Homologous recombination 5 3 3 5 5 3
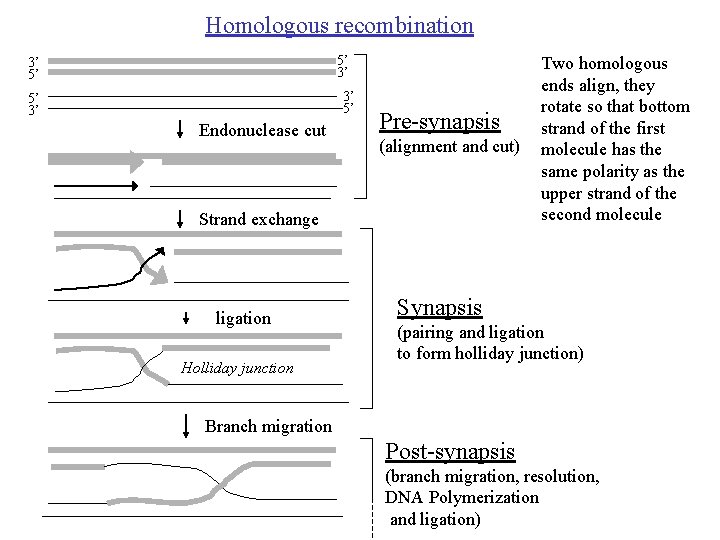
Homologous recombination 5’ 3’ 3’ 5’ 5’ 3’ Endonuclease cut Pre-synapsis (alignment and cut) Strand exchange ligation Holliday junction Two homologous ends align, they rotate so that bottom strand of the first molecule has the same polarity as the upper strand of the second molecule Synapsis (pairing and ligation to form holliday junction) Branch migration Post-synapsis (branch migration, resolution, DNA Polymerization and ligation)
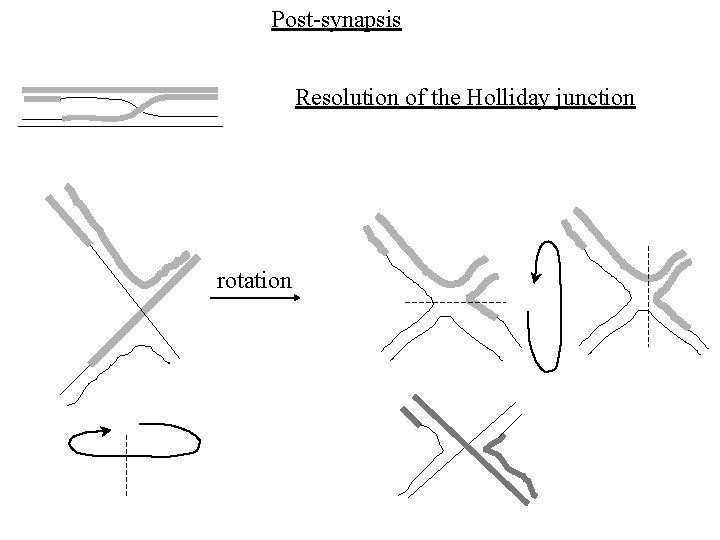
Post-synapsis Resolution of the Holliday junction rotation
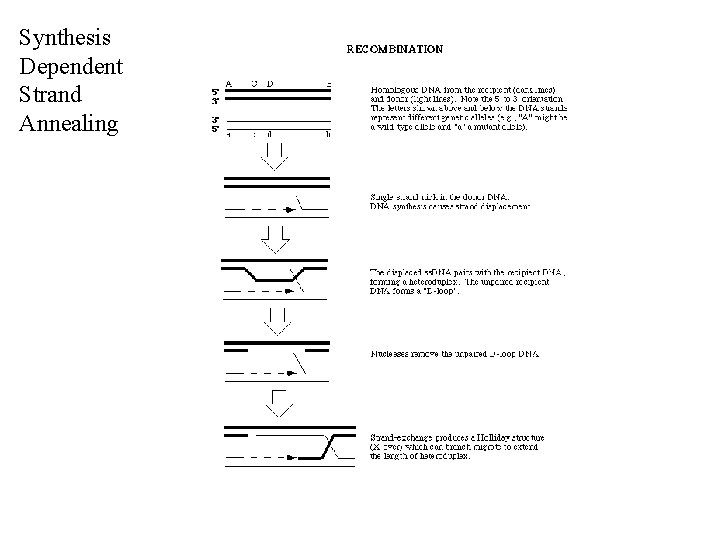
Synthesis Dependent Strand Annealing
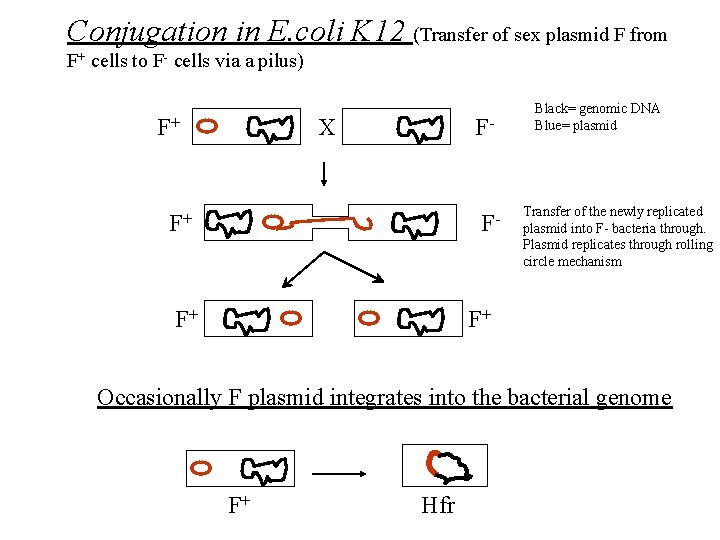
Conjugation in E. coli K 12 (Transfer of sex plasmid F from F+ cells to F- cells via a pilus) F+ F- X F+ F- F+ Black= genomic DNA Blue= plasmid Transfer of the newly replicated plasmid into F- bacteria through. Plasmid replicates through rolling circle mechanism F+ Occasionally F plasmid integrates into the bacterial genome F+ Hfr
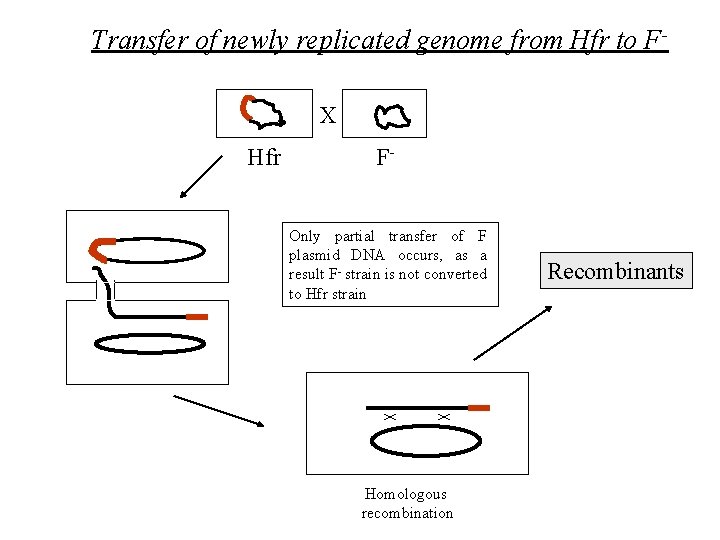
Transfer of newly replicated genome from Hfr to FX Hfr FOnly partial transfer of F plasmid DNA occurs, as a result F- strain is not converted to Hfr strain Homologous recombination Recombinants

Hfr x F-: a system for isolating genes involved in homologous recombination. Types of mutant screens A. Loss of function (Hypo-Rec mutants): 1. Reduced ability of F minus strain to produce recombinants when mated with Hfr lawn 2. Sensitivity to mitomycin C (a DNA damaging drug) 3. Sensitivity to UV B. Gain of function (suppressor mutations): mutations that increase the recombination proficiency of otherwise hypo-rec mutant.

rec. A (involved in synapsis) Rec. A protein requires ATP for binding to ss. DNA rapidly and cooperatively to generate a nucleo-protein filament (Rec. A bound ss. DNA). The nucleo-protein filament binds non-specifically to ds. DNA to generate DNA network. In these networks, ss. DNA synapses with homologous ds. DNA sequence. Thus, Rec. A promotes pairing and strand exchange. rec. B, rec. C, rec. D (pre-synapsis) Rec. B, Rec. C and Rec. D proteins are components of Rec. BCD enzyme, which is also called as Exo. V. Rec. BCD has an ATP dependent ds. DNA and ss. DNA exonuclease activity. It has an ATP-dependent unwinding activity on linear ds. DNA producing ss. DNA tails or long ss. DNA. During unwinding from right to left makes ss nicks near chi (crossover hotspot instigators) site (5’-GCTGGTGG-3’). It also contains D -loop cleavage activity.

rec. E (pre-synapsis) Also called as Exo. VIII. Has ATP-independent exonuclease activity. Degrades ds. DNA from 5’ end, producing long 3’ tails. It normally stays suppressed by the action of sbc. A protein. rec. E gene is activated in rec. BCD, sbc. A mutants. Thus Rec. E-mediated recombination is a secondary pathway. rec. F (pre-synapsis) Catalytic activity of Rec. F is unknown. It probably acts as a switch to divert ss. DNA from being replication templates to being a recombination substrates.

Post-synapsis genes for Homologous Recombination Ruv. AB - These enzymes catalyze branch migration, resulting in pairing of two homologous strands of DNA. Ruv. AB has DNA helicase activity that causes branch migration from a crossover in the direction that will extend the DNA heteroduplex. Ruv. C - These enzymes are Holliday junction specific endonucleases that cut two strands of DNA at a "cross-over" (Holliday junction). The resulting DNA strands can be rejoined by DNA ligase, completing the recombination reaction. Ruv. C interacts with Ruv. AB to form a Ruv. ABC complex that facilitates the branch migration and recombination reaction.

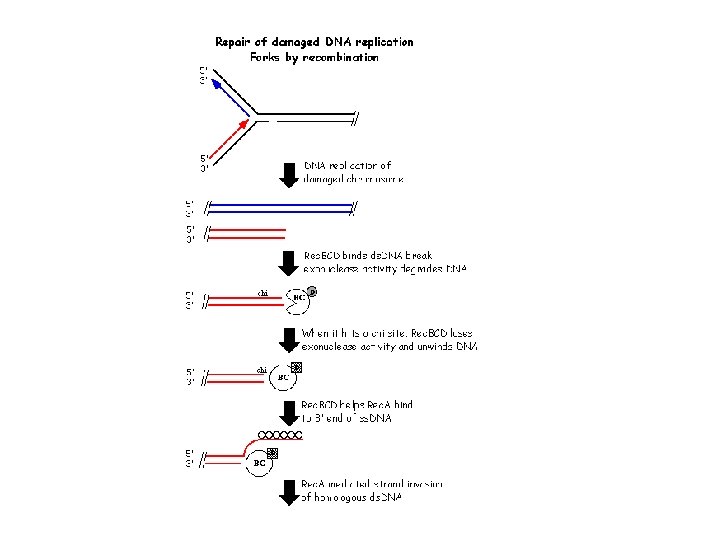
Pathways of homologous recombination 1. Rec. BCD pathway: predominant. 2. Rec. FOR pathway: secondary 3. Rec. E pathway ? (Differ in pre-synaptic steps. Have mechanistically identical synaptic and post-synaptic steps) Rec. BCD pathway The Rec. BCD complex initiates recombination between two homologous ds. DNA molecules when one of the molecules has a free, double-stranded end. The Rec. BCD complex binds to the free end and rapidly degrades the DNA. When Rec. BCD encounters a short DNA sequence called a chi site, the Rec. D subunit is altered such that the double-stranded exonuclease is inactivated -- the Rec. BC complex is converted to a helicase with 5' to 3' single-stranded nuclease activity that unwinds the donor DNA and degrades one strand of the DNA. In addition, the Rec. B subunit helps "load" Rec. A protein onto the 3' end of the resulting ss. DNA. The ss. DNA-Rec. A complex can invade a homologous ds. DNA molecule. (There are other enzymes (Rec. FOR pathway) in bacteria that can catalyze similar functions, so rec. BC mutants are not completely defective for recombination).

Rec. FOR pathway - The Rec. FOR pathway was discovered by looking for suppressors of rec. BC mutants (called sbc. BCD). Rec. F binds DNA and has a weak ATPase activity that is stimulated by Rec. R. Rec. O promotes the exchange of single-stranded DNA binding protein (SSB) with Rec. A protein.
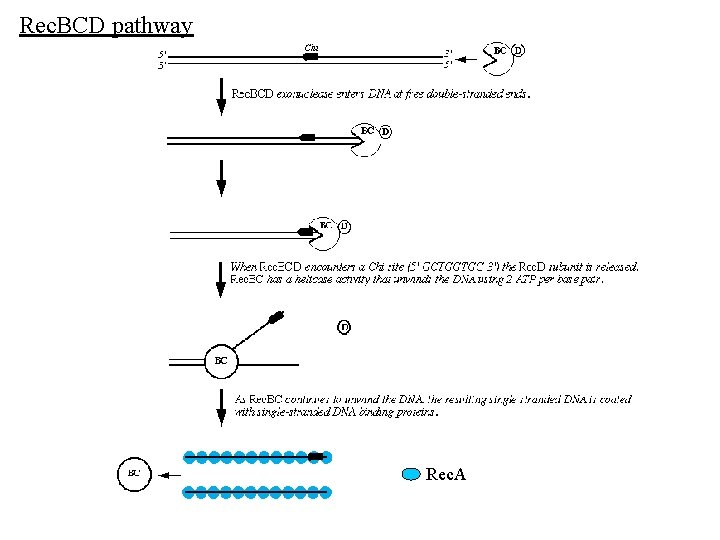
Rec. BCD pathway Rec. A

Relevance to crop genetic engineering 1. Gene integration via homologous recombination (HR) between the introduced DNA and the host chromosome will produce the most desirable transgenic product. However, HR-mediated foreign gene integration does not occur efficiently in higher plants. One of the burning topics in plant biology is to develop a strategy for obtaining HRmediated transgene integration. 2. Basic features of HR mechanism and the essential proteins (eg. Rec. A) are conserved between the kingdoms. rec. A-like genes are found in yeast, animals and plants. rec. A-like genes of yeast are called Rad 51 and Dmc 1. Plant homologs of dmc 1 have been identified. E. coli genetics (conjugation) Yeast genetics (mitosis/meiosis) Plant / Animal genetics (mitosis/meiosis)

Yeast proteins involved in homologous recombination (Mitosis/ Meiosis) Genetic screen= sensitivity of yeast cells to X-rays Rec. A counterpart Rad 51, Rad 52, Rad 54, Rad 55, and Rad 57: These proteins probably complex together to form a recombinosome. Meiosis specific Rad 51 is called DMC 1. Rad 51 is a Rec. A homolog. Thus, Rad 51 in the recombinosome is involved in ATP-dependent strand exchange Rec. BCD, F, E counterparts Rad 50, Mre 11, Xrs 2: Make a complex that is involved in the structural maintenance of recombination substrate and processing of DNA ends by a 5’— 3’ exonulease activity. Rad 50 and Mre 11 are also implicated to have roles in the illegitimate recombination. Mre 11 mutants of chicken show increased homologous recombination frequency (probably due to the suppression of illegitimate recombination)

Strategies for obtaining efficient HR in plants 1. Over-expression of E. coli genes involved in HR: [Bernd Reiss et al. PNAS | March 28, 2000 | vol. 97 | no. 7 | 3358 -3363 ] Over-expression of Rec. A: stimulated sister chromatid exchanges, increased intra-chromosomal recombination but not the frequency of gene targeting (GT is defined as recombination between foreign gene carrying homologous sequences and the complementary genomic sequences). Over-expression of Ruv. C (Ruv. C is involved in the resolution of Holliday junction): [Shalev et al. PNAS Vol. 96, Issue 13, 7398 -7402 1999] Increased intra-, extra-, and inter-chromosomal recombination but not GT frequency. 2. Modification of plant HR machinery: RAD 50 mutant: RAD 50 may be a common factor of illegitimate recombination and HR. RAD 50 is dispensable for HR. RAD 50 plant mutant may have suppressed illegitimate recombination. Intrachromosomal recombination indeed goes up in RAD 50 plant mutant. GT frequency is yet to be demonstrated in RAD 50 mutants. 3. Create ds breaks (DSB) in the target site: This is the most successful strategy. It is based on the finding that a ds break (DSB) in plant genome is preferentially repaired by HR process. This led to development of strategies for creating site-specific DSB, such as Zinc-Finger Nuclease-mediated double-stranded break induction.
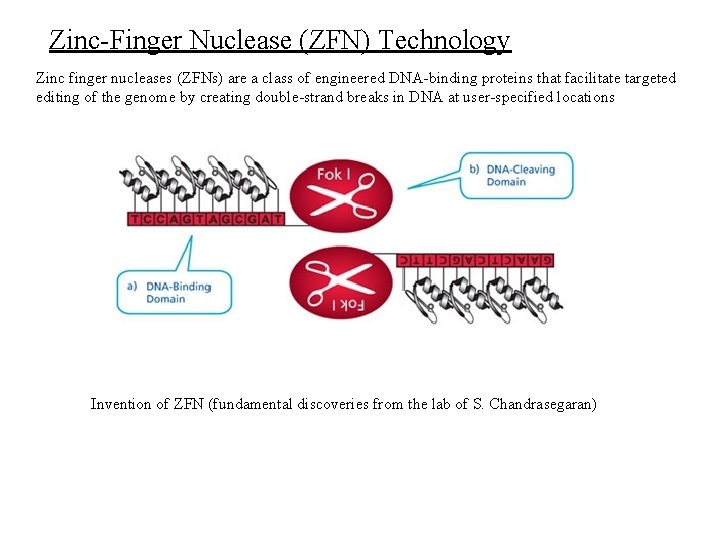
Zinc-Finger Nuclease (ZFN) Technology Zinc finger nucleases (ZFNs) are a class of engineered DNA-binding proteins that facilitate targeted editing of the genome by creating double-strand breaks in DNA at user-specified locations Invention of ZFN (fundamental discoveries from the lab of S. Chandrasegaran)

Srinivasan Chandrasegaran and Jeff Smith (1999) Biol. Chem. , Vol. 380, pp. 841 – 848. Minireview: Chimeric Restriction Enzymes: What Is Next? There are numerous bacterial enzymes that recognize an asymmetric sequence and cleave a short distance from that sequence. These are termed type IIS enzymes, where ‘s’ stands for shifted cleavage. These enzymes do not recognize any specific sequence at the site cut. For example, Fok. I restriction endonuclease recognizes the non-palindromic penta-deoxyribonucleotide 5’-GGATG-3’: 5’-CATCC-3’ in duplex DNA and cleaves 9/13 nucleotides downstream of the recognition site. This property implies the presence of two separate protein domains within Fok. I: one for sequence-specific recognition of DNA and the other for the endonuclease activity. Once the DNA-binding domain is anchored at the recognition site, a signal is transmitted to the endonuclease domain, probably, through allosteric interactions, and cleavage occurs. We reasoned that the type IIS enzymes probably are the ideal candidates for changing sequence specificities because one may be able to swap the recognition domain of these enzymes with other naturally occurring DNA-binding proteins which recognize longer sequences. Therefore, we undertook a detailed study of the Fok. I restriction-modification system from Flavobacterium okeanokoites R= recognition or DNA binding domain EN= endonuclease domain

Srinivasan Chandrasegaran and Jeff Smith (1999) Biol. Chem. , Vol. 380, pp. 841 – 848. Minireview: Chimeric Restriction Enzymes: What Is Next? The modular nature of Fok. I endonuclease suggested that it might be feasible to construct chimeric restriction enzymes with novel sequence-specificities by linking other DNA-binding proteins to the cleavage domain of Fok. I endonuclease. This indeed proved to be the case. We reported the construction of the first ‘chimeric’ restriction endonuclease by linking the Drosophila Ubx homeodomain to the cleavage domain of Fok. I (Kim and Chandrasegaran, 1994). We then reported the creation of novel sitespecific endonucleases by linking two different three zinc-finger proteins to the Fok. I cleavage domain (Kim et al. , 1996). Recently, we reported the creation of a novel site-specific endonuclease by linking the N-terminal 147 amino acids of the yeast Gal 4 protein to the cleavage domain of Fok. I (Kim et al. , 1998). Thus, we have shown that the three common eukaryotic DNA-binding motifs, namely the helix-turn-helix motif, the zinc finger motif and the basic helix-loop-helix protein containing a leucine zipper motif, can be converted into novel site-specific endonucleases by fusing them to the Fok. I cleavage domain. Such engineered chimeric nucleases have been shown to make specific cuts in vitro very close to the expected recognition sequences. Of these chimeric nucleases, the most important are those based on zinc-finger DNA-binding motifs. Because of their modular nature, the zinc finger proteins offer an attractive framework for designing chimeric restriction enzymes with tailor-made sequence-specificities. The Cys 2 -His 2 zinc finger proteins are a class of DNA-binding proteins that contain sequences of the form (Tyr, Phe)-Xaa-Cys-Xaa 2 -4 -Cys. Xaa 3 -Phe-Xaa 5 -Leu-Xaa 2 -His-Xaa 3 -5 -His, usually in tandem arrays. Xaa represents an unspecified amino acid. Each of these sequences binds a Zinc(II) ion to form the structural domain termed a zinc finger. These proteins bind to DNA by inserting an α-helix into the major groove of the double helix.
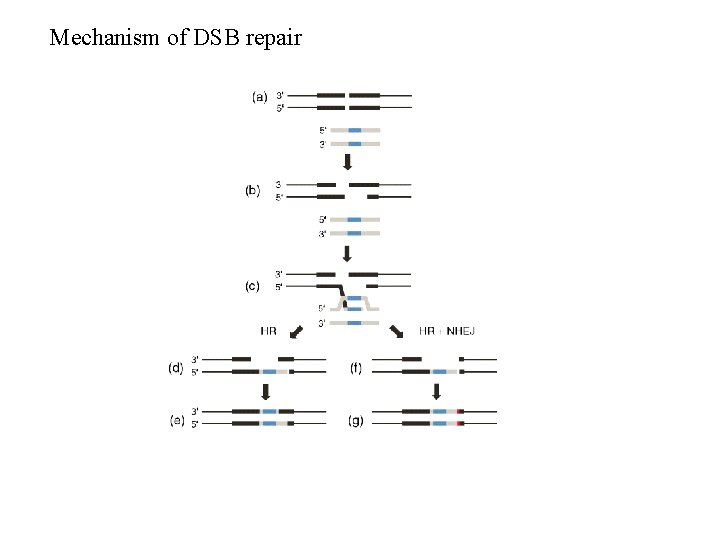
Mechanism of DSB repair
- Slides: 20