Honma Goto 2001 Elizabeth Samuels Heather Betz What

Honma & Goto 2001 Elizabeth Samuels + Heather Betz
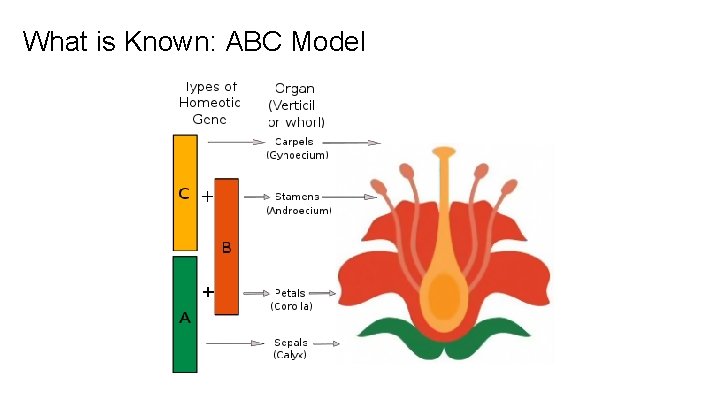
What is Known: ABC Model
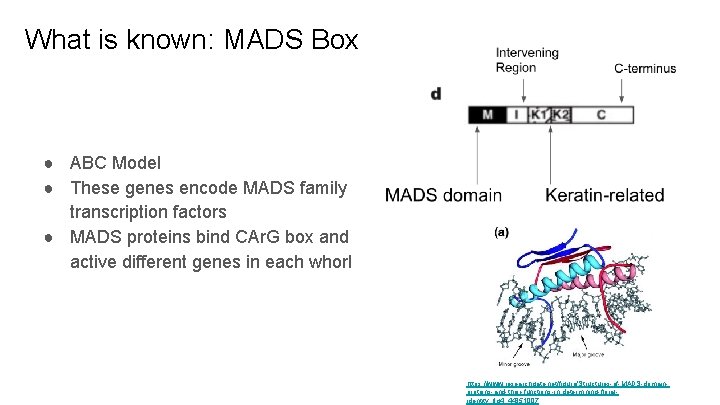
What is known: MADS Box ● ABC Model ● These genes encode MADS family transcription factors ● MADS proteins bind CAr. G box and active different genes in each whorl https: //www. researchgate. net/figure/Structures-of-MADS-domainproteins-and-their-functions-in-determining-floralidentity_fig 4_44851007
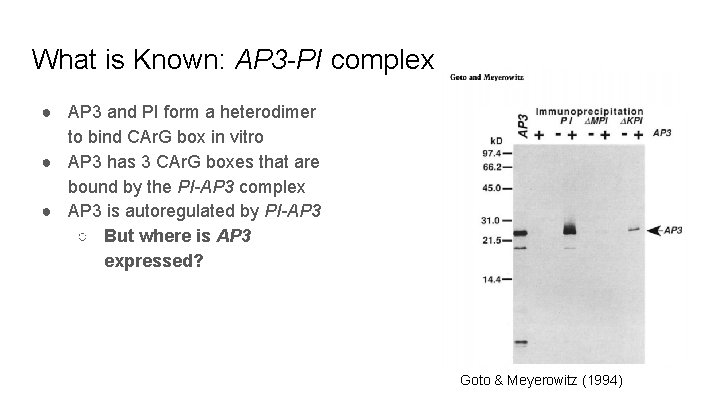
What is Known: AP 3 -PI complex ● AP 3 and PI form a heterodimer to bind CAr. G box in vitro ● AP 3 has 3 CAr. G boxes that are bound by the PI-AP 3 complex ● AP 3 is autoregulated by PI-AP 3 ○ But where is AP 3 expressed? Goto & Meyerowitz (1994)

Where is AP 3 expressed? If AP 3 and PI are sufficient to activate the AP 3 promoter Then we would expect that in a plant where AP 3 and PI are expressed constitutively (via 35 S promoter), the AP 3 promoter would also be active throughout the plant Figure 2 a Was that the case?
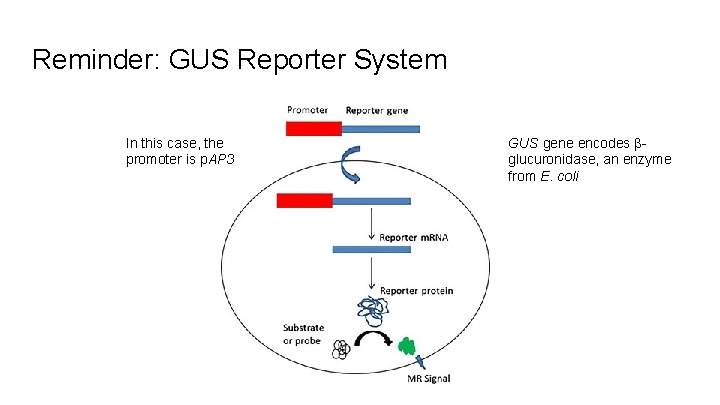
Reminder: GUS Reporter System In this case, the promoter is p. AP 3 GUS gene encodes βglucuronidase, an enzyme from E. coli

Where is AP 3 expressed? AP 3: GUS fusion gene in plants expressing 35 S: : PI 35 S: : AP 3 Only see blue in floral areas What does this mean? Figure 2 a
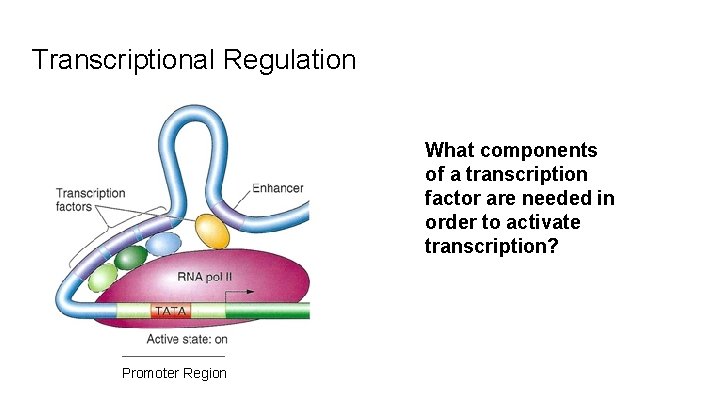
Transcriptional Regulation What components of a transcription factor are needed in order to activate transcription? Promoter Region
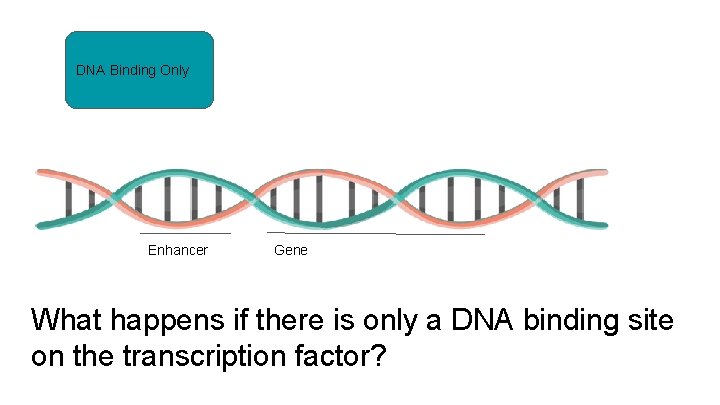
DNA Binding Only Enhancer Gene What happens if there is only a DNA binding site on the transcription factor?
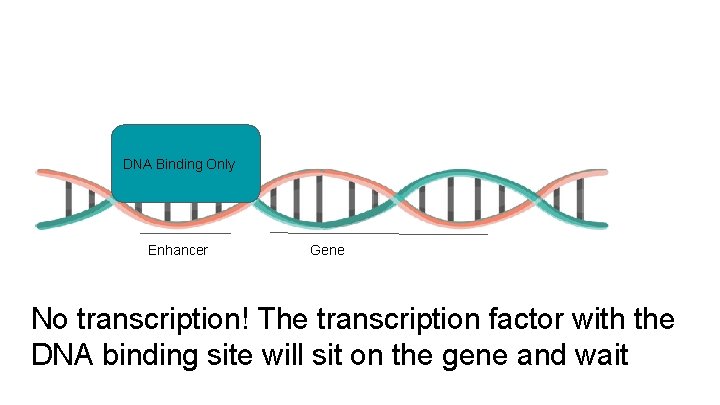
DNA Binding Only Enhancer Gene No transcription! The transcription factor with the DNA binding site will sit on the gene and wait
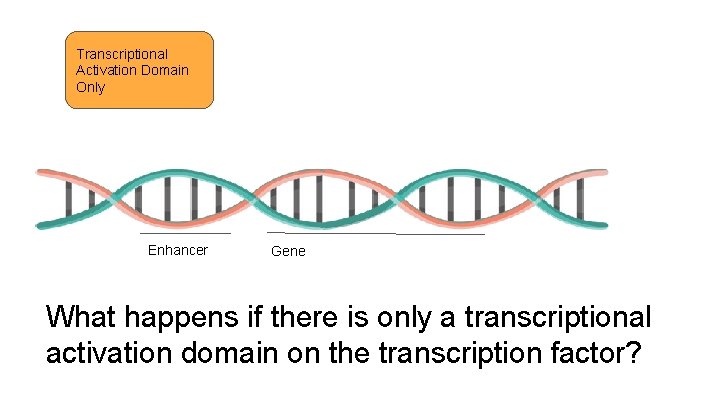
Transcriptional Activation Domain Only Enhancer Gene What happens if there is only a transcriptional activation domain on the transcription factor?
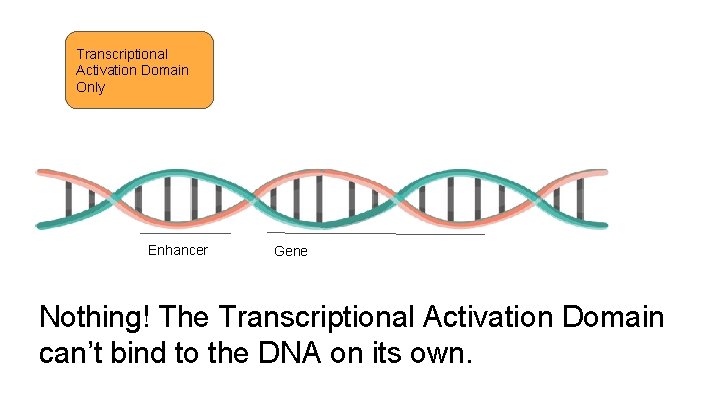
Transcriptional Activation Domain Only Enhancer Gene Nothing! The Transcriptional Activation Domain can’t bind to the DNA on its own.
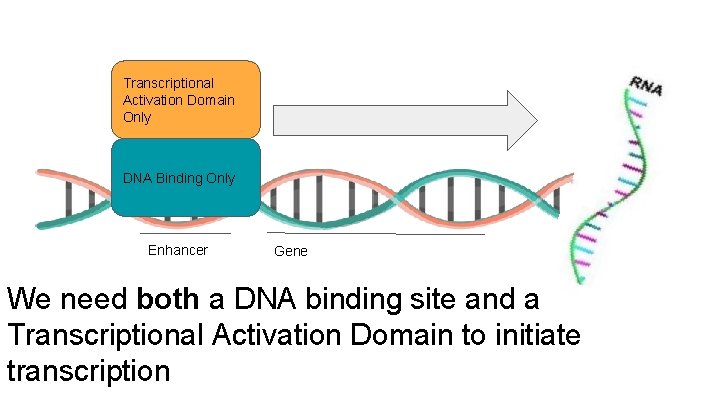
Transcriptional Activation Domain Only DNA Binding Only Enhancer Gene We need both a DNA binding site and a Transcriptional Activation Domain to initiate transcription

Back to AP 3 transcriptional activation AP 3 transcription is only activated in floral regions, even when PI-AP 3 complex is expressed throughout Recall, this figure is AP 3: : GUS fusion gene in plants Figure 2 a expressing 35 S: : PI 35 S: : AP 3

If ABC genes were sufficient to produce floral organs, we would expect ectopic expression to make organs in vegetative organs

Krizek and Meyerowitz, 1996 Figure 1, p 35 S: : PI p 35 s: : AP 3 plants compared to wildtype. BUT - Altering expression of ABC genes doesn’t turn vegetative organs into floral organs “Our results show that AG alone cannot convert leaves into floral organs. ” (Mizukami and Ma, 1992)
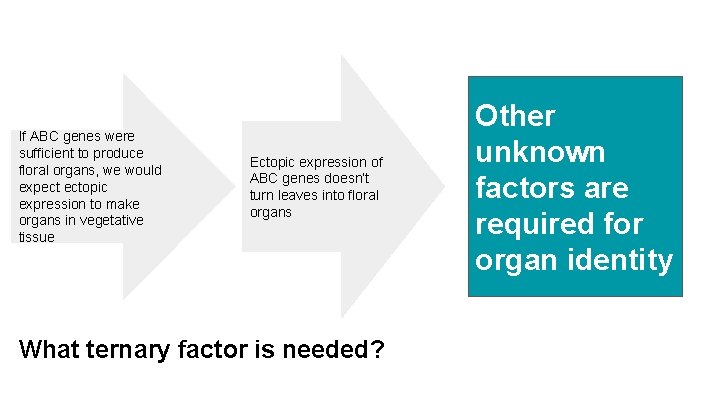
If ABC genes were sufficient to produce floral organs, we would expect ectopic expression to make organs in vegetative tissue Ectopic expression of ABC genes doesn’t turn leaves into floral organs What ternary factor is needed? Other unknown factors are required for organ identity
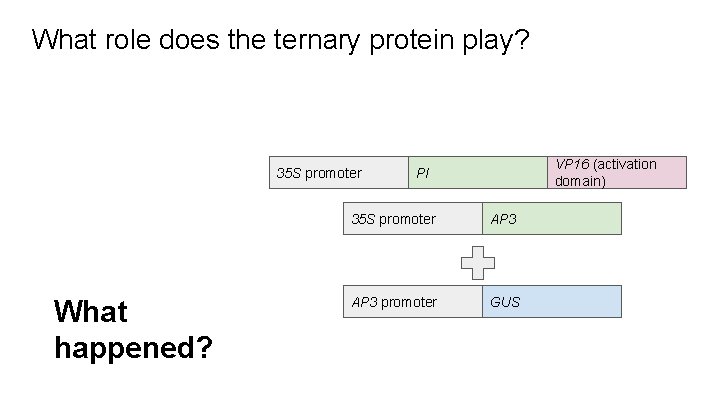
What role does the ternary protein play? 35 S promoter What happened? VP 16 (activation domain) PI 35 S promoter AP 3 promoter GUS
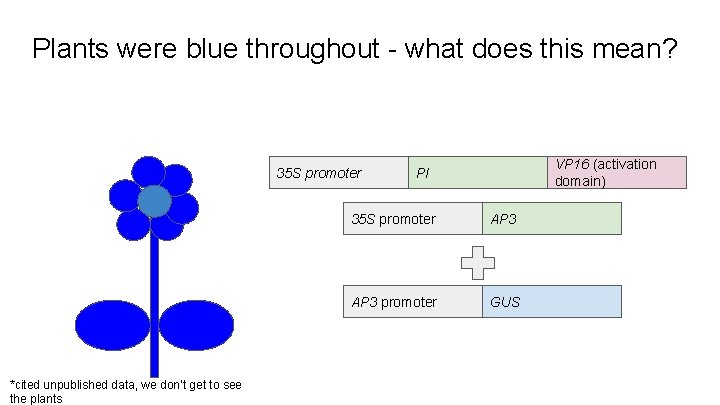
Plants were blue throughout - what does this mean? 35 S promoter *cited unpublished data, we don’t get to see the plants VP 16 (activation domain) PI 35 S promoter AP 3 promoter GUS
Hypothesis: A ternary (third) protein is responsible for modulating transcriptional activity
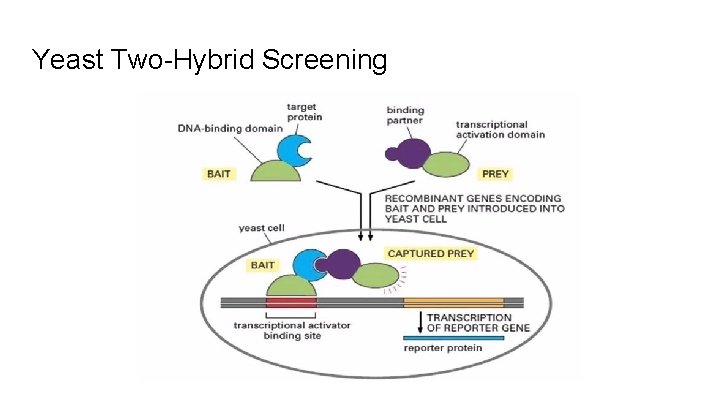
Yeast Two-Hybrid Screening
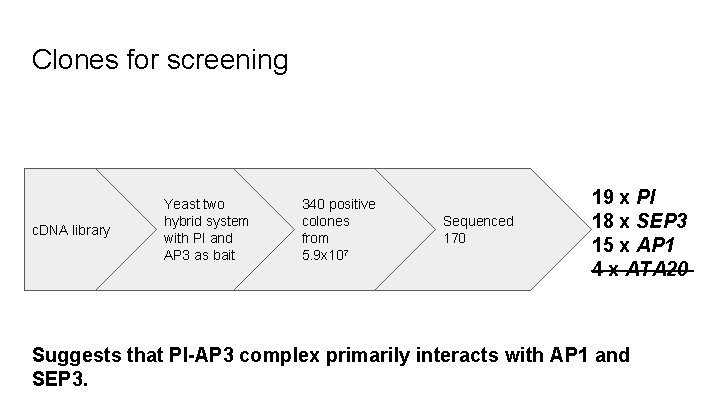
Clones for screening c. DNA library Yeast two hybrid system with PI and AP 3 as bait 340 positive colones from 5. 9 x 107 Sequenced 170 19 x PI 18 x SEP 3 15 x AP 1 4 x ATA 20 Suggests that PI-AP 3 complex primarily interacts with AP 1 and SEP 3.

SEP 3 ● SEP 3 (previously called AGL 9) ● MADS-box family of genes ○ SEP 1, SEP 2, SEP 3 ● SEP 3 Expressed in whorls 2 -4 ○ SEP 1 + 2 in all whorls
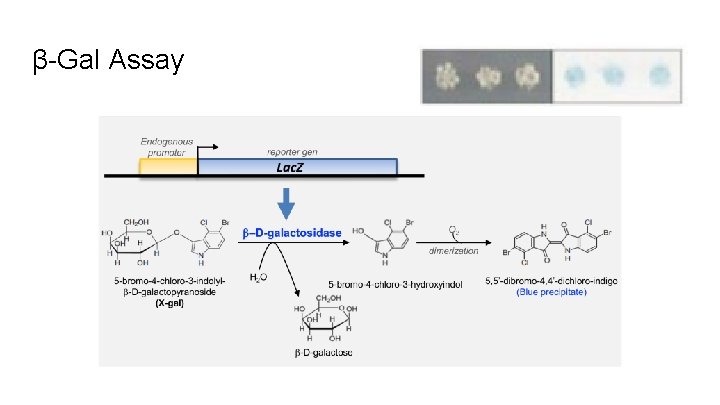
β-Gal Assay
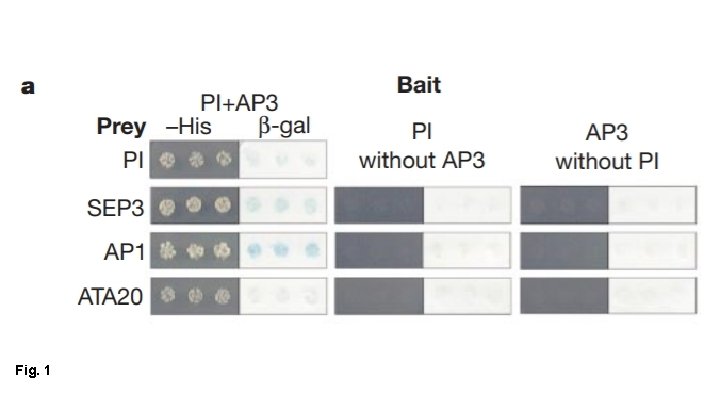
Fig. 1
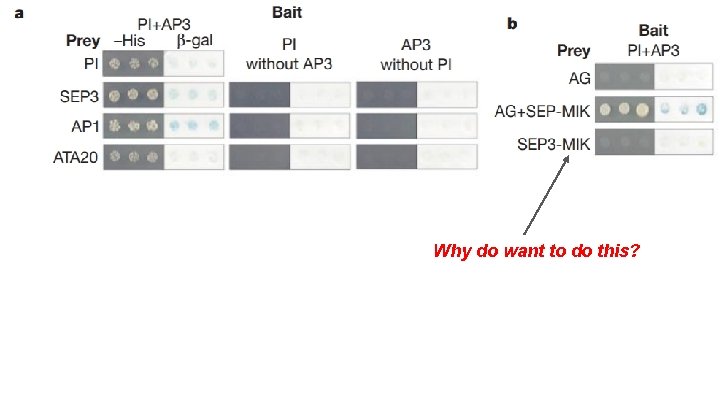
Why do want to do this?
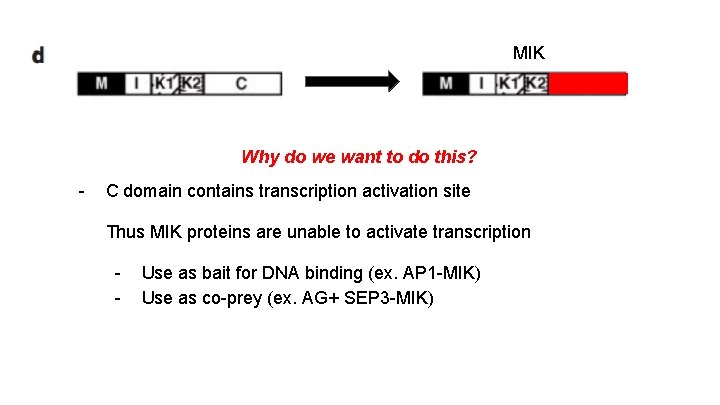
MIK Why do we want to do this? - C domain contains transcription activation site Thus MIK proteins are unable to activate transcription - Use as bait for DNA binding (ex. AP 1 -MIK) Use as co-prey (ex. AG+ SEP 3 -MIK)
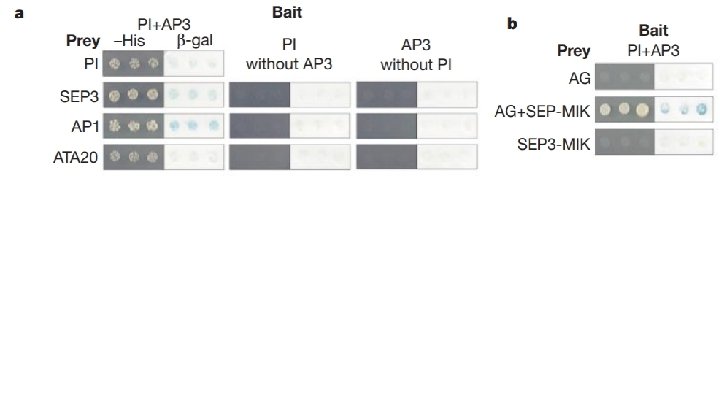
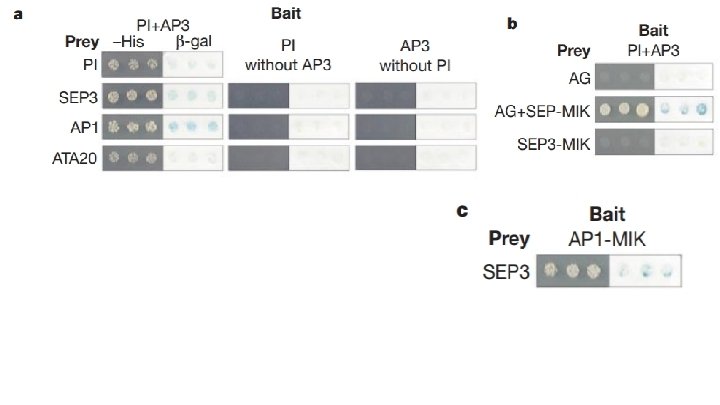
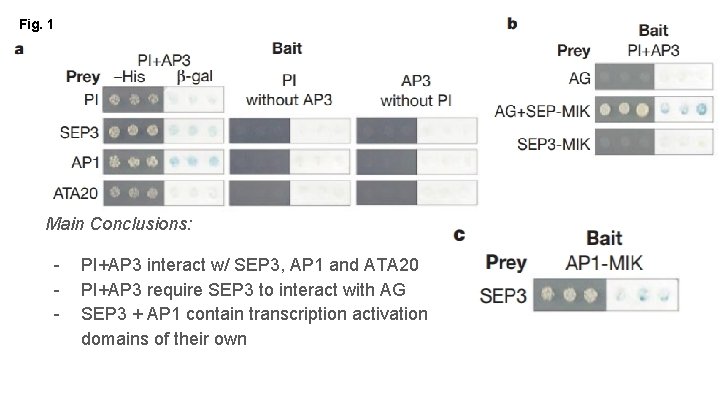
Fig. 1 Main Conclusions: - PI+AP 3 interact w/ SEP 3, AP 1 and ATA 20 PI+AP 3 require SEP 3 to interact with AG SEP 3 + AP 1 contain transcription activation domains of their own
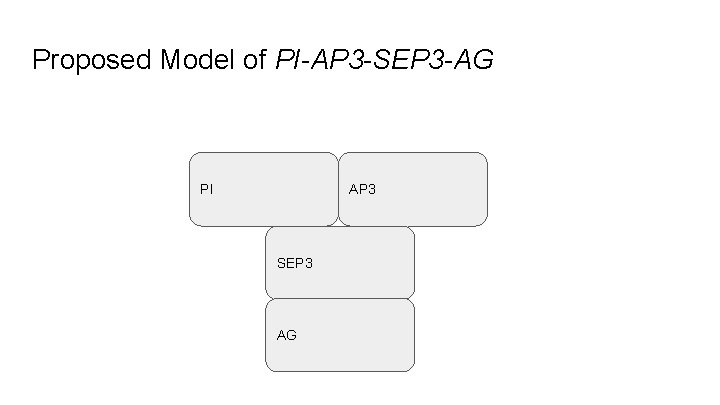
Proposed Model of PI-AP 3 -SEP 3 -AG PI AP 3 SEP 3 AG
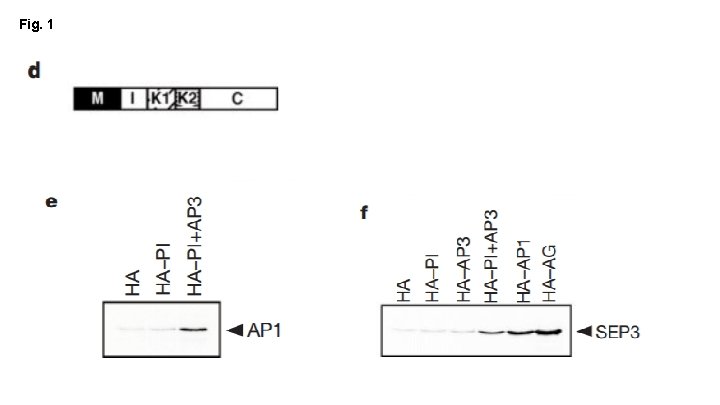
Fig. 1
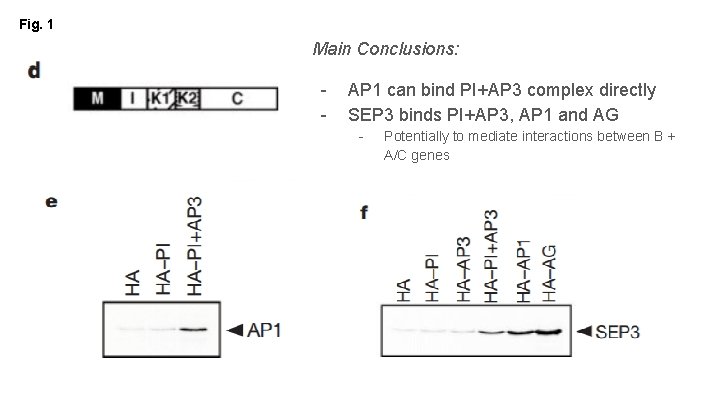
Fig. 1 Main Conclusions: - AP 1 can bind PI+AP 3 complex directly SEP 3 binds PI+AP 3, AP 1 and AG - Potentially to mediate interactions between B + A/C genes
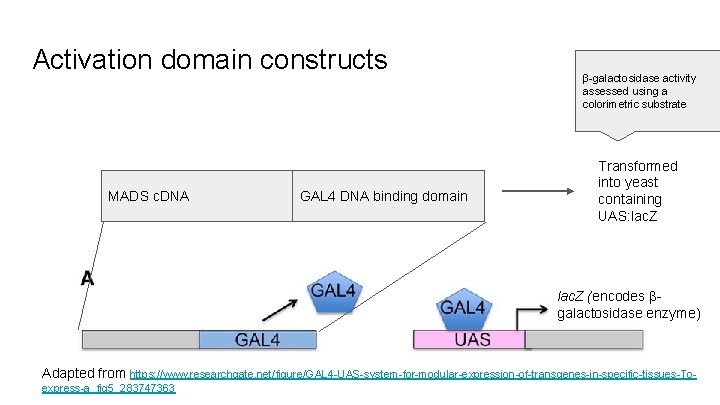
Activation domain constructs MADS c. DNA GAL 4 DNA binding domain β-galactosidase activity assessed using a colorimetric substrate Transformed into yeast containing UAS: lac. Z (encodes βgalactosidase enzyme) Adapted from https: //www. researchgate. net/figure/GAL 4 -UAS-system-for-modular-expression-of-transgenes-in-specific-tissues-Toexpress-a_fig 5_283747363
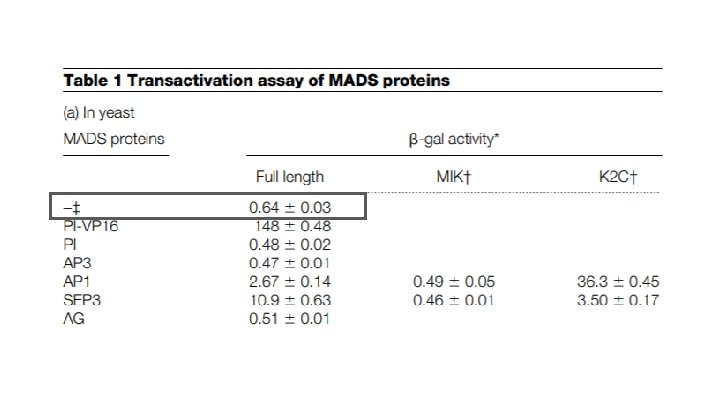
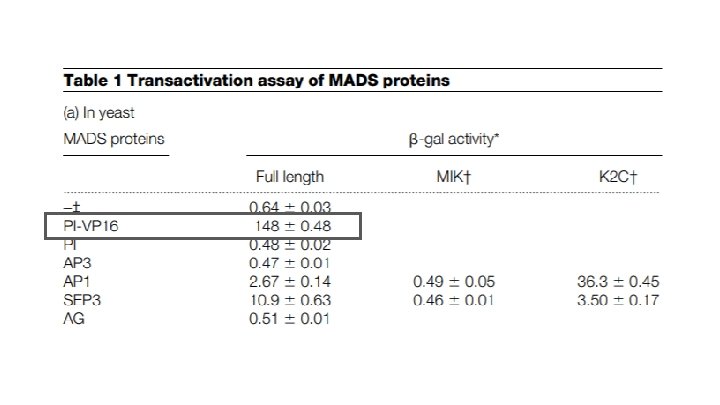
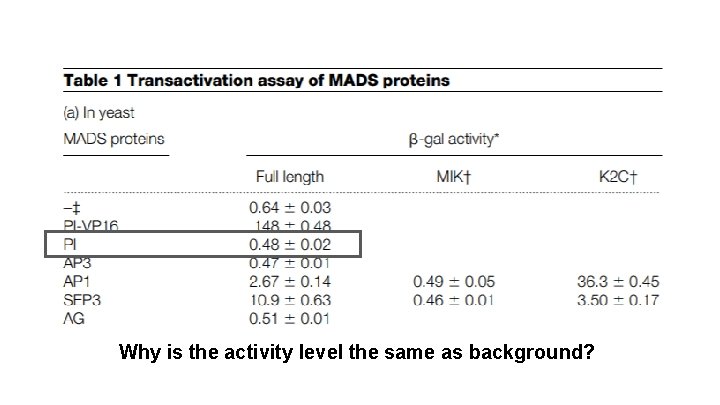
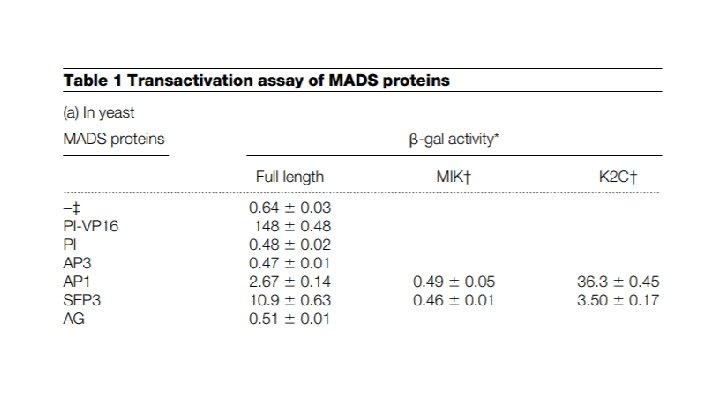
Why is the activity level the same as background?
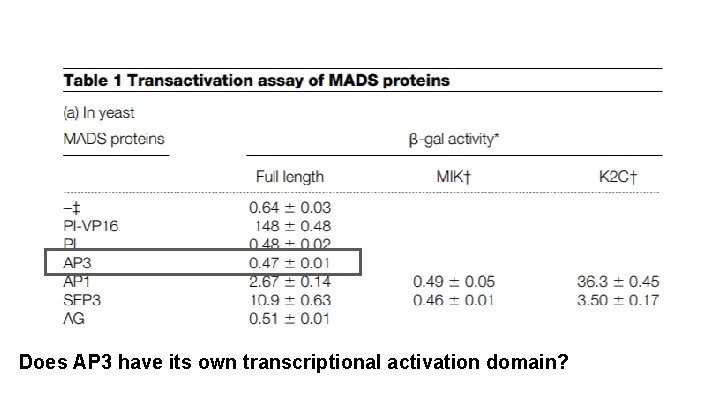
Does AP 3 have its own transcriptional activation domain?
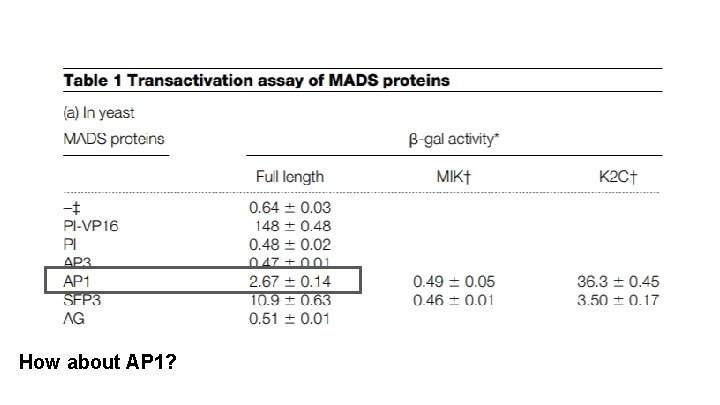
How about AP 1?

What do the MIK and K 2 C sections represent?
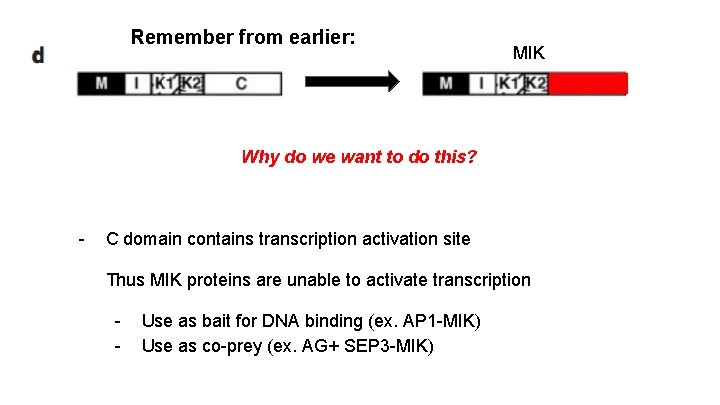
Remember from earlier: MIK Why do we want to do this? - C domain contains transcription activation site Thus MIK proteins are unable to activate transcription - Use as bait for DNA binding (ex. AP 1 -MIK) Use as co-prey (ex. AG+ SEP 3 -MIK)
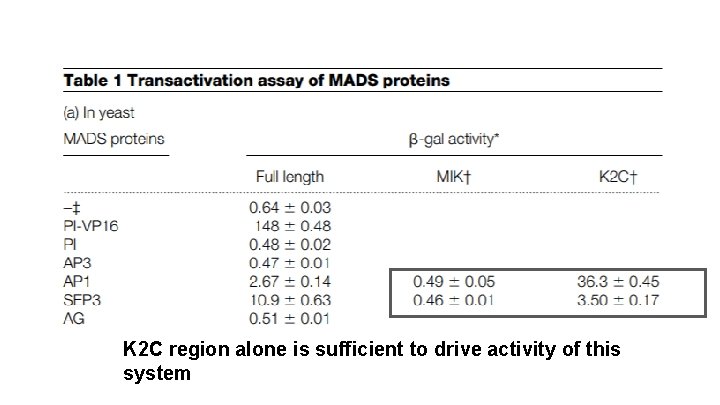
K 2 C region alone is sufficient to drive activity of this system

SEP 3 also has its own transcriptional activation domain
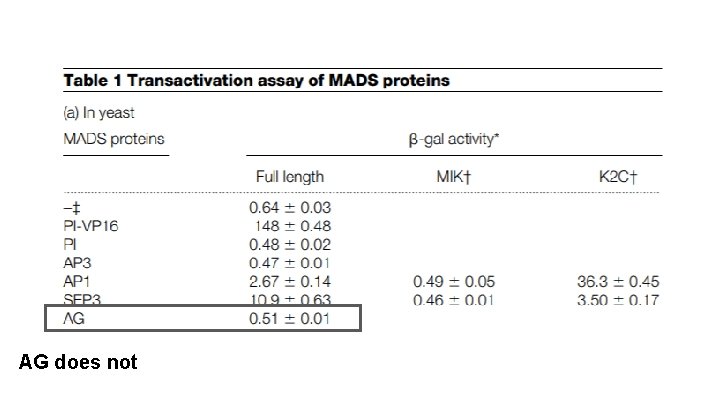
AG does not

Summary: 1. Only AP 1 and SEP 3 contain their own transcriptional activation domains 2. The activation domain is within the K 2 C region of the gene

Table 1(b) What is LUC?
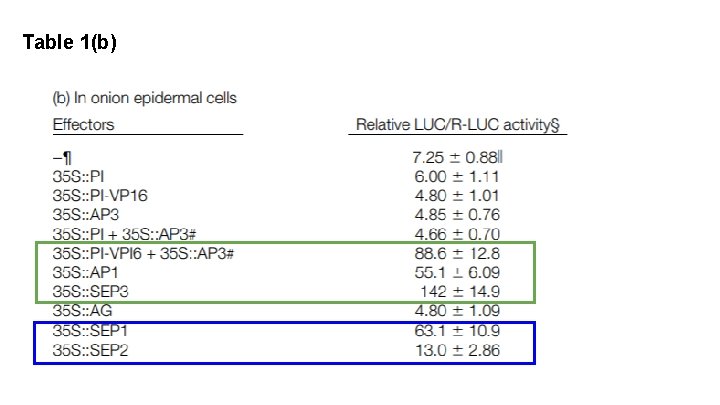
Table 1(b)

Fig. 2 35 S: : AP 3; 35 S: : PI AP 3: : GUS reporter 35 S: : AP 1 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 35 S: : SEP 3 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3

Fig. 2 35 S: : AP 3; 35 S: : PI AP 3: : GUS reporter 35 S: : AP 1 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 35 S: : SEP 3 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3
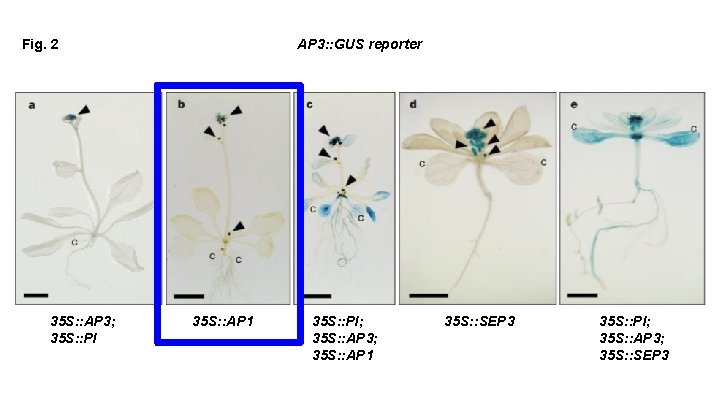
Fig. 2 35 S: : AP 3; 35 S: : PI AP 3: : GUS reporter 35 S: : AP 1 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 35 S: : SEP 3 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3
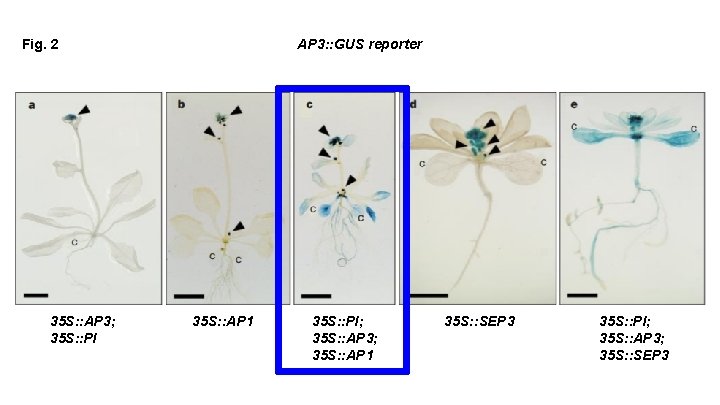
Fig. 2 35 S: : AP 3; 35 S: : PI AP 3: : GUS reporter 35 S: : AP 1 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 35 S: : SEP 3 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3
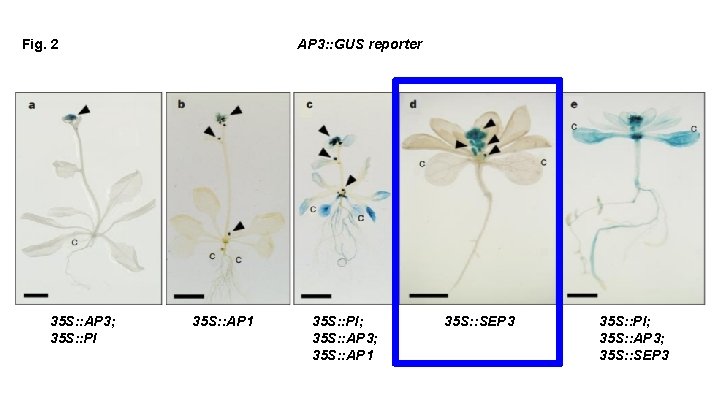
Fig. 2 35 S: : AP 3; 35 S: : PI AP 3: : GUS reporter 35 S: : AP 1 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 35 S: : SEP 3 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3
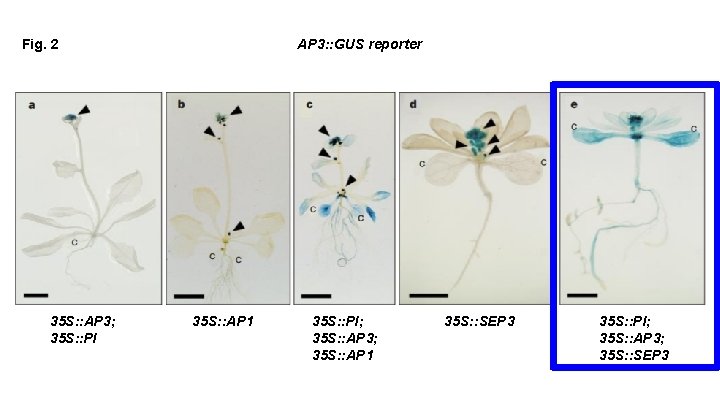
Fig. 2 35 S: : AP 3; 35 S: : PI AP 3: : GUS reporter 35 S: : AP 1 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 35 S: : SEP 3 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3
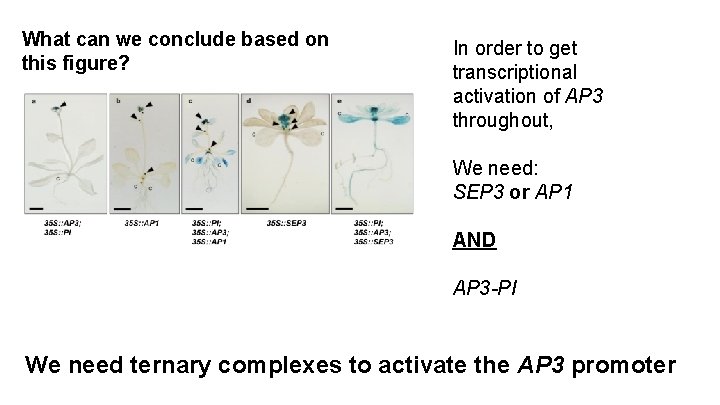
What can we conclude based on this figure? In order to get transcriptional activation of AP 3 throughout, We need: SEP 3 or AP 1 AND AP 3 -PI We need ternary complexes to activate the AP 3 promoter
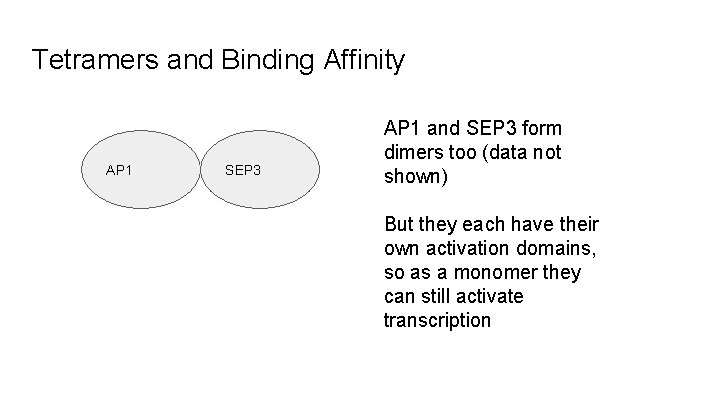
Tetramers and Binding Affinity AP 1 SEP 3 AP 1 and SEP 3 form dimers too (data not shown) But they each have their own activation domains, so as a monomer they can still activate transcription
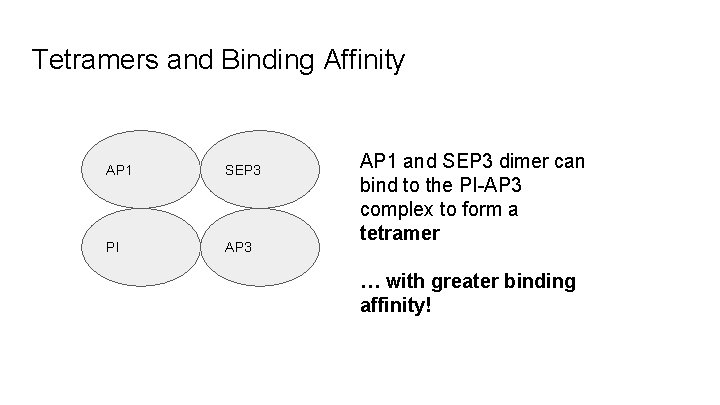
Tetramers and Binding Affinity AP 1 SEP 3 PI AP 3 AP 1 and SEP 3 dimer can bind to the PI-AP 3 complex to form a tetramer … with greater binding affinity!

Fig. 3 - Wild Type DOI: 10. 5772/53127

Fig. 3 - 35 S: : SEP 3 WT 35 S: : SEP 3

Fig. 3 - 35 S: : PI; 35 S: : AP 3 WT 35 S: : PI; 35 S: : AP 3

Fig. 3 - 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3 WT petal Mutant petalloid organ 35 S: : SEP 3 rosette leaf
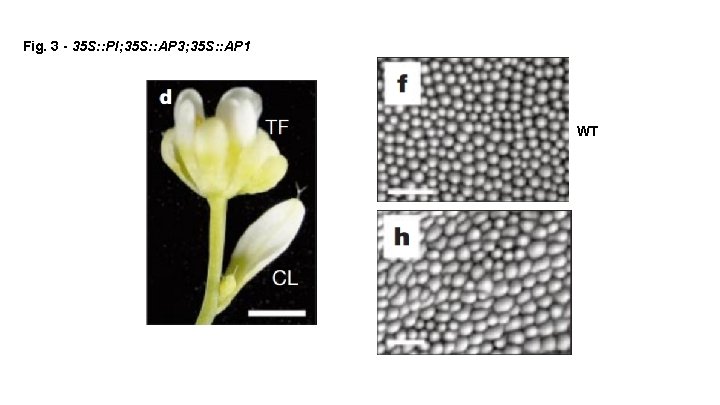
Fig. 3 - 35 S: : PI; 35 S: : AP 3; 35 S: : AP 1 WT
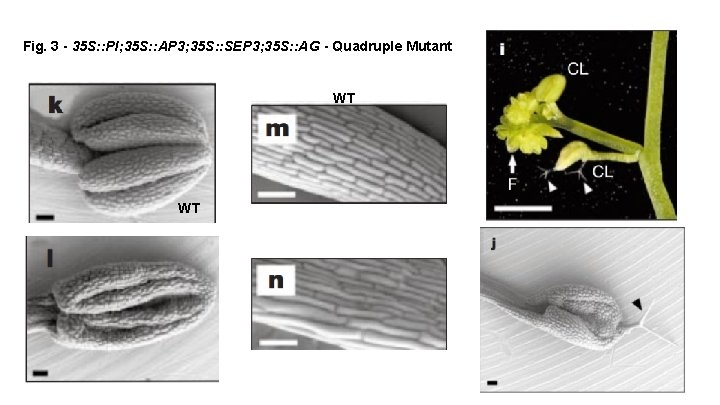
Fig. 3 - 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3; 35 S: : AG - Quadruple Mutant WT WT
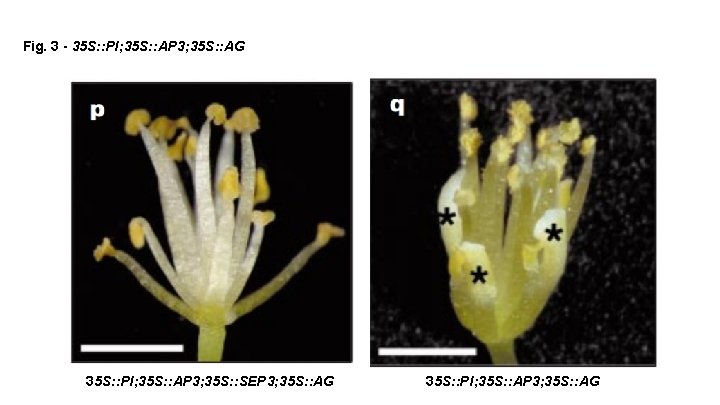
Fig. 3 - 35 S: : PI; 35 S: : AP 3; 35 S: : AG 35 S: : PI; 35 S: : AP 3; 35 S: : SEP 3; 35 S: : AG 35 S: : PI; 35 S: : AP 3; 35 S: : AG

Summary of Main Findings ● AP 1 and SEP 3 have transcriptional-activator domains, which are localized primarily within their C domains, and are able to supply them to PI-AP 3 or AG following complex formation. ● PI, AP 3 and AG do not have their own transcriptional activator domains ● Flower-specific expression of SEP 3 restrictions the ABC genes to floral areas ● AP 3 -PI requires SEP 3 to interact with AG ● Overexpression of SEP 3, AG, PI + AP 3 converts leaves into staminoid organs ● Overexpression of AP 1, PI + AP 3 converts leaves into petalloid organs The ABC Model is incomplete, and we must factor in SEP 3!
- Slides: 65