Homology and sequence alignment 1 Homology Similarity between

Homology and sequence alignment. 1
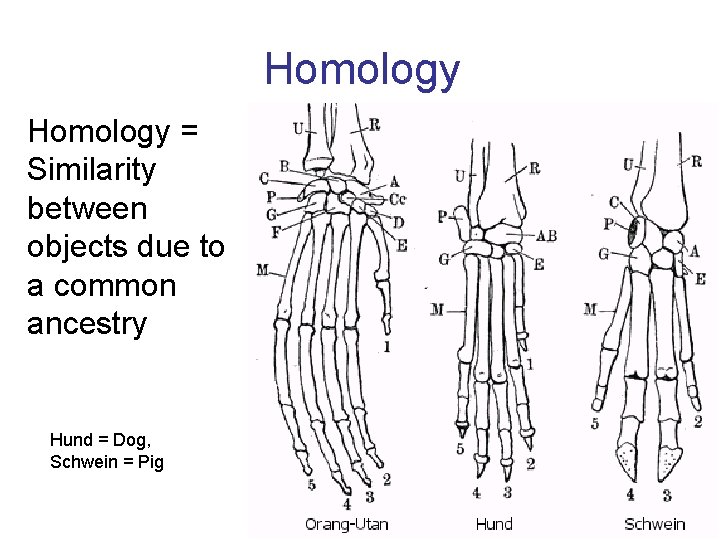
Homology = Similarity between objects due to a common ancestry Hund = Dog, Schwein = Pig

Sequence homology Similarity between sequences as a result of common ancestry. VLSPAVKWAKVGAHAAGHG ||| || |||| VLSEAVLWAKVEADVAGHG 3

Sequence alignment Alignment: Comparing two (pairwise) or more (multiple) sequences. Searching for a series of identical or similar characters in the sequences. 4

Why align? VLSPAVKWAKV ||| || |||| VLSEAVLWAKV 1. To detect if two sequence are homologous. If so, homology may indicate similarity in function (and structure). 2. Required for evolutionary studies (e. g. , tree reconstruction). 3. To detect conservation (e. g. , a tyrosine that is evolutionary conserved is more likely to be a phosphorylation site). 5
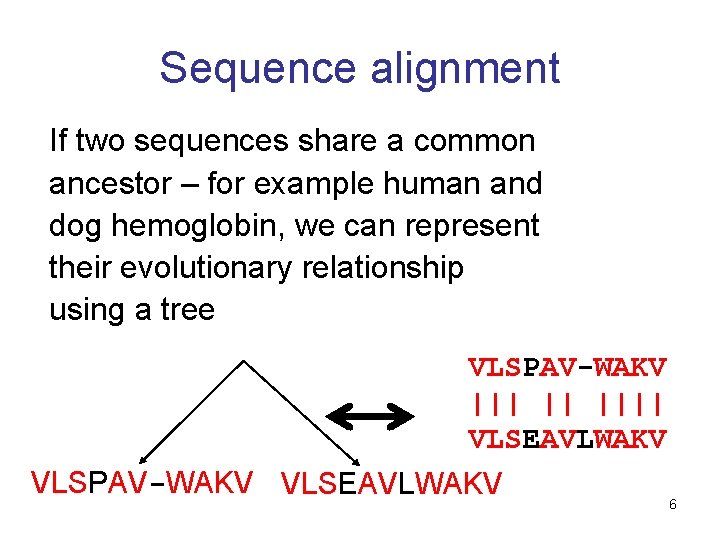
Sequence alignment If two sequences share a common ancestor – for example human and dog hemoglobin, we can represent their evolutionary relationship using a tree VLSPAV-WAKV ||| || |||| VLSEAVLWAKV VLSPAV-WAKV VLSEAVLWAKV 6
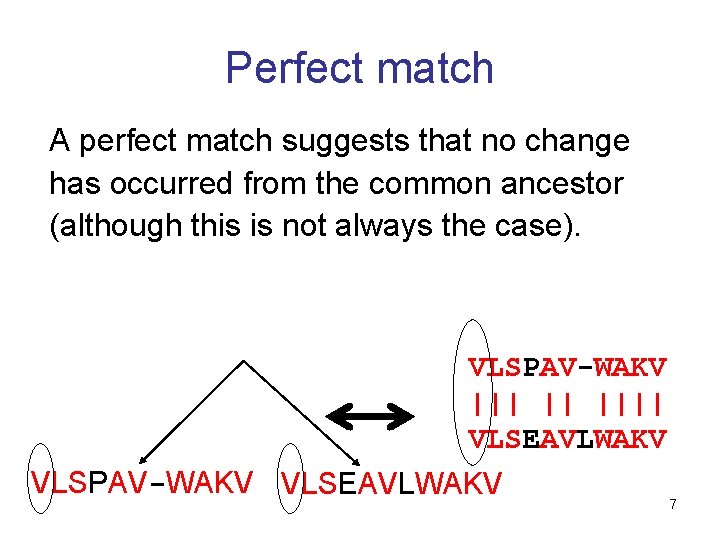
Perfect match A perfect match suggests that no change has occurred from the common ancestor (although this is not always the case). VLSPAV-WAKV ||| || |||| VLSEAVLWAKV VLSPAV-WAKV VLSEAVLWAKV 7

A substitution suggests that at least one change has occurred since the common ancestor (although we cannot say in which lineage it has occurred). VLSPAV-WAKV ||| || |||| VLSEAVLWAKV VLSPAV-WAKV VLSEAVLWAKV 8
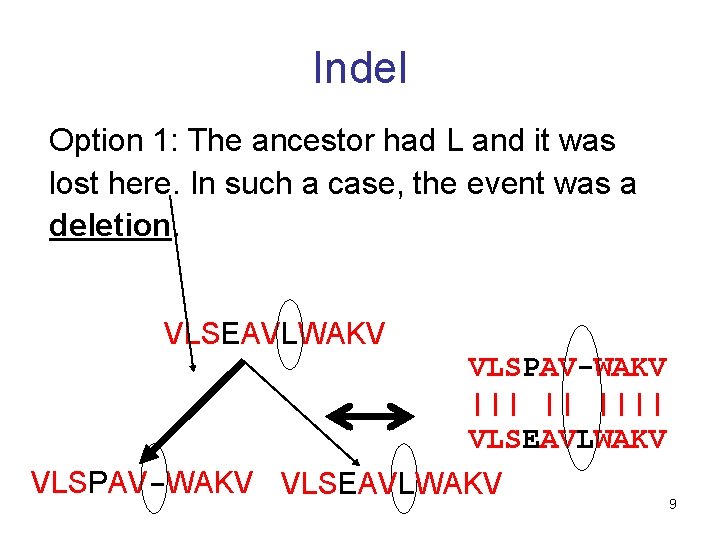
Indel Option 1: The ancestor had L and it was lost here. In such a case, the event was a deletion. VLSEAVLWAKV VLSPAV-WAKV ||| || |||| VLSEAVLWAKV VLSPAV-WAKV VLSEAVLWAKV 9
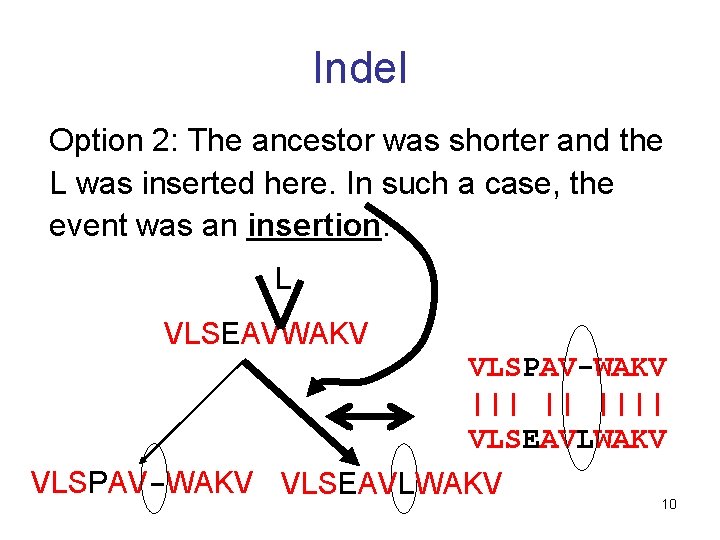
Indel Option 2: The ancestor was shorter and the L was inserted here. In such a case, the event was an insertion. L VLSEAVWAKV VLSPAV-WAKV ||| || |||| VLSEAVLWAKV VLSPAV-WAKV VLSEAVLWAKV 10
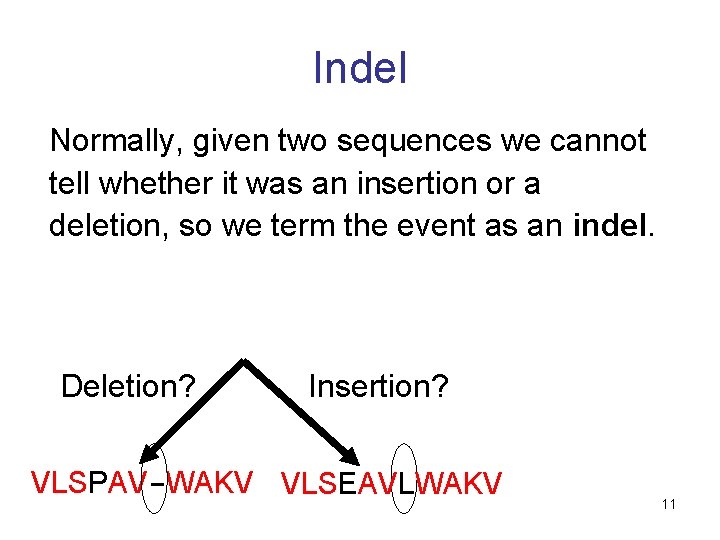
Indel Normally, given two sequences we cannot tell whether it was an insertion or a deletion, so we term the event as an indel. Deletion? Insertion? VLSPAV-WAKV VLSEAVLWAKV 11
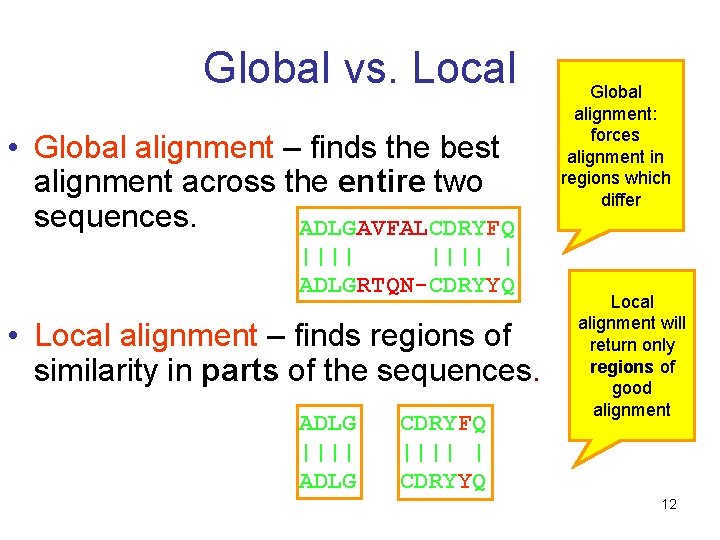
Global vs. Local • Global alignment – finds the best alignment across the entire two sequences. ADLGAVFALCDRYFQ |||| | ADLGRTQN-CDRYYQ • Local alignment – finds regions of similarity in parts of the sequences. ADLG |||| ADLG CDRYFQ |||| | CDRYYQ Global alignment: forces alignment in regions which differ Local alignment will return only regions of good alignment 12
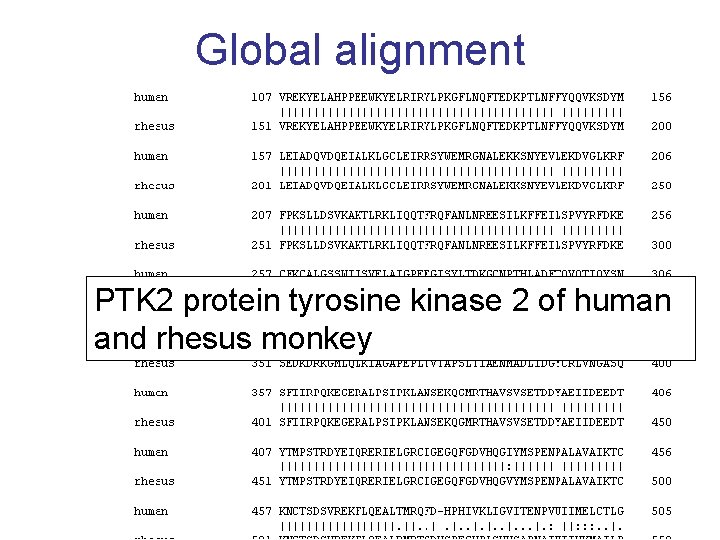
Global alignment PTK 2 protein tyrosine kinase 2 of human and rhesus monkey 13
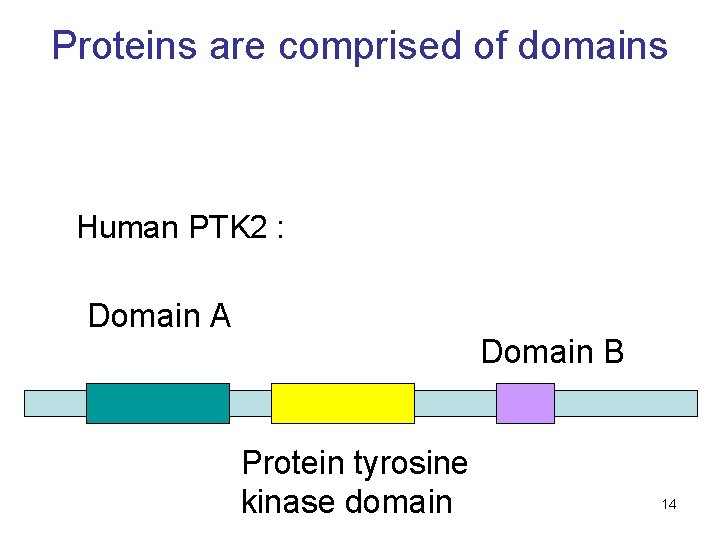
Proteins are comprised of domains Human PTK 2 : Domain A Domain B Protein tyrosine kinase domain 14
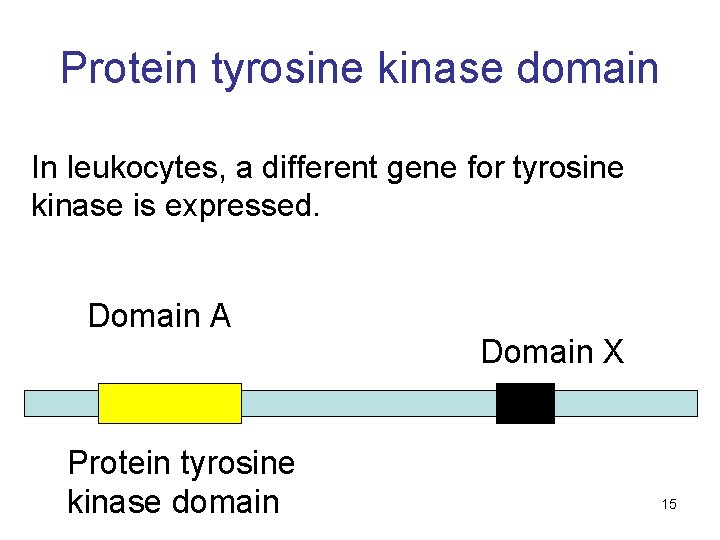
Protein tyrosine kinase domain In leukocytes, a different gene for tyrosine kinase is expressed. Domain A Protein tyrosine kinase domain Domain X 15
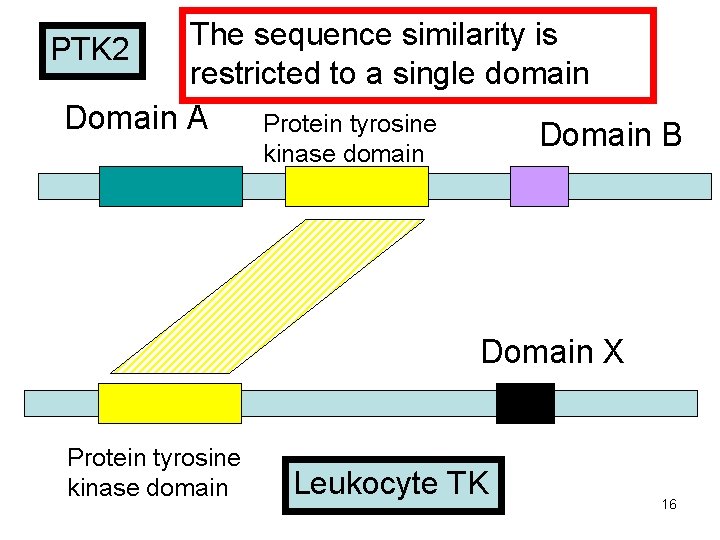
The sequence similarity is restricted to a single domain Domain A Protein tyrosine Domain B PTK 2 kinase domain Domain X Protein tyrosine kinase domain Leukocyte TK 16

Global alignment of PTK and LTK 17

Local alignment of PTK and LTK 18

Conclusions Use global alignment when the two sequences share the same overall sequence arrangement. Use local alignment to detect regions of similarity. 19

How alignments are computed 20

Pairwise alignment AAGCTGAATTCGAA AGGCTCATTTCTGA One possible alignment: AAGCTGAATT-C-GAA AGGCT-CATTTCTGA- 21

AAGCTGAATT-C-GAA AGGCT-CATTTCTGA- This alignment includes: 2 mismatches 4 indels (gap) 10 perfect matches 22
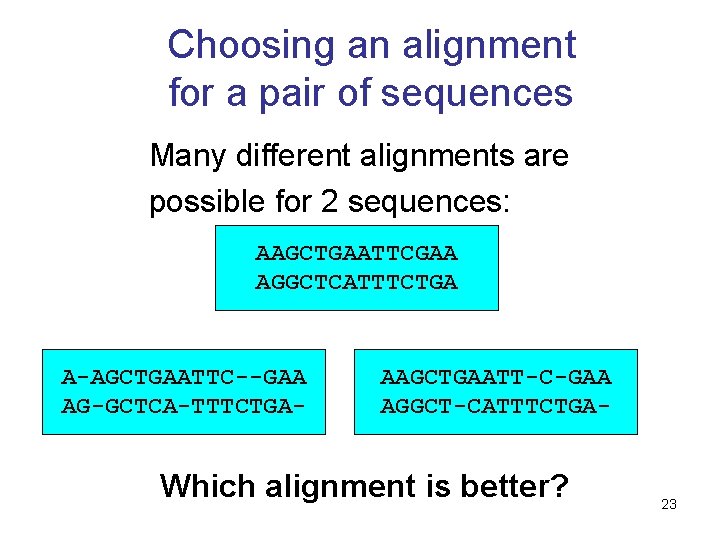
Choosing an alignment for a pair of sequences Many different alignments are possible for 2 sequences: AAGCTGAATTCGAA AGGCTCATTTCTGA A-AGCTGAATTC--GAA AG-GCTCA-TTTCTGA- AAGCTGAATT-C-GAA AGGCT-CATTTCTGA- Which alignment is better? 23
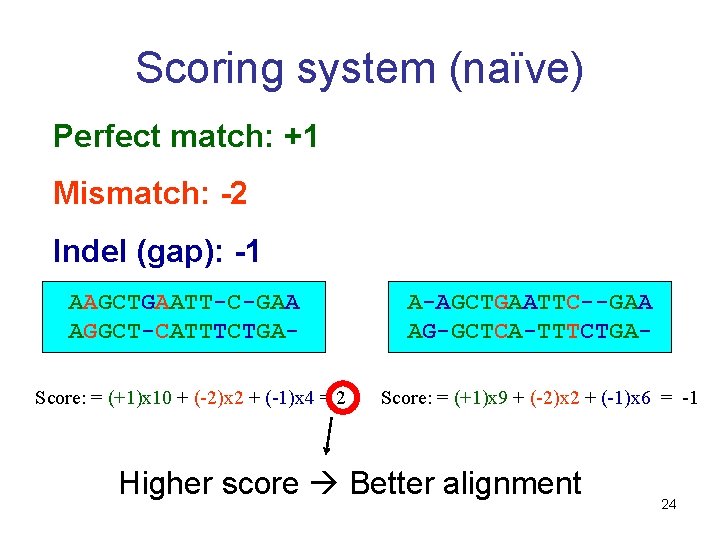
Scoring system (naïve) Perfect match: +1 Mismatch: -2 Indel (gap): -1 AAGCTGAATT-C-GAA AGGCT-CATTTCTGAScore: = (+1)x 10 + (-2)x 2 + (-1)x 4 = 2 A-AGCTGAATTC--GAA AG-GCTCA-TTTCTGAScore: = (+1)x 9 + (-2)x 2 + (-1)x 6 = -1 Higher score Better alignment 24

Alignment scoring - scoring of sequence similarity: Assumes independence between positions: each position is considered separately Scores each position: • Positive if identical (match) • Negative if different (mismatch or gap) Total score = sum of position scores Can be positive or negative 25

Scoring system • In the example above, the choice of +1 for match, -2 for mismatch, and -1 for gap is quite arbitrary • Different scoring systems different alignments • We want a good scoring system… 26
DNA scoring matrices Can take into account biological phenomena such as: • Transition-transversion 27
Amino-acid scoring matrices • Take into account physico-chemical properties 28

Scoring gaps (I) In advanced algorithms, two gaps of one amino-acid are given a different score than one gap of two amino acids. This is solved by giving a penalty to each gap that is opened. Gap extension penalty < Gap opening penalty 29

Homology versus chance similarity How to check if the score is significant? A. Take the two sequences Compute score. B. Take one sequence randomly shuffle it -> find score with the second sequence. Repeat 100, 000 times. If the score in A is at the top 5% of the scores in B the similarity is significant. 30

How close? • Rule of thumb: • Proteins are homologous if they are at least 25% identical (length >100) • DNA sequences are homologous if they are at least 70% identical 31

Twilight zone • < 25% identity in proteins – may be homologous and may not be…. • (Note that 5% identity will be obtained completely by chance!) 32

Searching a sequence database Idea: In order to find homologous sequences to a sequence of interest, one should compute its pairwise alignment against all known sequences in a database, and detect the best scoring significant homologs The same idea in short: Use your sequence as a query to find homologous sequences in a sequence database 33

Some terminology • Query sequence - the sequence with which we are searching • Hit – a sequence found in the database, suspected as homologous 34

Query sequence: DNA or protein? • For coding sequences, we can use the DNA sequence or the protein sequence to search for similar sequences. • Which is preferable? 35

Protein is better! • Selection (and hence conservation) works (mostly) at the protein level: CTTTCA = TTGAGT = Leu-Ser 36

Query type • Nucleotides: a four letter alphabet • Amino acids: a twenty letter alphabet • Two random DNA sequences will, on average, have 25% identity • Two random protein sequences will, on average, have 5% identity 37

Conclusion The amino-acid sequence is often preferable for homology search 38

How do we search a database? • If each pairwise alignment takes 1/10 of a second, and if the database contains 107 sequences, it will take 106 seconds = 11. 5 days to complete one search. • 150, 000 searches (at least!!) are performed per day. >82, 000 sequence records in Gen. Bank. 39

Conclusion • Using the exact comparison pairwise alignment algorithm between the query and all DB entries – too slow 40

Heuristic • Definition: a heuristic is a design to solve a problem that does not provide an exact solution (but is not too bad) but reduces the time complexity of the exact solution 41

BLAST 42

BLAST • BLAST - Basic Local Alignment and Search Tool • A heuristic for searching a database for similar sequences 43
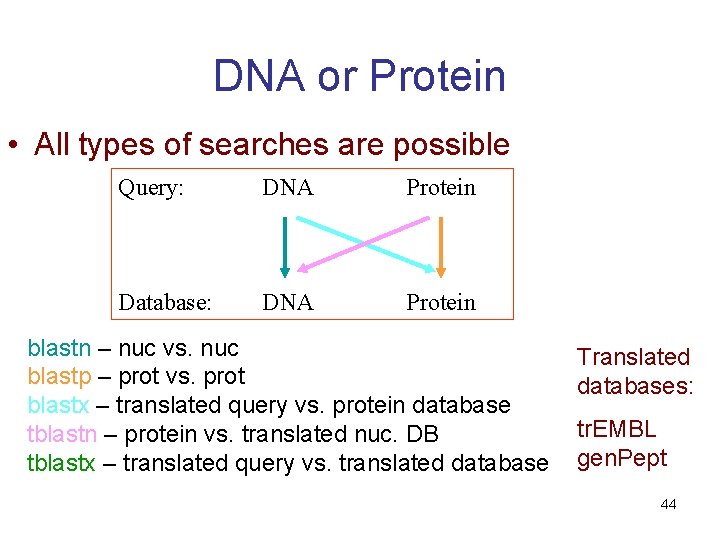
DNA or Protein • All types of searches are possible Query: DNA Protein Database: DNA Protein blastn – nuc vs. nuc blastp – prot vs. prot blastx – translated query vs. protein database tblastn – protein vs. translated nuc. DB tblastx – translated query vs. translated database Translated databases: tr. EMBL gen. Pept 44

BLAST - underlying hypothesis • The underlying hypothesis: when two sequences are similar there are short ungapped regions of high similarity between them • The heuristic: 1. Discard irrelevant sequences 2. Perform exact local alignment only with the remaining sequences 45

How do we discard irrelevant sequences quickly? • Divide the database into words of length w (default: w = 3 for protein and w = 7 for DNA) • Save the words in a look-up table that can be searched quickly WTDFGYPAILKGGTAC WTD TDF DFG FGY GYP … 46

BLAST: discarding sequences • When the user enters a query sequence, it is also divided into words • Search the database for consecutive neighboring words 47
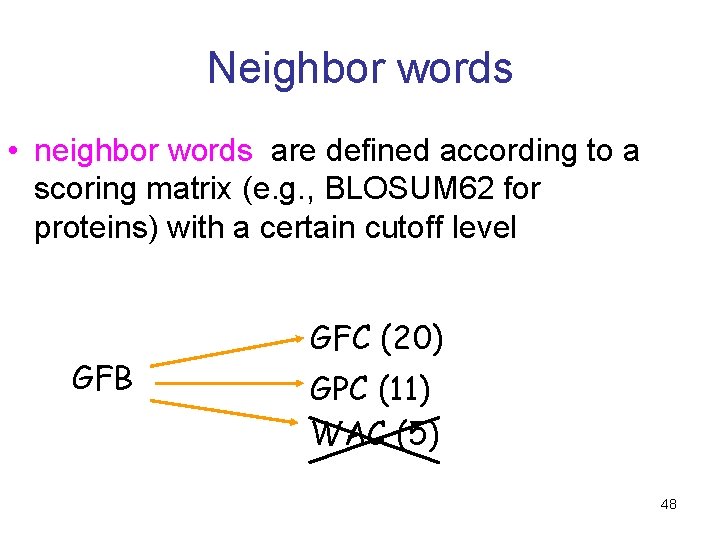
Neighbor words • neighbor words are defined according to a scoring matrix (e. g. , BLOSUM 62 for proteins) with a certain cutoff level GFB GFC (20) GPC (11) WAC (5) 48

E-value • The number of times we will theoretically find an alignment with a score ≥ Y of a random sequence vs. a random database Theoretically, we could trust any result with an E-value ≤ 1 In practice – BLAST uses estimations. E-values of 10 -4 and lower indicate a significant homology. E-values between 10 -4 and 10 -2 should be checked (similar domains, maybe non-homologous). E-values between 10 -2 and 1 do not indicate a good homology 49

Web servers for pairwise alignment

BLAST 2 sequences (bl 2 Seq) at NCBI Produces the local alignment of two given sequences using BLAST (Basic Local Alignment Search Tool) engine for local alignment • Does not use an exact algorithm but a heuristic
Back to NCBI
BLAST – bl 2 seq
Bl 2 Seq - query blastn – nucleotide blastp – protein

Bl 2 seq results

Bl 2 seq results Match Gaps Similarity Dissimilarity Low complexity
BLAST – programs Query: DNA Protein Database: DNA Protein
BLAST – Blastp
Blastp - results
Blastp – results (cont’)

Blast scores: • Bits score – A score for the alignment according to the number of similarities, identities, etc. • Expected-score (E-value) –The number of alignments with the same score one can “expect” to see by chance when searching a random database of a particular size. The closer the evalue is to zero, the greater the confidence that the hit is really a homolog

Blastp – acquiring sequences
blastp – acquiring sequences

Multiple sequence alignment VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGTSSNIGS--ITVNWYQQLPG LRLSCSSSGFIFSS--YAMYWVRQAPG LSLTCTVSGTSFDD--YYSTWVRQPPG PEVTCVVVDVSHEDPQVKFNWYVDG-ATLVCLISDFYPGA--VTVAWKADS-AALGCLVKDYFPEP--VTVSWNSG--VSLTCLVKGFYPSD--IAVEWWSNG-- Similar to pairwise alignment BUT n sequences are aligned instead of just 2 64

Multiple sequence alignment VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGTSSNIGS--ITVNWYQQLPG LRLSCSSSGFIFSS--YAMYWVRQAPG LSLTCTVSGTSFDD--YYSTWVRQPPG PEVTCVVVDVSHEDPQVKFNWYVDG-ATLVCLISDFYPGA--VTVAWKADS-AALGCLVKDYFPEP--VTVSWNSG--VSLTCLVKGFYPSD--IAVEWWSNG-- MSA = Multiple Sequence Alignment Each row represents an individual sequence Each column represents the ‘same’ position 65

Conserved positions • Columns in which all the sequences contain the same amino acids or nucleotides • Important for the function or structure VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGSSSNIGS--ITVNWYQQLPG LRLSCTGSGFIFSS--YAMYWYQQAPG LSLTCTGSGTSFDD-QYYSTWYQQPPG 66
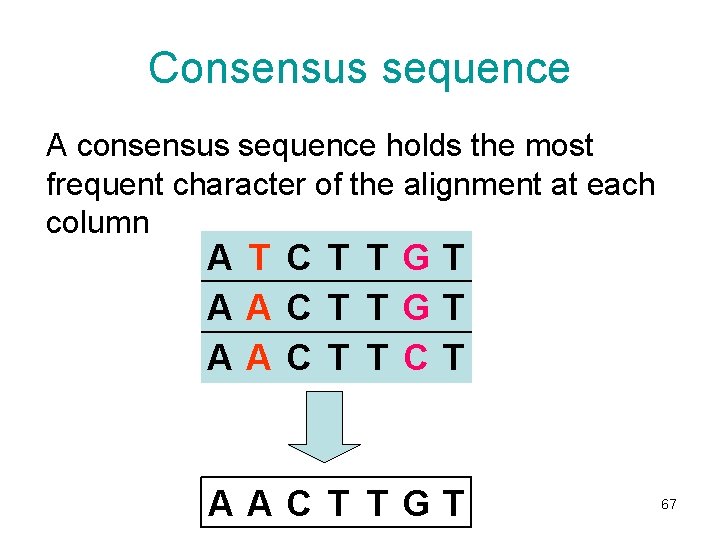
Consensus sequence A consensus sequence holds the most frequent character of the alignment at each column A T C T TGT AAC T T CT AAC T TGT 67

Profile = PSSM = Position Specific Score Matrix A T C T TG AAC T T C 1 2 A 1 . 67 00 C 0 0 G 0 0000 T 0 3 1 . 33 0 4 5 6 0 0 00 0. 33 0. 67 1 1 0 68

Alignment methods There is no available optimal solution for MSA – all methods are heuristics: • Progressive/hierarchical alignment (Clustal) • Iterative alignment (mafft, muscle) 69
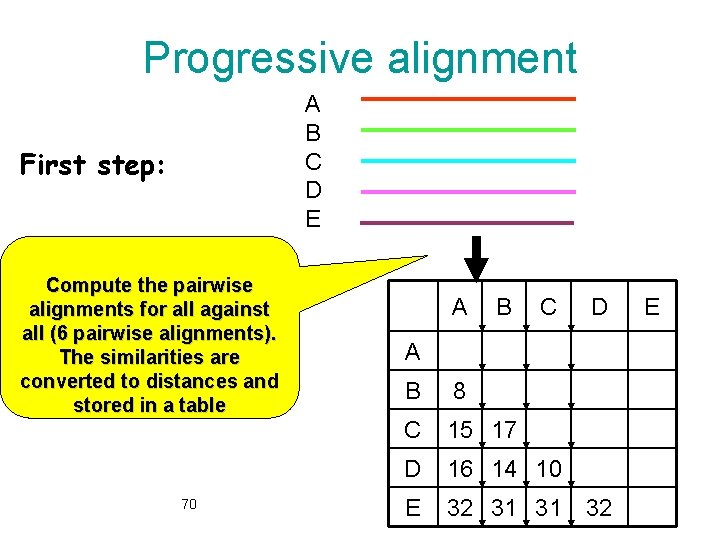
Progressive alignment A B C D E First step: Compute the pairwise alignments for all against all (6 pairwise alignments). The similarities are converted to distances and stored in a table 70 A B C D A B 8 C 15 17 D 16 14 10 E 32 31 31 32 E
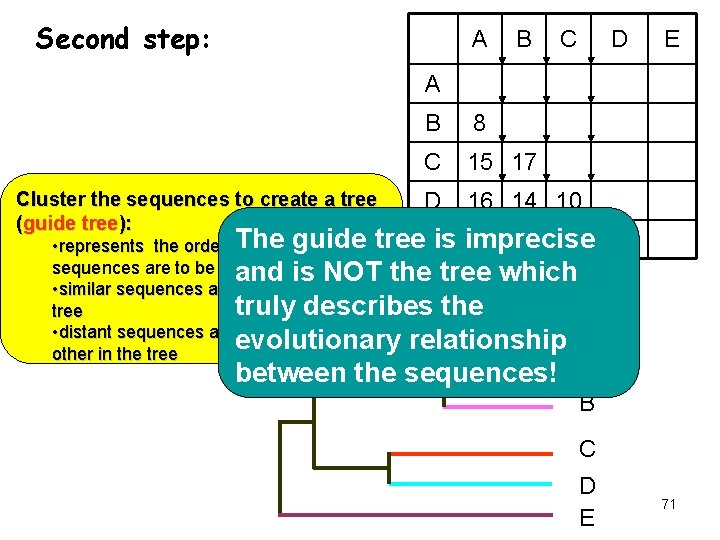
Second step: A B C D E A Cluster the sequences to create a tree (guide tree): B 8 C 15 17 D 16 14 10 The 32 31 31 • represents the order in whichguide pairs of tree. Eis imprecise sequences are to be aligned and is NOT the tree which • similar sequences are neighbors in the truly describes the tree • distant sequences are distant from each evolutionary relationship A other in the tree 32 between the sequences! B C D E 71
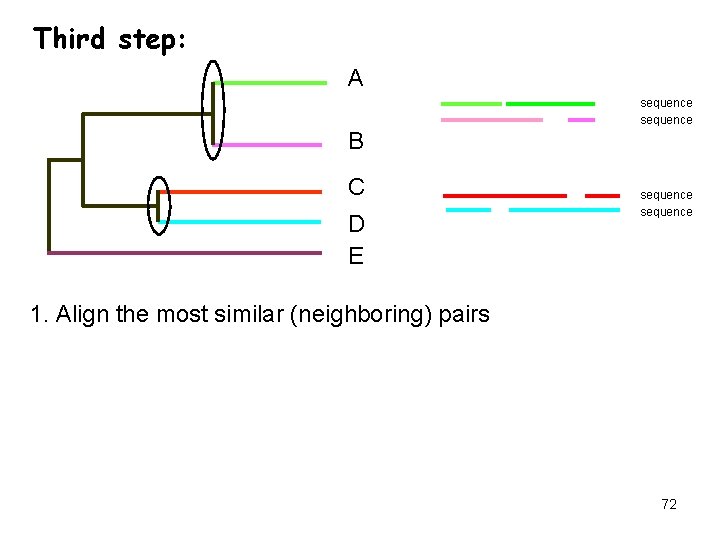
Third step: A sequence B C D E sequence 1. Align the most similar (neighboring) pairs 72
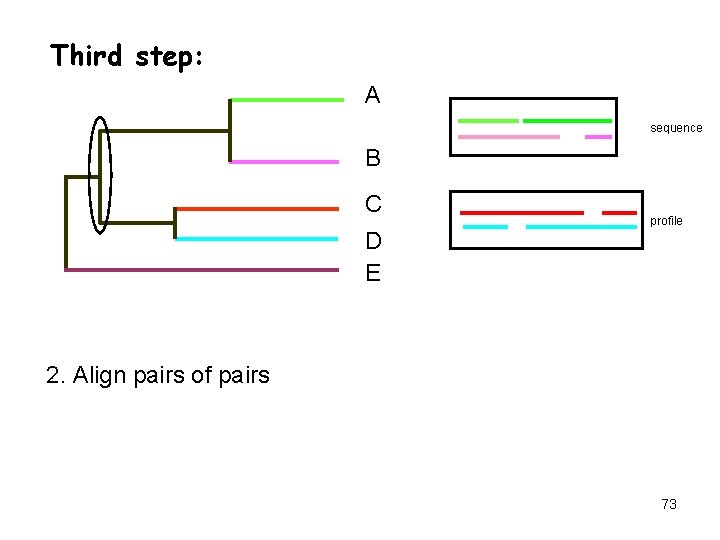
Third step: A sequence B C D E profile 2. Align pairs of pairs 73
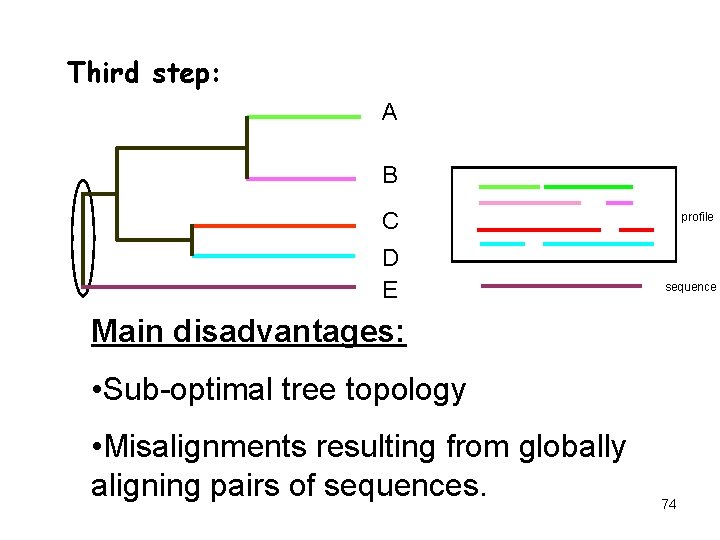
Third step: A B C D E profile sequence Main disadvantages: • Sub-optimal tree topology • Misalignments resulting from globally aligning pairs of sequences. 74
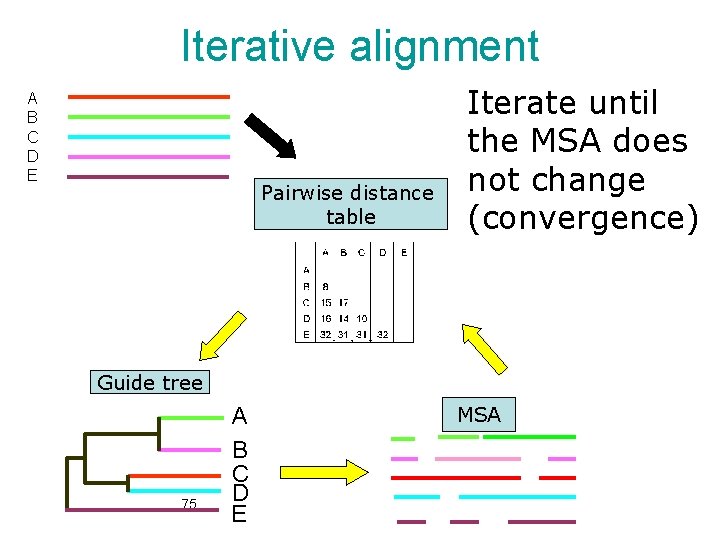
Iterative alignment A B C D E Pairwise distance table Iterate until the MSA does not change (convergence) Guide tree 75 A B C D E MSA

Case study: Using homology searching • The human kinome 76
Kinases and phosphatases 77

Multi-tasking enzymes • • • Signal transduction Metabolism Transcription Cell-cycle Differentiation Function of nervous and immune system • … • And more 78

How many kinases in the human genome? • 1950’s, discovery that reversible phosphorylation regulates the activity of glycogen phosphorylase • 1970’s, advent of cloning and sequencing produced a speculation that the vertebrate genome encodes as many as 1, 001 kinases 79

How many kinases in the human genome? • 2001 – human genome sequence … • As well – databases of Genbank, Swissprot, and db. EST • How can we find out how many kinases are out there? 80

The human kinome • In 2002, Manning, Whyte, Martinez, Hunter and Sudarsanam set out to: 1. Search and cross-reference all these databases for all kinases 2. Characterize all found kinases 81
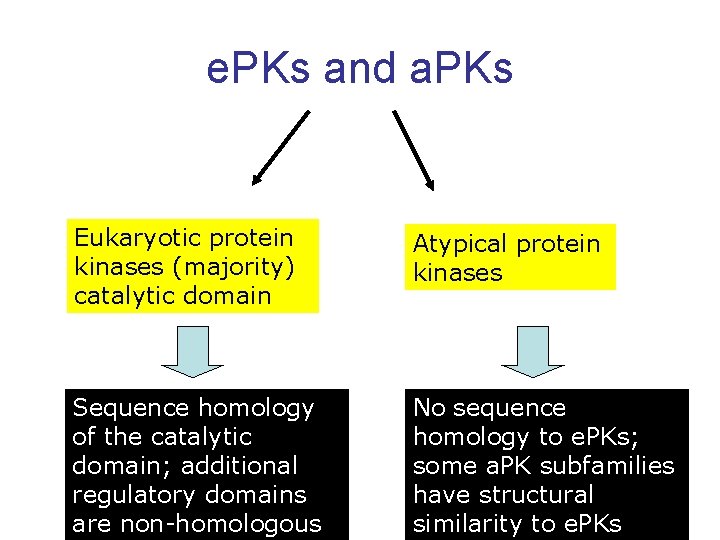
e. PKs and a. PKs Eukaryotic protein kinases (majority) catalytic domain Atypical protein kinases Sequence homology of the catalytic domain; additional regulatory domains are non-homologous No sequence homology to e. PKs; some a. PK subfamilies have structural 82 similarity to e. PKs

The search • Several profiles were built: based on the catalytic domain of: (a) 70 known e. PKs from yeast, worm, fly, and human with > 50% identity in the e. PK domain (b) each subfamily of known a. PKs • HMM-profile searches and PSI-BLAST searches were performed 83
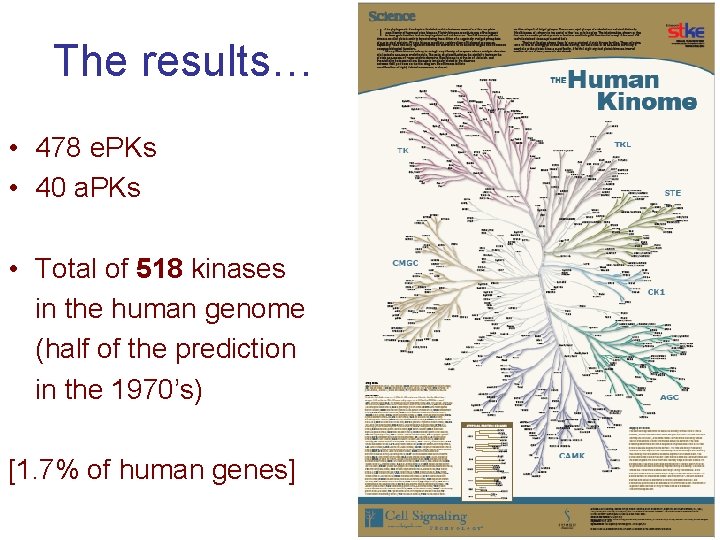
The results… • 478 e. PKs • 40 a. PKs • Total of 518 kinases in the human genome (half of the prediction in the 1970’s) [1. 7% of human genes] 84
- Slides: 84