HL 7 Clinical Genomics Implementation Roadmap The HL




- Slides: 8

HL 7 Clinical Genomics – Implementation Roadmap The HL 7 Clinical Genomics SIG Amnon Shabo (Shvo), Ph. D HL 7 Clinical Genomics SIG Co-chair and Modeling Facilitator HL 7 Structured Documents TC CDA R 2 Co-editor CCD Implementation Guide Co-editor

Haifa Research Lab V 2 : ||: V 3 Challenges § What should be the position of the HL 7 community towards independent modeling attempts of v 2 messages applied to new domains that have been solely modeled in v 3? § What’s the long term vision? § Gradual conversion to v 3 until v 2 is deprecated? Or § V 2/V 3 Co-existent? If so - how? § Divergence is the current situation: § V 3 and V 2 ‘languages’ have different semantics § V 2 - V 3 ‘attribute mapping’ does NOT help when semantics at the source level is different, e. g. , different structures, dynamics, etc. 2

Haifa Research Lab Proposed Approach § Recognize V 3 models as a single source of semantics § Refer to V 2 messages is a partial v 3 implementation § Automatically generate v 2 messages from v 3 models, much like the V 3 XML ITS § Acknowledge the incompleteness of the v 2 implementation, but § § Address specific requirements of use cases Let the operator of the v 3 v 2 generator make domain choices Document the gaps Imply missing semantics from the v 3 models 3

Haifa Research Lab A Proposed Roadmap for CG Implementations The Clinical Genomics semantics: represented by normative HL 7 V 3 Models V 2 message Implementation Technologies: generation V 3 XML ITS CDISC SDTM Genetic testing Family History Pharmacogemomics Implementation Balloted informative implementation specifications 4

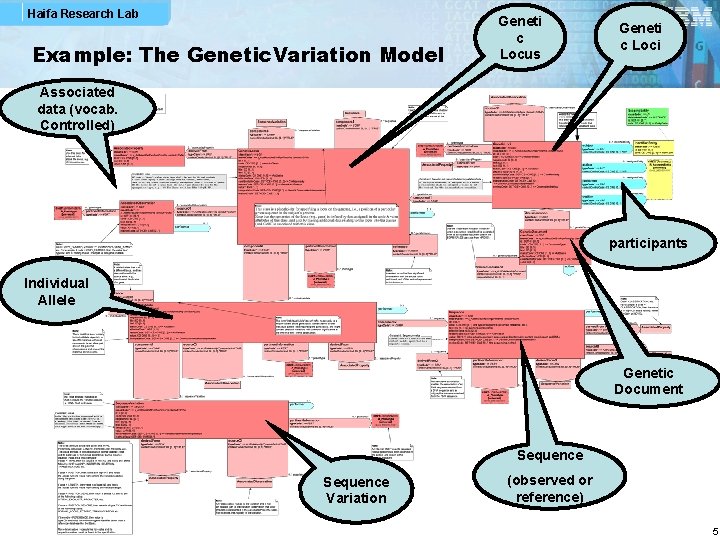
Haifa Research Lab Example: The Genetic. Variation Model Geneti c Locus Geneti c Loci Associated data (vocab. Controlled) participants Individual Allele Genetic Document Sequence Variation (observed or reference) 5

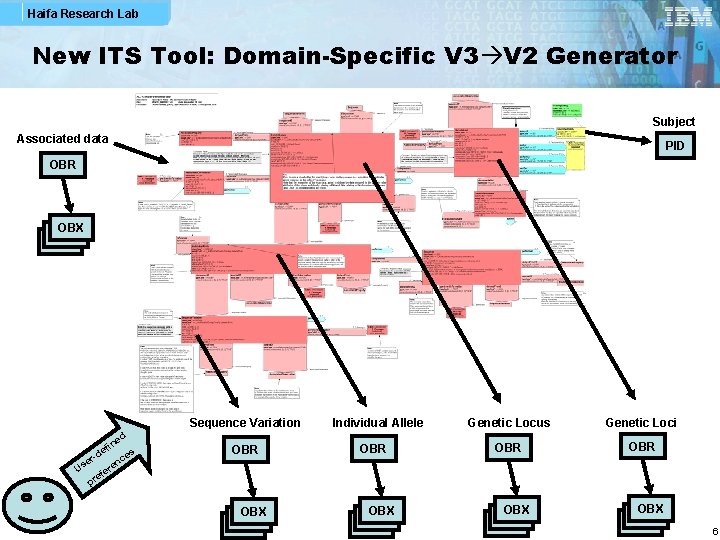
Haifa Research Lab New ITS Tool: Domain-Specific V 3 V 2 Generator Subject Associated data PID OBR OBX OBX Sequence Variation ed fin e s r-d nce e Us fere e pr Individual Allele Genetic Locus Genetic Loci OBR OBR OBX OBX OBX 6

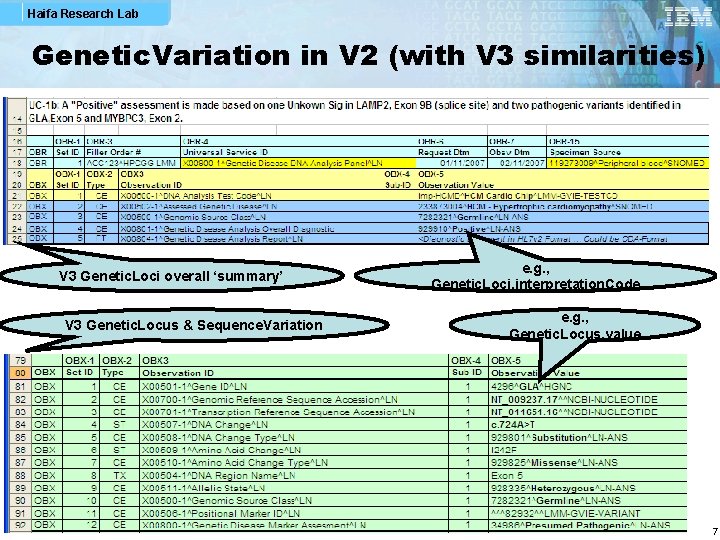
Haifa Research Lab Genetic. Variation in V 2 (with V 3 similarities) V 3 Genetic. Loci overall ‘summary’ V 3 Genetic. Locus & Sequence. Variation e. g. , Genetic. Loci. interpretation. Code e. g. , Genetic. Locus. value 7

Haifa Research Lab What Should the V 3 V 2 ITS Tool Facilitate? § Consistently capture v 3 structural codes & clone names in v 2 fields (is it doable? ) § The choice of v 2 ‘common patterns’, e. g. : § Patterns of implementing v 3 nesting clones (Sub-ID? Parent id? ) § Document common gaps like v 3 associations § Transform v 3 XML schemas to v 2 XML-encoded and on to v 2 ASCII 8