HIV RT INHIBITION in vitro and Cell Inhibition
HIV RT INHIBITION in vitro and Cell Inhibition of retroviral revertase (EC 2. 7. 7. 49) by isolated molecules from Plants. TITAS MALLICK, UNIVERSITY OF CALCUTTA

INTRODUCTION • AIDS A DEADLY DISEASE AIDS (acquired immunodeficiency syndrome) is a syndrome caused by a virus called HIV (human immunodeficiency virus). The disease alters the immune system, making people much more vulnerable to infections and diseases. This susceptibility worsens if the syndrome progresses. TITAS MALLICK, UNIVERSITY OF CALCUTTA

HIV STATISTICS 2017 GLOBAL HIV STATISTICS • 36. 9 million [31. 1 million– 43. 9 million] people globally were living with HIV in 2017. • 21. 7 million [19. 1 million– 22. 6 million] million people were accessing antiretroviral therapy in 2017. • 1. 8 million [1. 4 million– 2. 4 million] people became newly infected with HIV in 2017. • 940 000 [670 000– 1. 3 million] people died from AIDS-related illnesses in 2017. • 77. 3 million [59. 9 million– 100 million] people have become infected with HIV since the start of the epidemic. • 35. 4 million [25. 0 million– 49. 9 million] people have died from AIDS-related illnesses since the start of the epidemic. TITAS MALLICK, UNIVERSITY OF CALCUTTA

• HIV VIRUS Virus classification Group: Group VI (ss. RNA-RT) Order: Unassigned Family: Retroviridae Subfamily: Orthoretrovirinae Genus: Lentivirus TITAS MALLICK, UNIVERSITY OF CALCUTTA
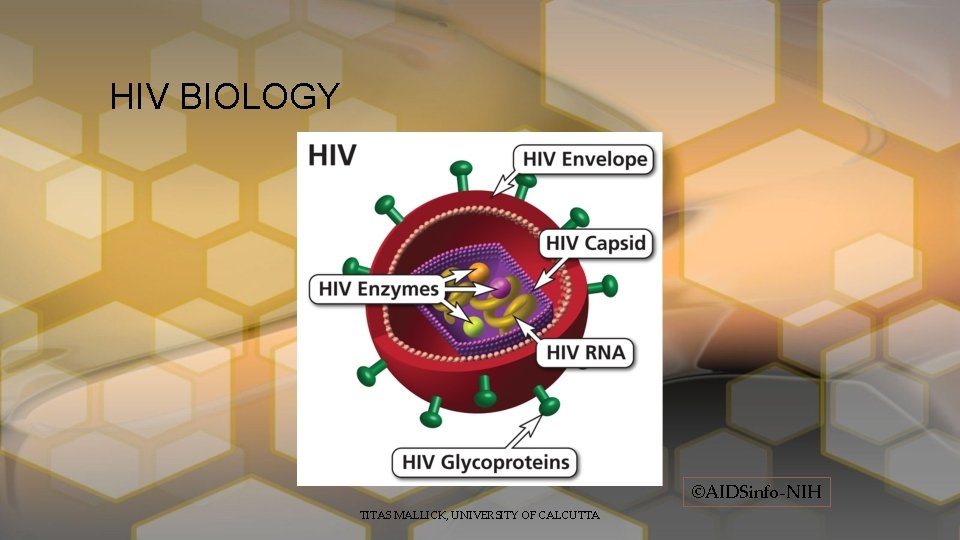
HIV BIOLOGY ©AIDSinfo-NIH TITAS MALLICK, UNIVERSITY OF CALCUTTA

Characters • Diameter 120 nm • 2 +ve sense strand RNA genome • 9 genes in total • 19 total proteins. (Gag, Pol, Env, Tat, Rev, Nef, Vif, Vpr, Vpu, Tev [10 th gene sometime present]) • P 24 coat protein • P 17 capsid protein TITAS MALLICK, UNIVERSITY OF CALCUTTA

HIV virology • Gag, Pol, Env is important for new virus production • Gag encodes for GP 120, which also get spliced into GP 41 • Tat encode p 16, P 14 transcriptase transactivator. • Tar element processed to mi. RNA and can regulate ERCC 1, IER 3 : important apoptotic gens • Rev bind pre. RNA element • Vif prevents APOBEC 3 G: a cellular protein that identifies poly C and poly U in viral tract and deactivate RT activity • Vpr induces cell cycle arrest at G 2/M checkpoint TITAS MALLICK, UNIVERSITY OF CALCUTTA

continue • Nef down regulate CD 4 as well as both Class I & II MHC • Vpu helps in packaging. • During packaging Psi elemnt must present in the 3’ end of the LTR • SLIP element helps in Gag-Pol frameshift during transcription. TITAS MALLICK, UNIVERSITY OF CALCUTTA

PROCESSES • GP 120 interact with CD 4, corecptor CCR 5 also plays vital role. (alternate of CD 4 -CCR 5 mechanism is Mannose specific C-type lectin mediated receptor DC-SIGV mediated infection) • GP 41 initiate initial injection of capsomeric proteins. • 2 -contact point stable attachment later established. • GP 120 interact with Integrin α 4β 7, initiate LFA-1 integrin mediated cell to cell spreading TITAS MALLICK, UNIVERSITY OF CALCUTTA
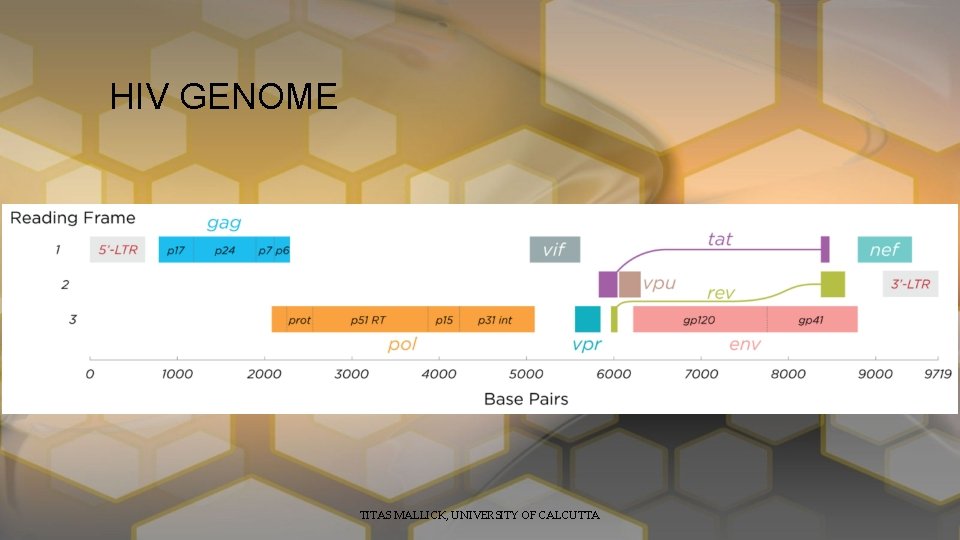
HIV GENOME TITAS MALLICK, UNIVERSITY OF CALCUTTA

continue • Viral RNA -> RT -> c. DNA -> Integration -> Transcription • Rev one among the first encoded protein have a NLS, first shuttle to the nucleus and with the help of Tat it helps the complete v. RNA +ve sense strand to shuttle to the cytoplasm • Gag, Env help to form the complete virus. General structure is like. LTR-U 3 -R-U 5 -LTR TITAS MALLICK, UNIVERSITY OF CALCUTTA

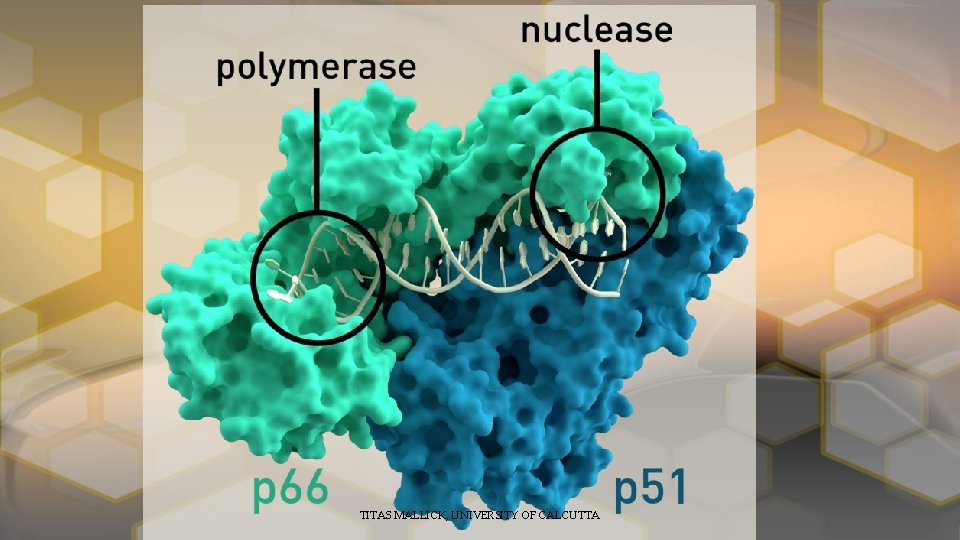
continue • RT developed from the Gag-Pol protein by protein splicing mechanism by the Cleavage by Viral Protease PR. • RT have two subunits p 66 and p 51, developed by a p 6 coding region frameshift read through. • P 66 is a 560 aminoacid protein • P 51 is a 440 aminoacid protein. TITAS MALLICK, UNIVERSITY OF CALCUTTA

HIV ENZYME SYSTEM • Reverse Transcriptase • Integrase • Protease TITAS MALLICK, UNIVERSITY OF CALCUTTA

Reverse Transcriptase TITAS MALLICK, UNIVERSITY OF CALCUTTA
TITAS MALLICK, UNIVERSITY OF CALCUTTA

Protein Seqence IV-1 (HXB 2): 10 20 30 40 50 60 70 80 90 100 | | | | | PISPIETVPV KLKPGMDGPK VKQWPLTEEK IKALVEICTE 110 120 130 140 150 160 170 180 190 200 | | | | | KKKKSVTVLD VGDAYFSVPL DEDFRKYTAF TIPSINNETP 210 220 230 240 250 260 270 280 290 300 | | | | | KIEELRQHLL RWGLTTPDKK HQKEPPFLWM GYELHPDKWT 310 320 330 340 350 360 370 380 390 400 | | | | | LELAENREIL KEPVHGVYYD PSKDLIAEIQ KQGQGQWTYQ 410 420 430 440 | | WWTEYWQATW IPEWEFVNTP PLVKLWYQLE KEPIVGAETF MEKEGKISKI GPENPYNTPV FAIKKKDSTK WRKLVDFREL NKRTQDFWEV QLGIPHPAGL GIRYQYNVLP QGWKGSPAIF QSSMTKILEP FRKQNPDIVI YQYMDDLYVG SDLEIGQHRT VQPIVLPEKD SWTVNDIQKL VGKLNWASQI YPGIKVRQLC KLLRGTKALT EVIPLTEEAE IYQEPFKNLK TGKYARMRGA HTNDVKQLTE AVQKITTESI VIWGKTPKFK LPIQKETWET TITAS MALLICK, UNIVERSITY OF CALCUTTA
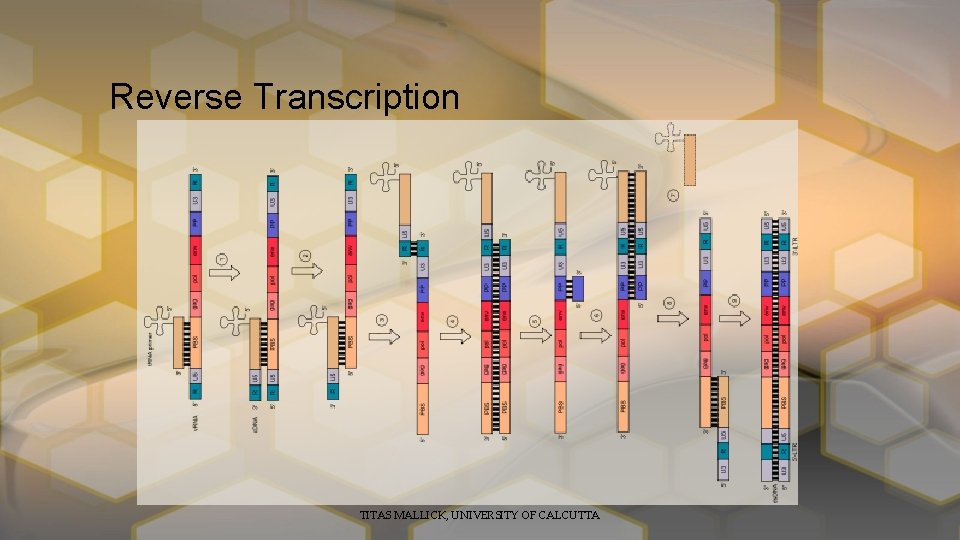
Reverse Transcription TITAS MALLICK, UNIVERSITY OF CALCUTTA

information • RT is a DNA/RNA dependant DNA polymerase. • Have a plam, finger and thumb domain. • Active Open conformation is denoted A form. • An intermediate form also characterized termed H form • Kd for DNA/DNA templte-Primer efficiency of RT is 4 μM whereas it is 14 μM for RNA/DNA • In active conformation it bends the DNA/RNA hybrid at -40 degree • DNA/DNA bending is also reported by some workers, thus making the RNA/DNA reverse transcription much efficient. TITAS MALLICK, UNIVERSITY OF CALCUTTA

continue • Important stage in the procedure is host t. RNAlys 3 is used as primer • Two sharp strand transfer is needed to generate both the +ve and ve sense strand RNA. • One theory suggests as c. DNA synthesis processes overtime the RTc matures by conformational changes into a complex called PIC, that later imported into the nucleus and get involved in some host gene interfering processes. TITAS MALLICK, UNIVERSITY OF CALCUTTA

Main project: identification of RT inhibitor • Accumulation of biomass. • Isolation and purification of RT. • Quantification. • Plant Crude extraction. • Protein isolation. • Exclusion and Screening. • RT functional assay. • Pico-green assay. • In-vivo assay. TITAS MALLICK, UNIVERSITY OF CALCUTTA

BEGINNING WITH CULTURE • The HIV RT GENE is placed in a vector in it’s MCS site under an IPTG inducible enhancer. • The plasmid was transferred in the E. coli strain BL 21(DE 3) • The first batch of culture begun with reviving the cells in liquid media for overnight in 37 degree. • Then the primary culture was established in 37 degree. • From the late log phase of the primary culture the secondary culture was seeded, when it reached a OD of 0. 6 then IPTG added for induction. TITAS MALLICK, UNIVERSITY OF CALCUTTA

TO DO • The secondary cultures must be centrifuged for the collection of the biomass. • This phase should be repeated several time for collection of proper amount of biomass. • The induced RT then must be isolated from the biomass • For this process we must use hexa-His tag affinity chromatography • Then this proteins must be assayed for quantification (Barford’s), dialysis must be performed for condensation of the proteins. TITAS MALLICK, UNIVERSITY OF CALCUTTA

PLANT SCREENING • We must begin with a rough initial screening and then proceed from a material with previous reports of RT inhibition. • Choice of plants 1. Catharanthus roseus (L. ) G. Don (Apocynaceae) 2. Moringa (Moringaceae) 3. Carica papaya L. (Caricaceae) 4. Ocimum basilicum L. (Lamiaceae) TITAS MALLICK, UNIVERSITY OF CALCUTTA

Next Stages • The Proteins must be isolated from the plant extract after proper cleaning and drying of the plant leaves. • The protein fractionation must be done by salting in. • Then other processes is done. • Nickel column chromatography is performed for separation of different phases of the solution. • Different fractions are dissolved in different grades of non-polar to polar solvents. • Each fractions are tested for the RT inhibition activity in vitro possibly through Pico-green Assay. TITAS MALLICK, UNIVERSITY OF CALCUTTA

Functional RT assay • Isolated RT in vitro tested by rt. PCR and gel electrophoresis if it is functional or not. • MLV RT used as positive control. TITAS MALLICK, UNIVERSITY OF CALCUTTA

Pico-green assay • Pico green binds with only DNA/DNA or DNA/RNA hybrid but not with ss. RNA or ss. DNA. • Only pico green shows a high pick. • Pico green+RNA shows low pick. • Pico green+RNA+RT shows fall in pick if RT functional. • Pico green+RNA+RT+inhibitor tested if show fall in the pick it indicates the inhibitor inhibiting RT. TITAS MALLICK, UNIVERSITY OF CALCUTTA

Next Stages • After a fraction showing its inhibition action further exclusion stages are performed. • To identify a single molecule interacting with the RT for its inhibition we must use first molecular docking and GCMS for proper identification of the molecule, TITAS MALLICK, UNIVERSITY OF CALCUTTA

NEXT • After the in vitro assays we must go for some in cell assay where we should introduce both the RT and the inhibitory protein in the cell line and will test for its action. • Before working in cell culture the compunds CC 50 level is measured. The compounds are tested only below the cytotoxicity level. TITAS MALLICK, UNIVERSITY OF CALCUTTA

Current status • Primary culture had been seeded • Secondary culture was inoculated • Induction was done. • The work stopped here. TITAS MALLICK, UNIVERSITY OF CALCUTTA

THANK YOU. TITAS MALLICK, UNIVERSITY OF CALCUTTA

reference. • Wikipedia • Structure and function of HIV-1 reverse transcriptase: molecular mechanisms of polymerization and inhibition: Stefan G. Sarafianos et al. NCBI-NLM-NIH • HIV-1 Reverse Transcription: Wei-Shau Hu et al. NCBI-NLM-NIH • Strand transfer events during HIV-1 reverse transcription: Basu VP et al. NCBI-NLMNIH. • Jmol Proteopedia. • http: //www. bioafrica. net/proteomics • The Mechano-Chemistry of Reverse Transcriptase: Omri Malik et al. ; The CELL TITAS MALLICK, UNIVERSITY OF CALCUTTA
- Slides: 31