Homology modelling Intro Proteins Modelling 8 Steps Detect



















- Slides: 32

Homology modelling ? Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate NMR ? X-ray ? ©CMBI 2002

Homology Modelling ! Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

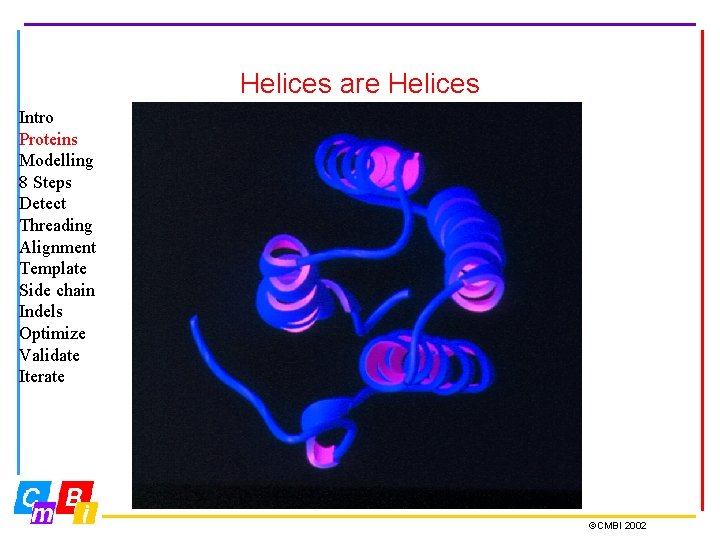
Helices are Helices Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

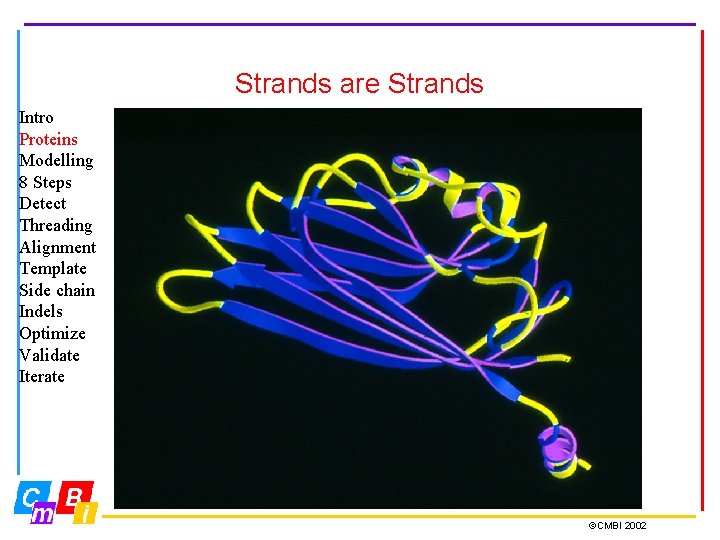
Strands are Strands Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

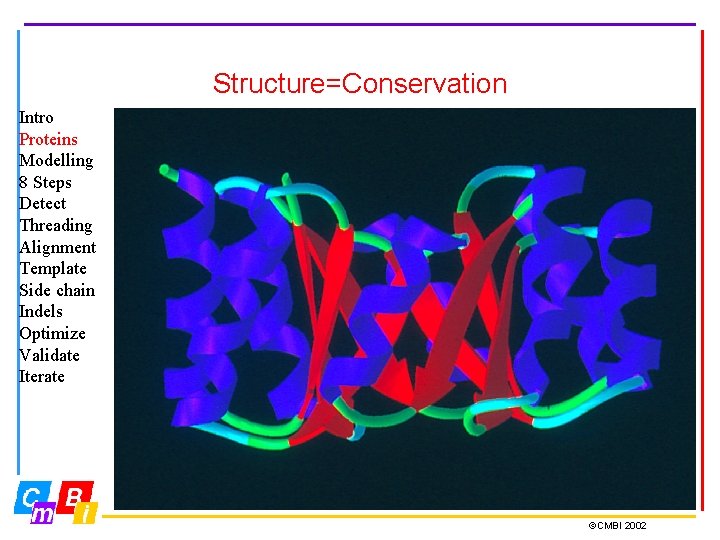
Structure=Conservation Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

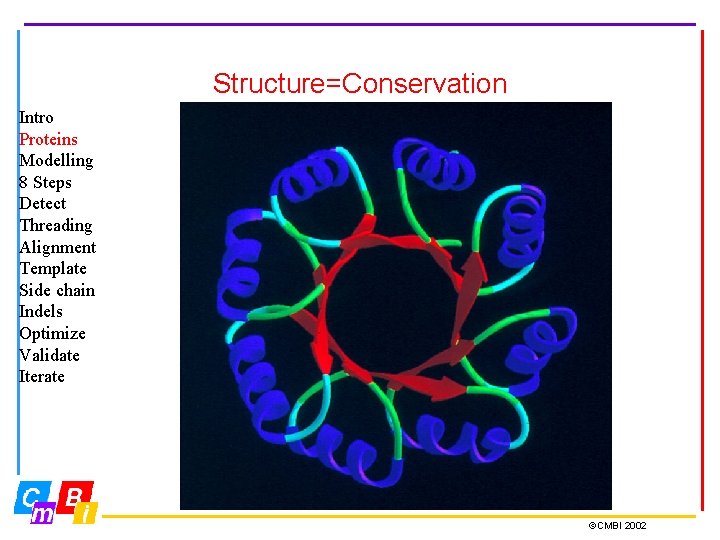
Structure=Conservation Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

Modelling beats X-ray Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

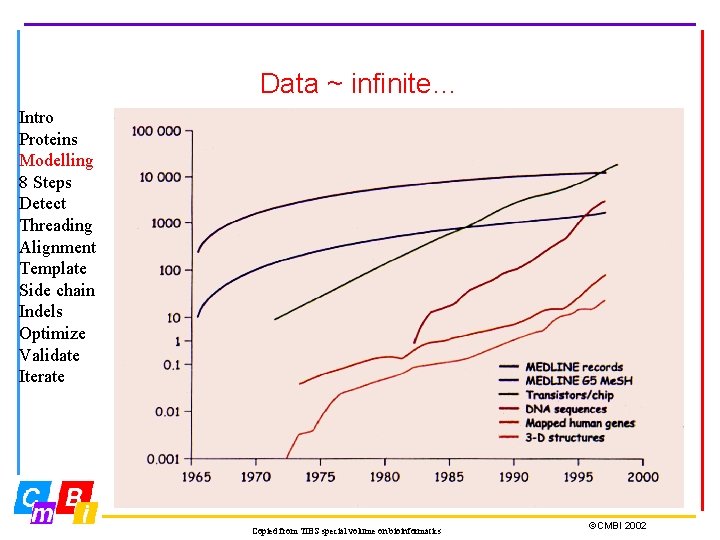
Data ~ infinite… Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Copied from TIBS special volume on bioinformatics ©CMBI 2002

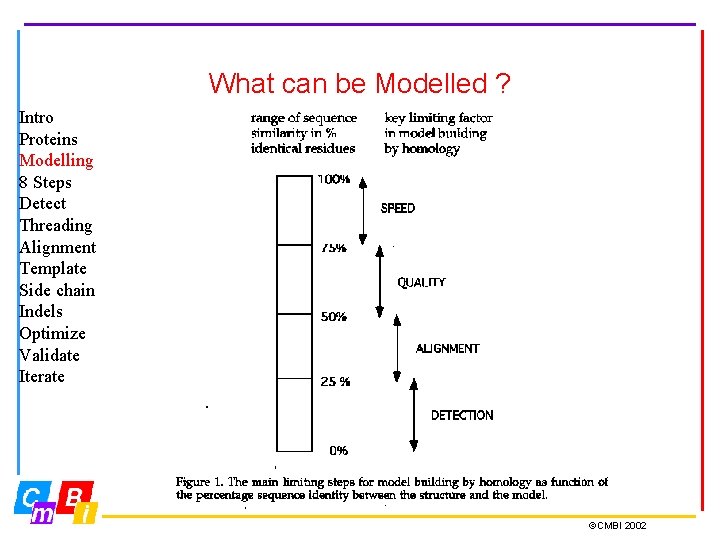
What can be Modelled ? Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

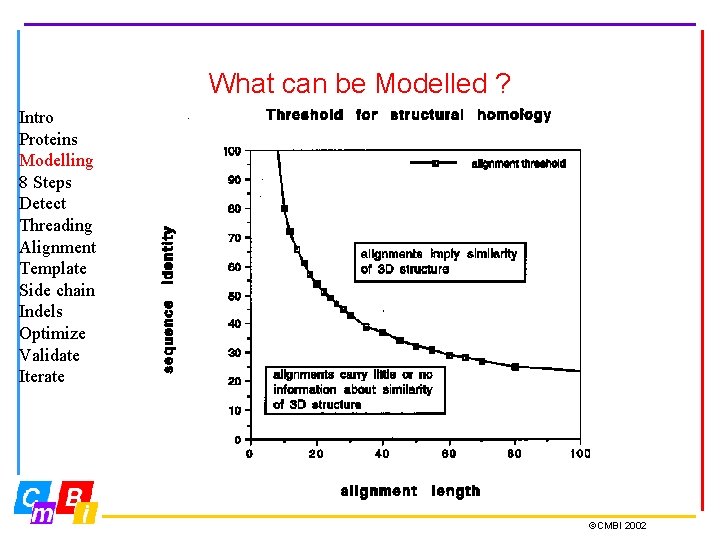
What can be Modelled ? Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

The ‘ 8’ Steps of Modelling Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate 1 2 3 4 5 6 7 8 9 Detect template Get alignment Optimize template Exchange side chains Deal with insertions/deletions Optimize model Validate Iterate ©CMBI 2002

Template detection Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Normally BLAST is good enough. If BLAST doesn’t find a template, you should not want to build a model anyway. When desperate, use PSIBLAST (on PDB + Swiss. Prot), or use threading. ©CMBI 2002

Threading Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Threading means using information from the template structure to: • detect homology • to improve an alignment. ©CMBI 2002

Threading Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Small residues ©CMBI 2002

Threading Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Alcoholic residues ©CMBI 2002

Threading Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate The folded ‘protein’ ©CMBI 2002

Threading Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Two aligned ‘proteins’ ©CMBI 2002

Alignment Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate BLAST the model sequence. BLAST the template sequence. Select 50 -100 representatives. Do multiple sequence alignment. Keep only model and template from that multiple sequence alignment. ©CMBI 2002

How to align: Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ASASAS YPYPYP (three ways…) ©CMBI 2002

How to align: Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ASASASAYAYAY-YPYPYP (two ways…) ©CMBI 2002

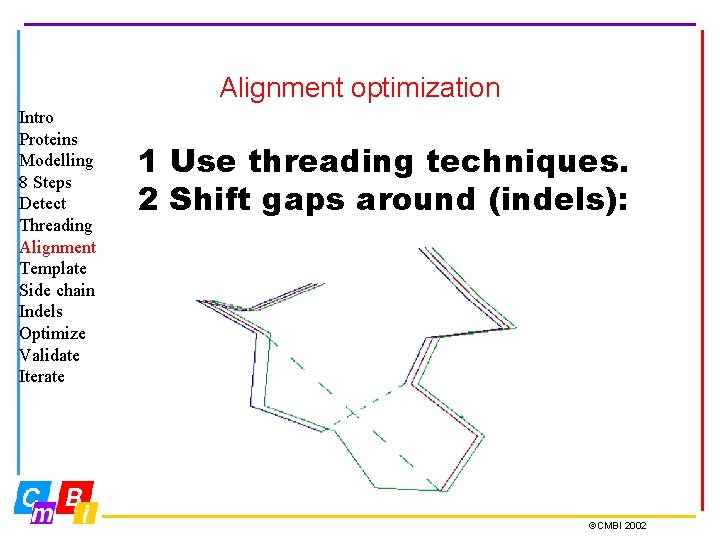
Alignment optimization Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate 1 Use threading techniques. 2 Shift gaps around (indels): ©CMBI 2002

Select ‘best’ template Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

Deal with errors Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

Exchange side chains Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Keep template rigid. Determine best rotamer. Do NOT optimize rotamers. If best rotamer doesn’t fit, start thinking. If the model is bad, you had the wrong template, or the wrong alignment. Make sure your model can exist. ©CMBI 2002

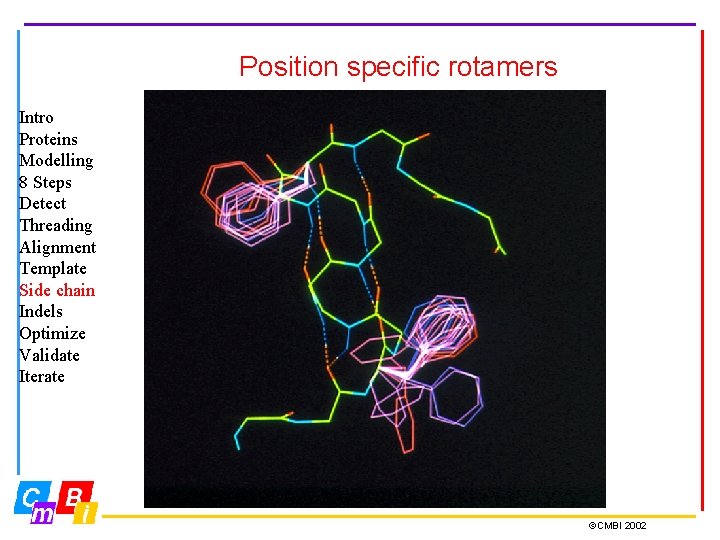
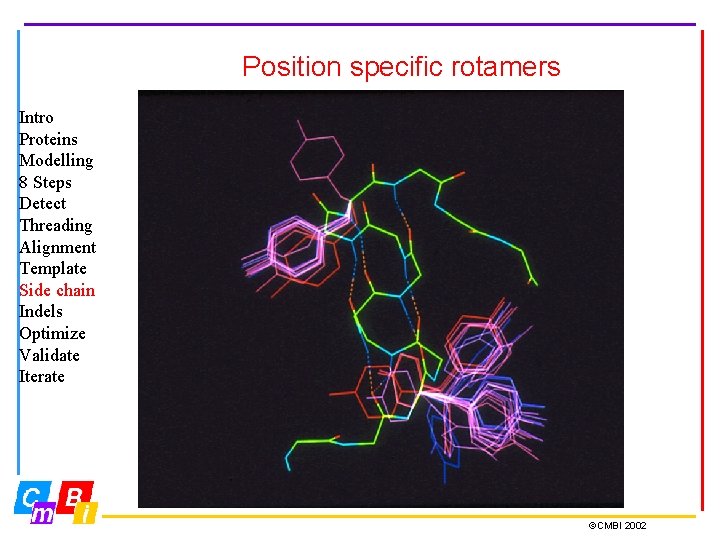
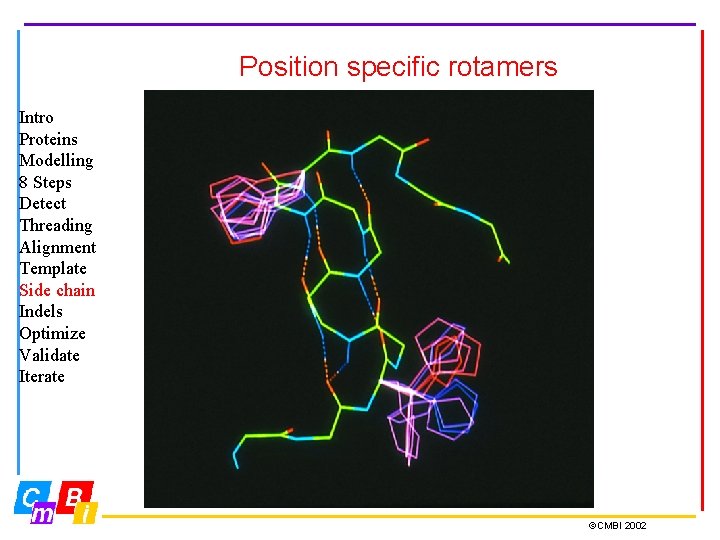
Position specific rotamers Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

Position specific rotamers Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

Position specific rotamers Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate ©CMBI 2002

Insertions - Deletions Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Insertions are impossible Deletions: Move gap around in template till end point distance is short. If this is not possible, you have either the wrong template, or the wrong alignment. ©CMBI 2002

Model Optimization Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Do NOT use molecular dynamics (unless you know that you know what you are doing): ©CMBI 2002

Model Optimization Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Use 25 – 50 steps energy minimization, or use a force field that has been especially designed for the optimization of homology models. ©CMBI 2002

Validation Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Use the WHAT_CHECK server. (Next seminar). Use your brains. Compare with structures. Compare with Meta-server. ©CMBI 2002

The ‘ 8’ Steps of Modelling Intro Proteins Modelling 8 Steps Detect Threading Alignment Template Side chain Indels Optimize Validate Iterate Depending on the WHAT_CHECK result, you can restart at each of the previous steps in the modelling process. ©CMBI 2002