HIV RT INHIBITION invitro in cell culture Materials

HIV RT INHIBITION (in-vitro , / in cell culture) Materials and methods TITAS MALLICK © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 1

Recap of previous presentation • Presentation 1 Basic HIV biology, AIDS, the real life scenario, global statistics and importance of new drug development and an overview of HIV biology; i. e. the infection and replication of the virus. • AIDS A DEADLY DISEASE AIDS (acquired immunodeficiency syndrome) is a syndrome caused by a virus called HIV (human immunodeficiency virus). The disease alters the immune system, making people much more vulnerable to infections and diseases. This susceptibility worsens if the syndrome progresses. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 2

Cont. 2017 GLOBAL HIV STATISTICS • 36. 9 million [31. 1 million– 43. 9 million] people globally were living with HIV in 2017. • 21. 7 million [19. 1 million– 22. 6 million] million people were accessing antiretroviral therapy in 2017. • 1. 8 million [1. 4 million– 2. 4 million] people became newly infected with HIV in 2017. • 940 000 [670 000– 1. 3 million] people died from AIDS-related illnesses in 2017. • 77. 3 million [59. 9 million– 100 million] people have become infected with HIV since the start of the epidemic. • 35. 4 million [25. 0 million– 49. 9 million] people have died from AIDS-related illnesses since the start of the epidemic. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 3

Cont. • Presentation 2 History of HIV drug development, its drawback and importance of new drug development. LUC MONTAGNIER, FRANCOISE BARRE-SINOUSSI’s effort to identify the causal organism of AIDS, previously called as GRID And finally ROBERT GALLO’s isolation of the virus. Early treatment by Suramin and AZT(azidothymidine) as HIV inhibitor. Their drawback; cytotoxicity and other and introduction of modern era of treatment. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 4

Cont. • HAART, and PREP 1. 2 nucleoside reverse transcriptase inhibitors (NRTI) 2. 1 non-nucleoside reverse transcriptase inhibitor (NNRTI) 3. 2 protease inhibitor (PI) 4. 1 Integrase nuclear strand transfer inhibitors (INSTI) • 21 st century approaches like CRISPER, BONE MARROW TRANSPLANT and VACCINATION. Their problems, cost-effectiveness, and the main need for drug development. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 5

The basic need for new drug research • The HIV replication is a very error prone process by nature, the DNA dependent RNA polymerase incorporates wrong bases during replication, most of them though causes lethal mutations so the resultant virus doesn’t mature, but some of them end up altering the RT protein structure (or other protein) that doesn’t make the enzyme non functional but alters the site of drug binding. So the drug doesn’t work for long, that makes it necessary to create new drugs that can effectively attack the new and mutated proteins of the virus. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 6

1. 2. 3. 4. 5. | Plant sources may be potent source for anti-retroviral drugs | Low cytotoxicity is expected. Expected to be much efficient. Low cost expected. Can be major component of HAART. Already reported plants crude extract having anti-retroviral effect. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 7

Objectives of the project. OBJECTIVE 1 Extraction and purification of HIV RT The HIV RT gene is cloned in a BL 21(DE 3) Vector in E. coli strain PET 28 A fused with Hexa Histidine tag under a IPTG inducible promoter. Secondary culture, induction, and nickel affinity chromatography can be used to successfully isolate and purify HIV RT. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 8

• PET 28 RT 66 cultured for the RT 66 isolation. • Primary culture overnight. • Secondary culture from the primary culture done. • Induced by IPTG. • The cells are harvested as pellet after centrifugation at 6 K for 6’’ from the overnight grown secondary culture. • The pellet are washed with a pellet wash buffer. (COMP: 0. 2 M Na. Cl and 10 m. M Tris) • Sonicated with lysis buffer (Comp: Binding Buffer + 0. 05% NP 40 + Lysozyme + PMSF) © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 9

Cont. • Initially the IMAC column recharged with Ni. So 4. • Ni-IMAC done to extract the His-tagged RT 66 • First the binding done with the binding buffer (comp: 50 m. M Tris PH 8. 0, 500 m. M Na. Cl, 5 m. M Imidazole) • Washing buffer (comp: 20 m. M Tris PH 7. 5, 500 m. M Na. Cl, 75 m. M imidazole) used to wash off other proteins. • Bradford test used to check the protein conc in washes. • Elution buffer (comp: 20 m. M Tris PH 7. 5, 500 m. M Na. Cl, 500 m. M imidazole) used to elute and collect the protein. • The protein eluted in the previous stage put in 2 step dialysis (comp Dialysis Buffer 1 - 50 m. M Tris, 100 m. M Na. Cl, 30% glycerol, comp Dialysis Buffer 2 - 50 m. M Tris, 100 m. M Na. Cl, Glycerol 50%) © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 10

cont. • As same process the PBR 28 ART 51 cultured and protein purification done for RT 51 • I quantified the RT 51 by Bradford standard curve. • The RT 51 I quantified have a conc of 3. 70 ug/ml • THE RT 51 and RT 66 will be mixed equimolarly to get functional RT. • RT 66 shown functionality in in-vitro transcription itself. • To check the proteins in SDS-PAGE Negative stating of the gel is performed for quick visualization © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 11
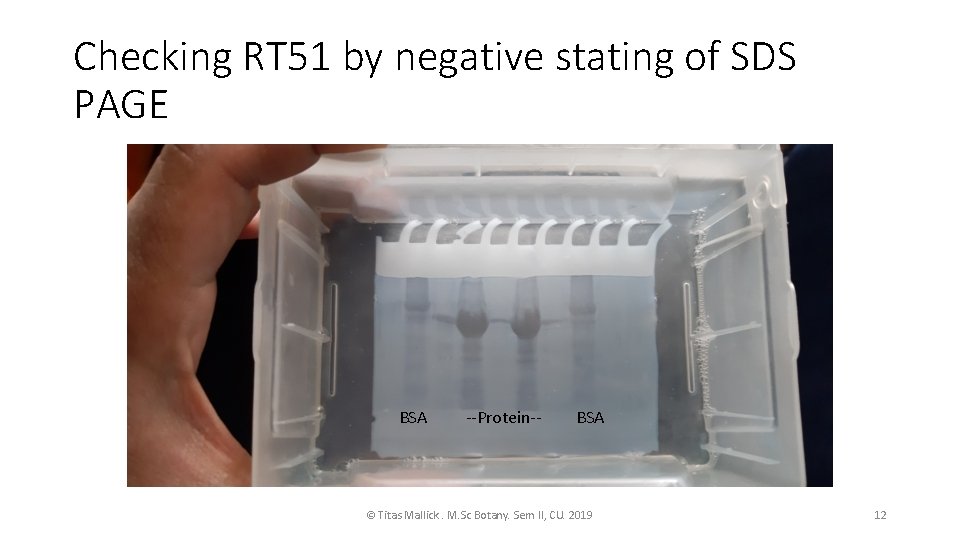
Checking RT 51 by negative stating of SDS PAGE BSA --Protein-- BSA © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 12

Procedure of negative staining of SDS PAGE • After the gel electrophoresis the gel is placed on a clear tray in mili. Q water. • The gel is hydrated for 2 min. • Decant the water. 0. 2 M Imidazole with 0. 1% SDS in mili. Q added, and the gel is placed on a rocker for 15 min. • 0. 2 M Zn. SO 4 addedon the gel. • Within 2 min banding pattern appears. • Before staining with Coomassie the gel is washed multiple times with mili. Q till it turn transparent. • Normal staining and distaining is performed. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 13
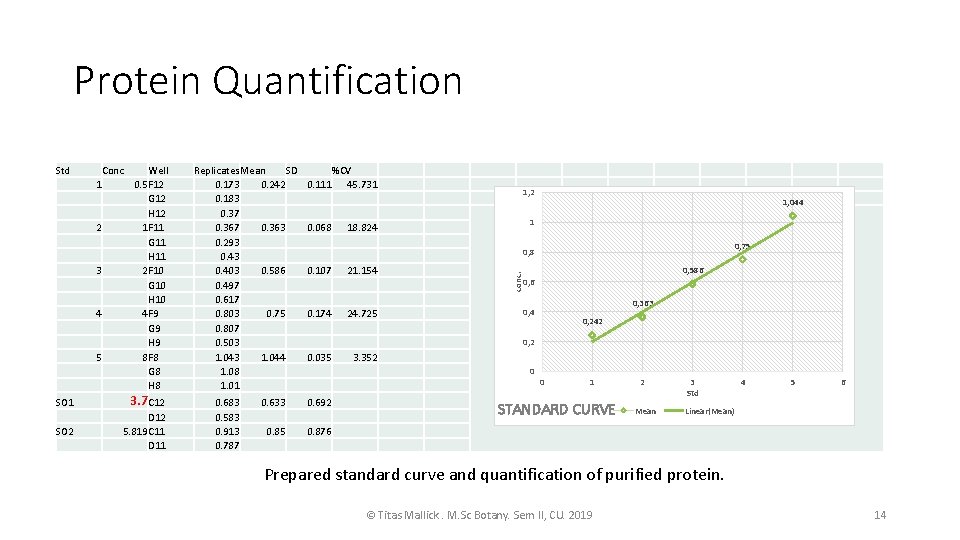
Protein Quantification Conc 1 2 3 4 5 Well 0. 5 F 12 G 12 H 12 1 F 11 G 11 H 11 2 F 10 G 10 H 10 4 F 9 G 9 H 9 8 F 8 G 8 H 8 SO 1 3. 7 C 12 SO 2 D 12 5. 819 C 11 D 11 Replicates Mean SD 0. 173 0. 242 0. 183 0. 37 0. 363 0. 293 0. 403 0. 586 0. 497 0. 617 0. 803 0. 75 0. 807 0. 503 1. 044 1. 08 1. 01 0. 683 0. 583 0. 913 0. 787 %CV 0. 111 45. 731 0. 068 1, 2 1, 044 1 18. 824 0, 75 0, 8 0. 107 21. 154 0. 174 24. 725 conc. Std 0, 586 0, 363 0, 4 0, 242 0, 2 0. 035 3. 352 0 0. 633 0. 692 0. 85 0. 876 0 1 STANDARD CURVE 2 Mean 3 Std 4 5 6 Linear(Mean) Prepared standard curve and quantification of purified protein. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 14

OBJECTIVE 2 Isolation and purification of plant crude extract Choice of plant for the project is used to prepare the crude extract. Column extraction processes are used to for the isolation of concentrated crude protein from the plant. Protein fractionations are made in different grade of polar to nopolar solvents. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 15

Procedure of Plant crude extraction. A D B F E A. Selection of Plant, B. Collection, C. Drying and grinding, D. Column extraction, E. Rotary evaporation, F. Collection and. M. Sc drying of. Sem crude © Titas Mallick Botany. II, CU. extract. 2019 16 C

Isolation of plant crude extract • Catharanthus roseus and Ocimum gigatium chosen as choice of plant. The leaf and branches are dried and powder is made. The Powder loaded in the column. Three separate solvents 1. Petroleum Benzene 2. Ethyl Acetate 3. Methanol according to their polarity used for the extraction from the plant materials. The extracts are then dried and collected and weighed. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 17

OBJECTIVE 3 Assay of RT to check its’ functionality. High throughput Functional RT assay to be done to check the RT activity. For the project we should use the DNA that is in a RELA-GFP vector for other experiment to be cloned under a T 7 promoter in a second vector. Then in vitro transcription is used to check RT functionality RT PCR performed to check the RT functionality. Before RT PCR an in-vitro transcription is performed to generate RNAs. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 18

In vitro transcription • Template DNA, forward and reverse primer, ATP, CTP, GTP UTP and 10 x buffer used for this process. DNA dependent RNA polymerase is used as the enzyme to generate RNA. DEPC treated H 2 O is used in this process. • After the incubation the RNA is quantified and this RNA is used for the RT PCR. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 19

RT PCR • To check the RT activity this assay is performed. • The enzyme purified (RT 66) is used in this procedure. The first step of this assay include a Reverse transcription process, and the DNA developed in this process is amplified using the PCR and checked in the gel. Positive result indicate the RT is functional. • The first stage of this process is c. DNA synthesis, RNA from the in vitro transcription, a primer d. NTPs is used, RT 66 is used as the enzyme. And 5 x Buffer and water is used. • After that the c. DNA is amplified using PCR, d. NTPs, Mg. Cl 2, 10 x Buffer, 2 primers, water and enzyme is used. After the incubation the product is checked in gel. RT 66 HAS SHOWN ACTIVITY. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 20

OBJECTIVE 4 Identification of the functional RT inhibitor molecule. First the crude plant extracts are to be used to check their effectiveness against the HIV RT. And then the crudes are to be fractionated further and each fractions are needed to be screened for functional RT inhibition. The further fractionations to be done unless a single molecule could be identified as HIV RT inhibitor. High throughput RT assays needed to be done for the screening process. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 21

High throughput pico-green assay • Pico green assay was performed in the lab. But due to some issues related to the instruments it failed and instead the radioactive primer extension assay was started. • STATUS OF PICO GREEN ASSAY: STOPPED. • Procedure: first a optimization and a standard curve generation is crucial. Reaction is performed with 2800, 28, 2. 8, 0. 28, 0. 028 pico mole of DNA, TE buffer and Pico green dye (1: 2000). • Dye control, DNA control and RNA control with all the sets are prepared in duplicates. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 22

Cont. • After the completion of optimization and standard curve preparation pico green should be used in reaction with RT alongside control RT inhibitors to check different plant crudes for their RT inhibition activity. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 23

Radioactive primer extension assay to check RT activity • Poly RA is used as the RNA with oligo DT primer. HIV RT is used as the enzyme. • Radio labeled 3 Hd. TTP (Tritium T) added as nucleotides. • First optimization to be done with different amount of enzymes, RNA. • Once after the completion of the optimization process. Different crudes and fractions are tested for HIV RTi activity, and the reading is taken through a Saintillation counter. • STATUS: NOT STARTED. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 24

Materials and process • First Template Primer dimer is needed to be prepared in a ratio of 1: 5. It is then diluted 200 fold with polymerization buffe (includes Tris-Cl 120 m. M PH 8. 1, KCl 120 m. M, Mg. Cl 2 16 m. M, 0. 5 M DTT and H 2 O) and incubated in 37 degree C for 1 hr. • Then Reaction sets are prepared with Reaction Buffer (includes 50 m. M Tris CL p. H 7. 8, 5 m. M Mg. Cl 2, 60 m. M KCl, 100 ug/ml BSA, 1 m. M DTT, 0. 5 uci 3 Hd. TTPs), Poly RA-dt, RT, Water. • Incubated in 37 degree for 1, 3, 5… mins. The reaction is stopped with EDTA and incubated on ice for 30 mins. • Transferred on a filter paper. Washed with 1 ml 5% TCA. The filter paper washed with TCA, Water, 70% Ethanol x 2, dried. • 1 m. L saintilletion fluid added and reading taken in saintillation counter. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 25

OBJECTIVE 5 Checking the RT inhibitor in cellular condition. The molecule showing HIV RT inhibition activity is tested for it’s cytotoxicity level by a CC 50 test. Then the molecule is introduced to a cell culture system with single cycle HIV virus along with the RT inhibitor for testing it in cellular condition. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 26

Test of HIV inhibitor activity in cell culture • 293 T cells are co-transfected with PNL 43ΔGFP (RT backbone with Green fluroscent Protein) and VSVG coat. • After 48 hours of incubation the produced virus are separated and it is used to infect Huh cells. The infected cells are seen green in fluorescent microscopy. • After 24 hr the cells should be treated with Control HIV inhibitors and test Crude extracts and fractions to check anti HIV activity. • In case of positive result the healthy cell must not be green. • Status: This is expected to be started soon. © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 27

• Discussion © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 28

• THANK YOU © Titas Mallick. M. Sc Botany. Sem II, CU. 2019 29
- Slides: 29