Hiram Clawson U C Santa Cruz Genome Browser

Hiram Clawson U. C. Santa Cruz Genome Browser 09 June 2016

Genome Assemblies In the beginning: a. one b. two c. many Acceptance: UCSC NCBI Ensembl Standardization: A: GENBANK B: REFSEQ www. ncbi. nlm. nih. gov/assembly/ up to date copy of this presentation: genomewiki. ucsc. edu/index. php/File: Bejerano. Lab_2016 -06 -09. pptx

Naming schemes • • UCSC: chr 1, chr 2, … chr 22, chr. X, chr. Y, chr. M Ensembl: 1, 2, … 22, X, Y, MT Genbank: CM 000663. 2, CM 000664. 2 Refseq: NC_000001. 11, NC_000002. 12 chr 16_KI 270728 v 1_random chr 6_GL 000256 v 2_alt chr. Un_KI 270442 v 1

UCSC genomes • • hg 38 – 38 th human genome assembly mm 10 – 10 th Mus musculus assembly gor. Gor 4 – 4 th Gorilla gorilla pan. Pan 2 – 2 nd Pan paniscus (bonobo) hgsql -e 'select name, organism, scientific. Name, tax. Id from db. Db; ' hgcentral Count: 277 total public site, 164 species, 516 total hgwdev, 331 species

Assembly hubs Prototype experiment • genome-preview. cse. ucsc. edu/gbdb/hubs/refseq/ • 33, 772 species, 50, 718 assemblies • genome-preview. cse. ucsc. edu/gbdb/hubs/genbank/ • 36, 944 species, 62, 096 assemblies • • from downloads: ftp: //ftp. ncbi. nlm. nih. gov/genomes/refseq/ ftp: //ftp. ncbi. nlm. nih. gov/genomes/genbank/ (can also rsync from these sources)
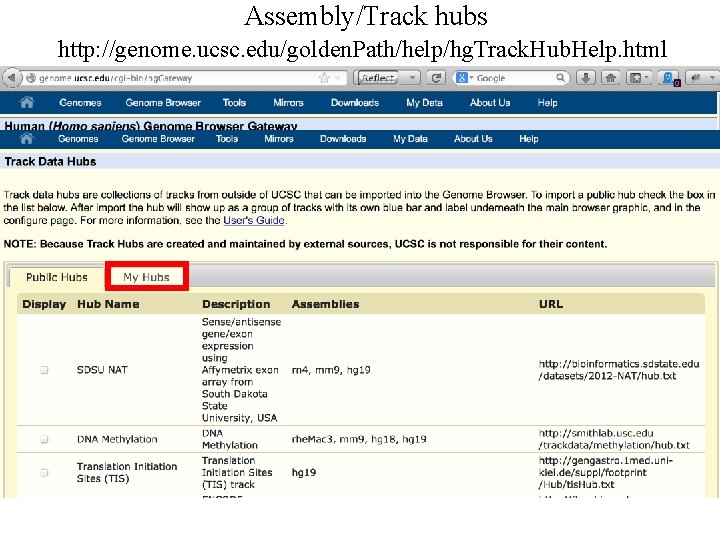
Assembly/Track hubs http: //genome. ucsc. edu/golden. Path/help/hg. Track. Hub. Help. html
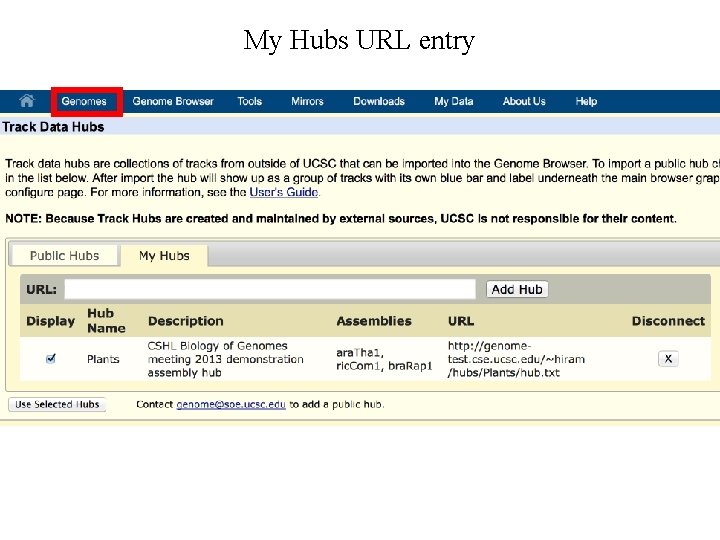
My Hubs URL entry
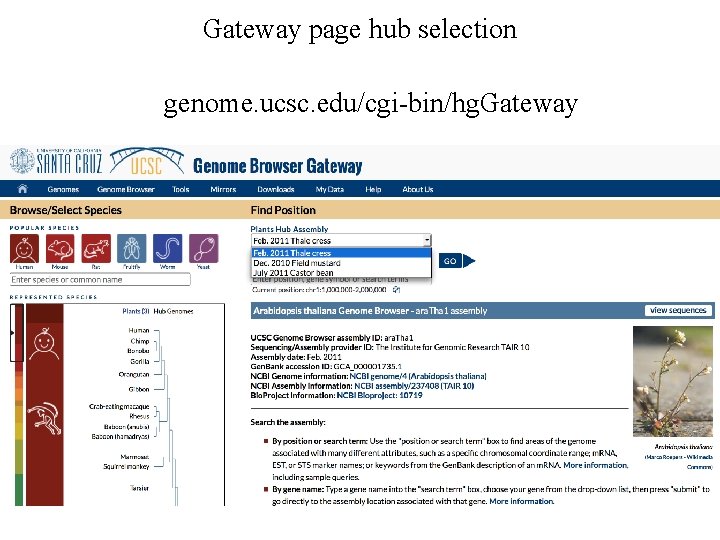
Gateway page hub selection genome. ucsc. edu/cgi-bin/hg. Gateway
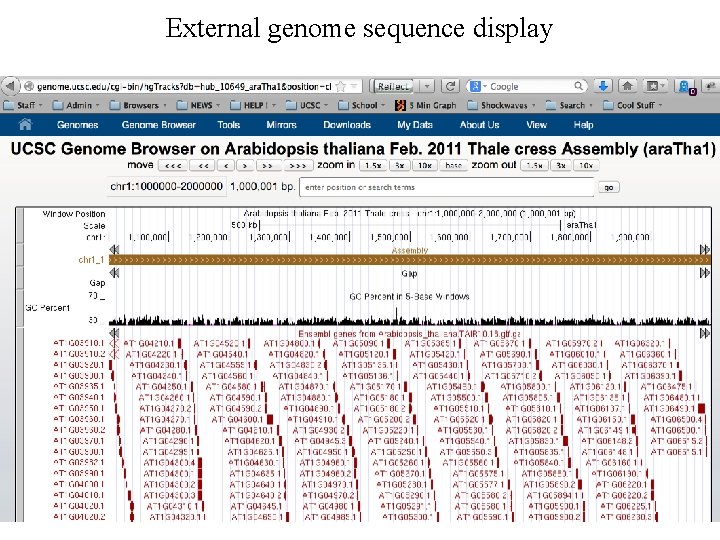
External genome sequence display
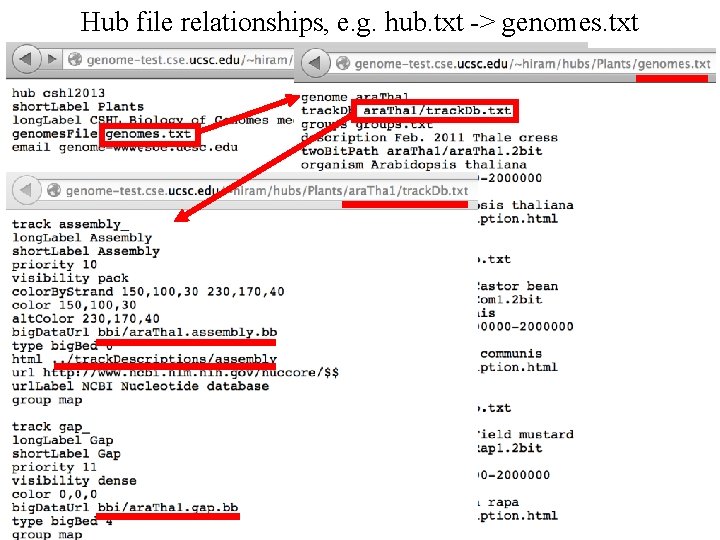
Hub file relationships, e. g. hub. txt -> genomes. txt

genome to genome alignments 1. establish systematic directory layout for all work (this is a bookkeeping exercise) 2. partition genomes a. 20 to 40 million bases target genome, 10, 000 overlap b. 10 to 20 million bases query genome limit number of sequences in one chunk to ~100 to 400 thus: ~250 target pieces, ~400 query pieces, cluster jobs: ~100, 000 = 250 * 400 3. run lastz on each query chunk to each target chunk www. bx. psu. edu/~rsharris/lastz/ results in psl file format. A couple of hours to perhaps two weeks of cluster time. (much hand waving here, many options for alignment)

‘chain’ psl alignments 3. run axt. Chain on all psl files for each chromosome: zcat. . /psl. Parts/$1*. psl. gz | axt. Chain -psl -verbose=0 -score. Scheme=human_chimp. v 2. q -min. Score=3000 -linear. Gap=medium stdin pan. Pan 2. 2 bit gor. Gor 4. 2 bit stdout | chain. Anti. Repeat pan. Pan 2. 2 bit gor. Gor 4. 2 bit stdin $2 www. pnas. org/content/100/20/11484. abstract

construct ‘nets’ from chains # Make nets ("no. Class", i. e. without rmsk/class stats which are added later): chain. Pre. Net pan. Pan 2. gor. Gor 4. pan. Pan 2. chrom. sizes gor. Gor 4. chrom. sizes stdout | chain. Net stdin -min. Space=1 pan. Pan 2. chrom. sizes gor. Gor 4. chrom. sizes stdout /dev/null | net. Syntenic stdin no. Class. net # Make lift. Over chains: net. Chain. Subset -verbose=0 no. Class. net pan. Pan 2. gor. Gor 4. all. chain. gz stdout | chain. Stitch. Id stdin stdout | gzip -c > pan. Pan 2. gor. Gor 4. over. chain. gz www. pnas. org/content/100/20/11484. abstract

loading chain/net tracks # Load chains: hg. Load. Chain -t. Index pan. Pan 2 chain. Gor 4 pan. Pan 2. gor. Gor 4. all. chain. gz # Add gap/repeat stats to the net file using database tables: net. Class -verbose=0 -no. Ar no. Class. net pan. Pan 2 gor. Gor 4 pan. Pan 2. gor. Gor 4. net # Load nets: net. Filter -min. Gap=10 pan. Pan 2. gor. Gor 4. net | hg. Load. Net -verbose=0 pan. Pan 2 net. Gor 4 stdin # Measure genome to genome ‘coverage’ feature. Bits pan. Pan 2 chain. Gor 4 Link www. pnas. org/content/100/20/11484. abstract 1, 536 pair-wise lastz/chain/net alignments: hgwdev-hiram. cse. ucsc. edu/cgi-bin/100 way/lastz. pl
- Slides: 14