years of evolution and recognize likely developments in the years ahead More studies need to be done to test the origin and transfer of virulence genes

Bibliography • Dobrindt, U. , Chowdary, M. G. , Krumbholz, G. , & Hacker, J. (2010). Genome dynamics and its impact on evolution of Escherichia coli. Medical Microbiology and Immunology, 199(3), 145– 154. http: //doi. org/10. 1007/s 00430 -0161 -2 • Frost, L. S. , Leplae, R. , Summers, A. O. , & Toussaint, A. (2005). Mobile genetic elements: the agents of open source evolution. Nature Reviews Microbiology, 3(9), 722– 732. http: //doi. org/10. 1038/nrmicro 1235 • Gyles, C. , & Boerlin, P. (2014). Horizontally Transferred Genetic Elements and Their Role in Pathogenesis of Bacterial Disease. Veterinary Pathology, 51(2), 328– 340. http: //doi. org/10. 1177/0300985813511131 • Ibarra, J. A. , & Steele-Mortimer, O. (2009). Salmonella--the ultimate insider. Salmonella virulence factors that modulate intracellular survival. Cellular Microbiology, 11(11), 1579– 86. http: //doi. org/10. 1111/j. 1462 -5822. 2009. 01368. x • Ochman, H. , Lawrence, J. G. , & Groisman, E. A. (2000). Lateral gene transfer and the nature of bacterial innovation. Nature, 405(6784), 299– 304. http: //doi. org/10. 1038/35012500 • Smillie, C. , Garcillán-Barcia, M. P. , Francia, M. V. , Rocha, E. P. C. , & de la Cruz, F. (2010). Mobility of plasmids. Microbiology and Molecular Biology Reviews MMBR, 74(3), 434– 52. http: //doi. org/10. 1128/MMBR. 00020 -10
- Slides: 17