Homologous recombination deficiency HRD of high grade serous
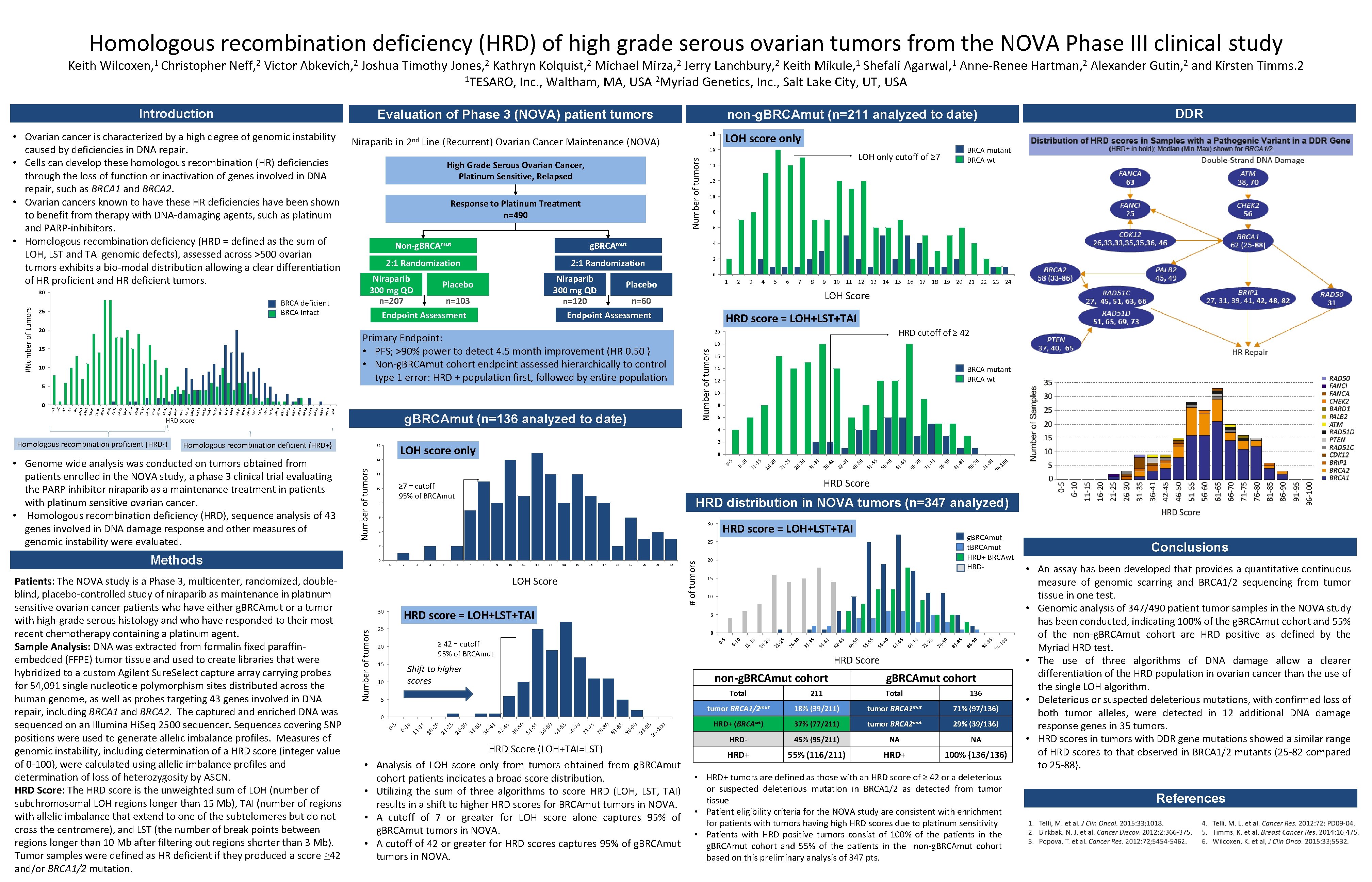
Homologous recombination deficiency (HRD) of high grade serous ovarian tumors from the NOVA Phase III clinical study Keith Wilcoxen, 1 Christopher Neff, 2 Victor Abkevich, 2 Joshua Timothy Jones, 2 Kathryn Kolquist, 2 Michael Mirza, 2 Jerry Lanchbury, 2 Keith Mikule, 1 Shefali Agarwal, 1 Anne-Renee Hartman, 2 Alexander Gutin, 2 and Kirsten Timms. 2 1 TESARO, Inc. , Waltham, MA, USA 2 Myriad Genetics, Inc. , Salt Lake City, UT, USA Introduction Evaluation of Phase 3 (NOVA) patient tumors Niraparib in 2 nd Line (Recurrent) Ovarian Cancer Maintenance (NOVA) Response to Platinum Treatment n=490 2: 1 Randomization Niraparib Placebo 300 mg QD n=103 n=207 Endpoint Assessment BRCA deficient BRCA intact 20 10 5 100 98 -99 96 -97 94 -95 92 -93 90 -91 88 -89 86 -87 82 -83 84 -85 80 -81 78 -79 76 -77 74 -75 72 -73 70 -71 68 -69 66 -67 64 -65 62 -63 60 -61 58 -89 56 -57 54 -55 52 -53 50 -51 48 -49 46 -47 44 -45 42 -43 40 -41 38 -39 36 -37 32 -33 30 -31 28 -29 26 -27 24 -25 22 -23 20 -21 34 -35 HRD score 10 8 6 4 2: 1 Randomization 2 0 Niraparib Placebo 300 mg QD n=60 n=120 Endpoint Assessment 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 LOH Score HRD score = LOH+LST+TAI HRD cutoff of ≥ 42 20 18 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 18 -19 8 16 -17 7 10 -11 12 -13 6 14 -15 5 8 -9 4 6 -7 3 4 -5 2 2 -3 1 0 -1 0 12 g. BRCAmut Primary Endpoint: • PFS; >90% power to detect 4. 5 month improvement (HR 0. 50 ) • Non-g. BRCAmut cohort endpoint assessed hierarchically to control type 1 error: HRD + population first, followed by entire population 15 BRCA mutant BRCA wt LOH only cutoff of ≥ 7 14 Number of tumors #Number of tumors 30 25 16 High Grade Serous Ovarian Cancer, Platinum Sensitive, Relapsed Non-g. BRCAmut LOH score only 18 Number of tumors • Ovarian cancer is characterized by a high degree of genomic instability caused by deficiencies in DNA repair. • Cells can develop these homologous recombination (HR) deficiencies through the loss of function or inactivation of genes involved in DNA repair, such as BRCA 1 and BRCA 2. • Ovarian cancers known to have these HR deficiencies have been shown to benefit from therapy with DNA-damaging agents, such as platinum and PARP-inhibitors. • Homologous recombination deficiency (HRD = defined as the sum of LOH, LST and TAI genomic defects), assessed across >500 ovarian tumors exhibits a bio-modal distribution allowing a clear differentiation of HR proficient and HR deficient tumors. DDR non-g. BRCAmut (n=211 analyzed to date) g. BRCAmut (n=136 analyzed to date) 16 BRCA mutant BRCA wt 14 12 10 8 6 4 30 4 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 HRD score = LOH+LST+TAI # of tumors 2 0 -1 0 96 91 -9 5 -9 0 86 81 -8 5 -8 0 76 71 -7 5 -7 0 66 -6 5 61 -6 0 -5 5 56 46 51 -4 5 42 36 -4 1 -3 5 31 26 -3 0 -2 5 21 -2 0 -5 0 g. BRCAmut t. BRCAmut HRD+ BRCAwt HRD- 20 15 10 5 25 15 0 -1 0 96 -9 5 91 -9 0 86 -8 5 81 -8 0 76 -7 5 71 -7 0 66 -6 5 61 -6 0 56 -5 5 51 -5 0 46 -4 5 42 -4 1 36 -3 5 31 -3 0 26 -2 5 21 -2 0 16 11 -1 5 0 -1 0 96 -9 5 91 -9 0 86 -8 5 81 -8 0 76 71 -7 5 -7 0 66 -6 5 61 -6 0 56 -5 5 51 -5 0 46 -4 5 42 -4 1 36 -3 5 31 -3 0 26 -2 5 21 -2 0 16 -1 5 non-g. BRCAmut cohort 5 0 - 11 HRD Score Shift to higher scores 10 6 - 5 ≥ 42 = cutoff 95% of BRCAmut 20 10 0 0 - Number of tumors HRD score = LOH+LST+TAI 25 30 • 16 6 LOH Score • -1 5 HRD distribution in NOVA tumors (n=347 analyzed) 8 1 • 11 HRD Score ≥ 7 = cutoff 95% of BRCAmut 10 0 • 10 5 0 - 12 2 Methods Patients: The NOVA study is a Phase 3, multicenter, randomized, doubleblind, placebo-controlled study of niraparib as maintenance in platinum sensitive ovarian cancer patients who have either g. BRCAmut or a tumor with high-grade serous histology and who have responded to their most recent chemotherapy containing a platinum agent. Sample Analysis: DNA was extracted from formalin fixed paraffinembedded (FFPE) tumor tissue and used to create libraries that were hybridized to a custom Agilent Sure. Select capture array carrying probes for 54, 091 single nucleotide polymorphism sites distributed across the human genome, as well as probes targeting 43 genes involved in DNA repair, including BRCA 1 and BRCA 2. The captured and enriched DNA was sequenced on an Illumina Hi. Seq 2500 sequencer. Sequences covering SNP positions were used to generate allelic imbalance profiles. Measures of genomic instability, including determination of a HRD score (integer value of 0 -100), were calculated using allelic imbalance profiles and determination of loss of heterozygosity by ASCN. HRD Score: The HRD score is the unweighted sum of LOH (number of subchromosomal LOH regions longer than 15 Mb), TAI (number of regions with allelic imbalance that extend to one of the subtelomeres but do not cross the centromere), and LST (the number of break points between regions longer than 10 Mb after filtering out regions shorter than 3 Mb). Tumor samples were defined as HR deficient if they produced a score ≥ 42 and/or BRCA 1/2 mutation. 0 14 610 • Genome wide analysis was conducted on tumors obtained from patients enrolled in the NOVA study, a phase 3 clinical trial evaluating the PARP inhibitor niraparib as a maintenance treatment in patients with platinum sensitive ovarian cancer. • Homologous recombination deficiency (HRD), sequence analysis of 43 genes involved in DNA damage response and other measures of genomic instability were evaluated. 2 LOH score only 16 6 - Homologous recombination deficient (HRD+) Number of tumors Homologous recombination proficient (HRD-) HRD Score (LOH+TAI=LST) Analysis of LOH score only from tumors obtained from g. BRCAmut cohort patients indicates a broad score distribution. Utilizing the sum of three algorithms to score HRD (LOH, LST, TAI) results in a shift to higher HRD scores for BRCAmut tumors in NOVA. A cutoff of 7 or greater for LOH score alone captures 95% of g. BRCAmut tumors in NOVA. A cutoff of 42 or greater for HRD scores captures 95% of g. BRCAmut tumors in NOVA. g. BRCAmut cohort Total 211 Total 136 tumor BRCA 1/2 mut 18% (39/211) tumor BRCA 1 mut 71% (97/136) HRD+ (BRCAwt) 37% (77/211) tumor BRCA 2 mut 29% (39/136) HRD- 45% (95/211) NA NA HRD+ 55% (116/211) HRD+ 100% (136/136) • HRD+ tumors are defined as those with an HRD score of ≥ 42 or a deleterious or suspected deleterious mutation in BRCA 1/2 as detected from tumor tissue • Patient eligibility criteria for the NOVA study are consistent with enrichment for patients with tumors having high HRD scores due to platinum sensitivity • Patients with HRD positive tumors consist of 100% of the patients in the g. BRCAmut cohort and 55% of the patients in the non-g. BRCAmut cohort based on this preliminary analysis of 347 pts. Conclusions • An assay has been developed that provides a quantitative continuous measure of genomic scarring and BRCA 1/2 sequencing from tumor tissue in one test. • Genomic analysis of 347/490 patient tumor samples in the NOVA study has been conducted, indicating 100% of the g. BRCAmut cohort and 55% of the non-g. BRCAmut cohort are HRD positive as defined by the Myriad HRD test. • The use of three algorithms of DNA damage allow a clearer differentiation of the HRD population in ovarian cancer than the use of the single LOH algorithm. • Deleterious or suspected deleterious mutations, with confirmed loss of both tumor alleles, were detected in 12 additional DNA damage response genes in 35 tumors. • HRD scores in tumors with DDR gene mutations showed a similar range of HRD scores to that observed in BRCA 1/2 mutants (25 -82 compared to 25 -88). References
- Slides: 1