Herpes Simplex Virus HSV and EpsteinBarr Virus EBV
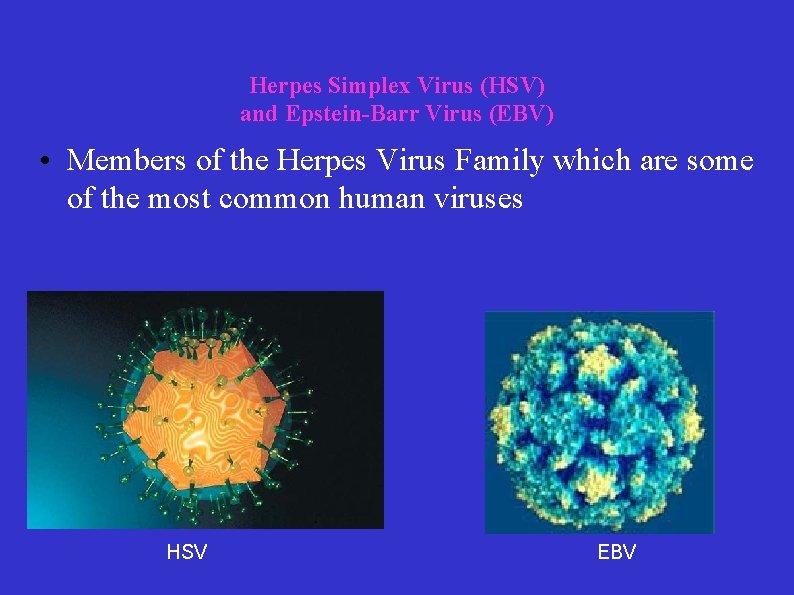
Herpes Simplex Virus (HSV) and Epstein-Barr Virus (EBV) • Members of the Herpes Virus Family which are some of the most common human viruses HSV EBV

How was the HSV Discovered • HSV is an ancient disease with descriptions of orolabial herpes appearing in records from the fifth century BCE. • In the 1736 Astruc (physician to King of France) identified Herpes as a cause of genital infection. • Over the next 50 years many different strains of herpes were discovered. • In 1893 intimate human-to-human transmission was identified. • In the 1940’s neonatal HSV infection was described and finally identified in 1968.
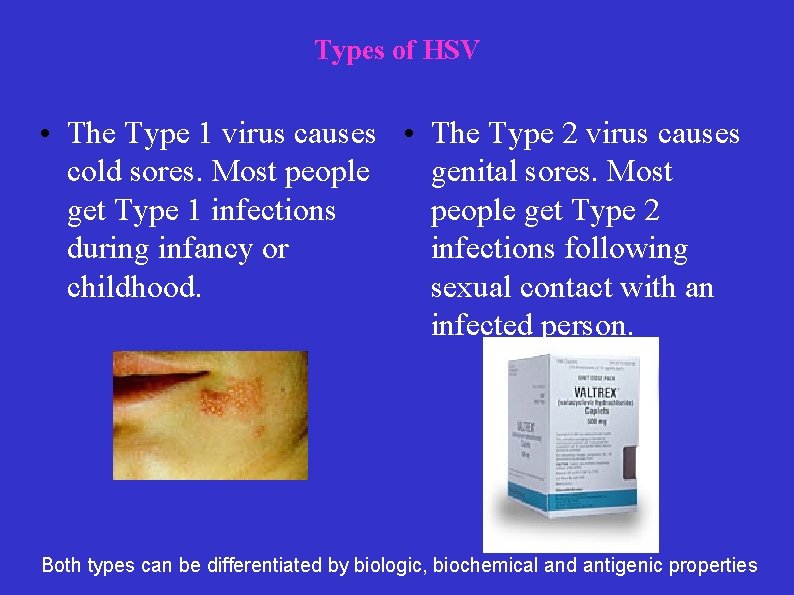
Types of HSV • The Type 1 virus causes • The Type 2 virus causes cold sores. Most people genital sores. Most get Type 1 infections people get Type 2 during infancy or infections following childhood. sexual contact with an infected person. Both types can be differentiated by biologic, biochemical and antigenic properties

HSV Viral Structure • Composed of a ds. DNA (152 kbp) nucleoprotein core • Core is surrounded by an icosahedral protein capsid • 100 nm Capsid is enclosed in an outer envelope consisting of at least 8 glycoproteins. • Envelope spikes ~8 nm long • The virus requires a moist environment for survival.

Humans, HSV Hosts simians, cattle, cats and chickens. Of these, only herpes virus simiae is harmful to us.
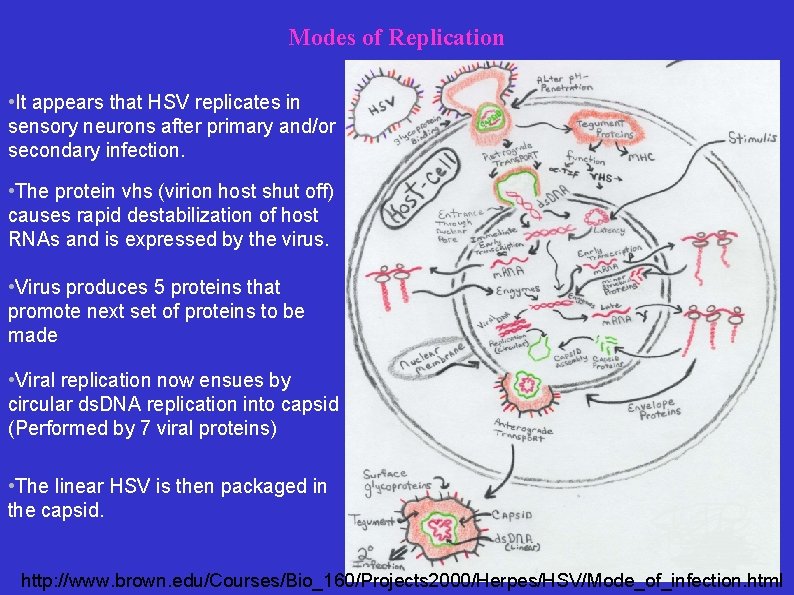
Modes of Replication • It appears that HSV replicates in sensory neurons after primary and/or secondary infection. • The protein vhs (virion host shut off) causes rapid destabilization of host RNAs and is expressed by the virus. • Virus produces 5 proteins that promote next set of proteins to be made • Viral replication now ensues by circular ds. DNA replication into capsid (Performed by 7 viral proteins) • The linear HSV is then packaged in the capsid. http: //www. brown. edu/Courses/Bio_160/Projects 2000/Herpes/HSV/Mode_of_infection. html
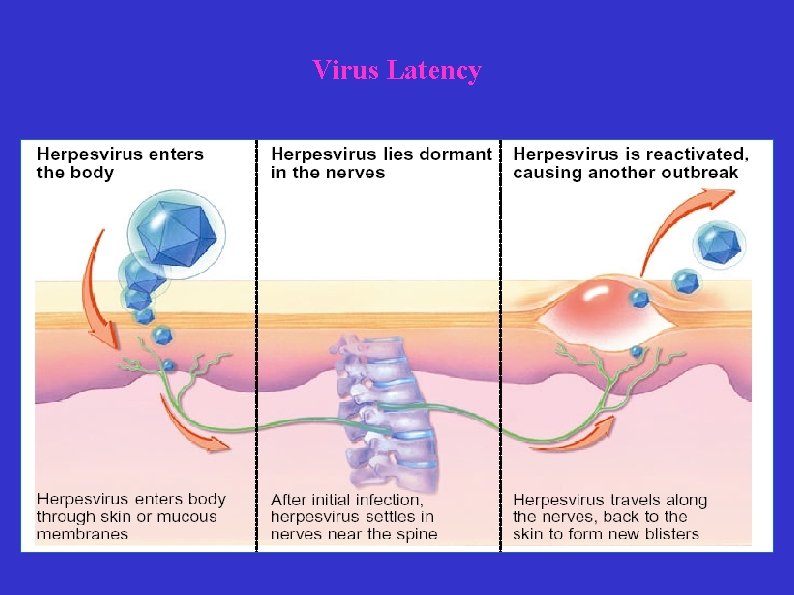
Virus Latency
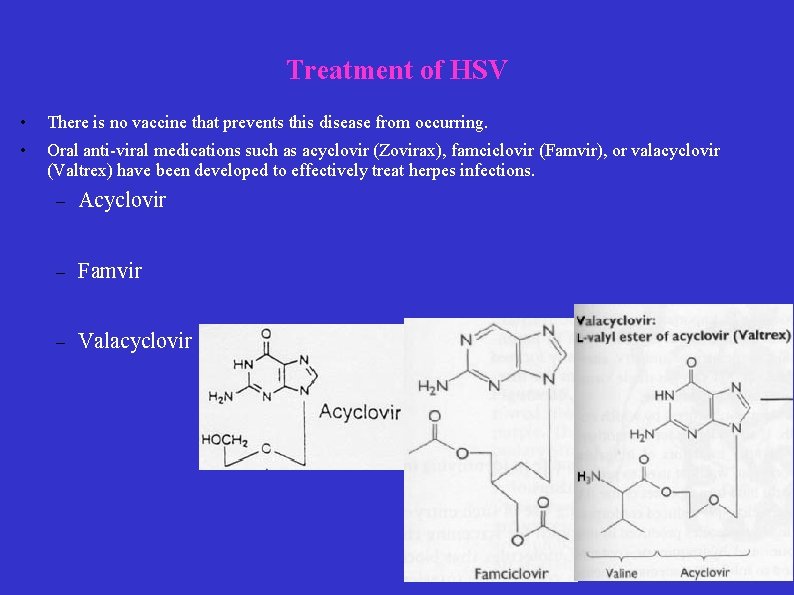
Treatment of HSV • There is no vaccine that prevents this disease from occurring. • Oral anti-viral medications such as acyclovir (Zovirax), famciclovir (Famvir), or valacyclovir (Valtrex) have been developed to effectively treat herpes infections. – Acyclovir – Famvir – Valacyclovir

Discovery of EBV and Important Dates • 1958 - Burkitt's lymphoma described in the malaria belt of east Africa • 1964 - Epstein and Barr discover EBV through electron microscopy of cells cultured from Burkitt's lymphoma tissue • 1968 - EBV demonstrated as the etiological agent of infectious mononucleosis

EBV Viral Structure • A core containing a linear, ds. DNA molecule of about 175 kbp. • An icosahedral capsid, approximately 100 -110 nm in diameter, containing 162 capsomeres with a hole running down the long axis. • An amorphous, sometimes asymmetric material that surrounds the capsid, designated as the tegument • An envelope containing viral glycoprotein spikes on its surface.

Modes of Transmission • Intimate Contact – kissing, sharing food, coughing
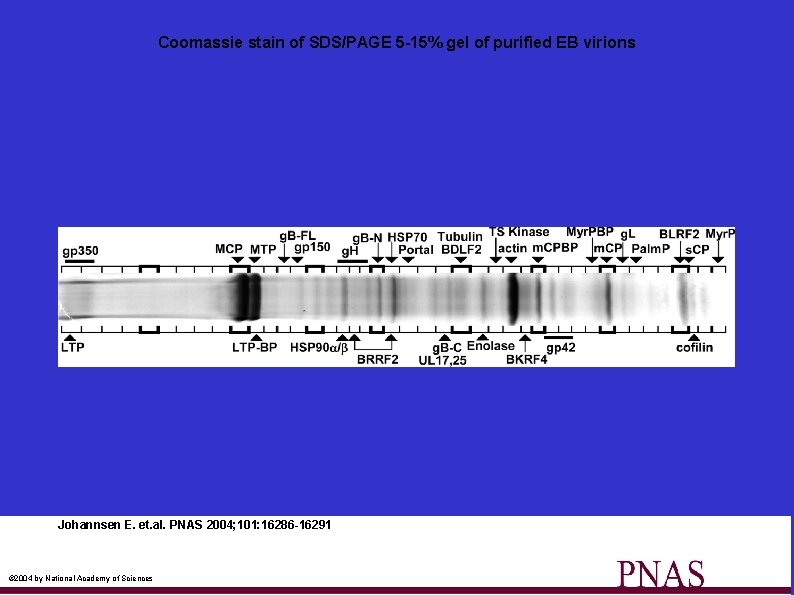
Coomassie stain of SDS/PAGE 5 -15% gel of purified EB virions Johannsen E. et. al. PNAS 2004; 101: 16286 -16291 © 2004 by National Academy of Sciences
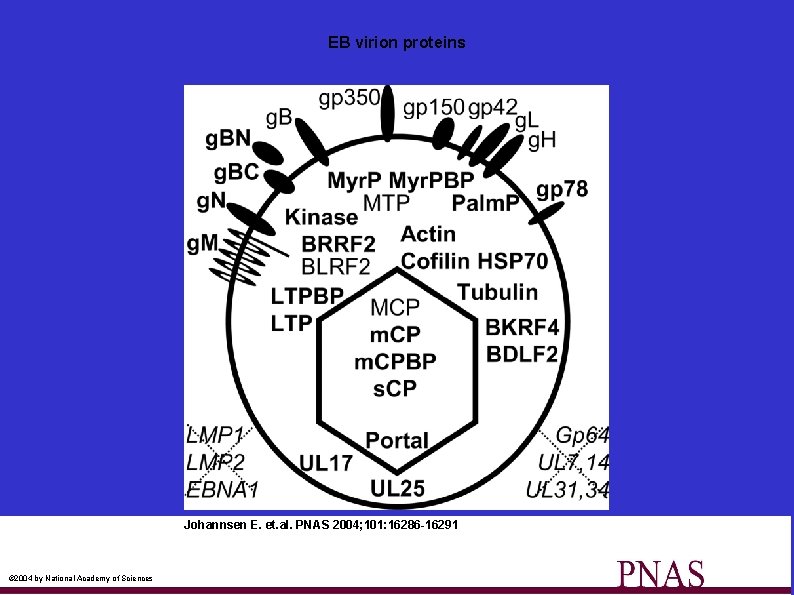
EB virion proteins Johannsen E. et. al. PNAS 2004; 101: 16286 -16291 © 2004 by National Academy of Sciences
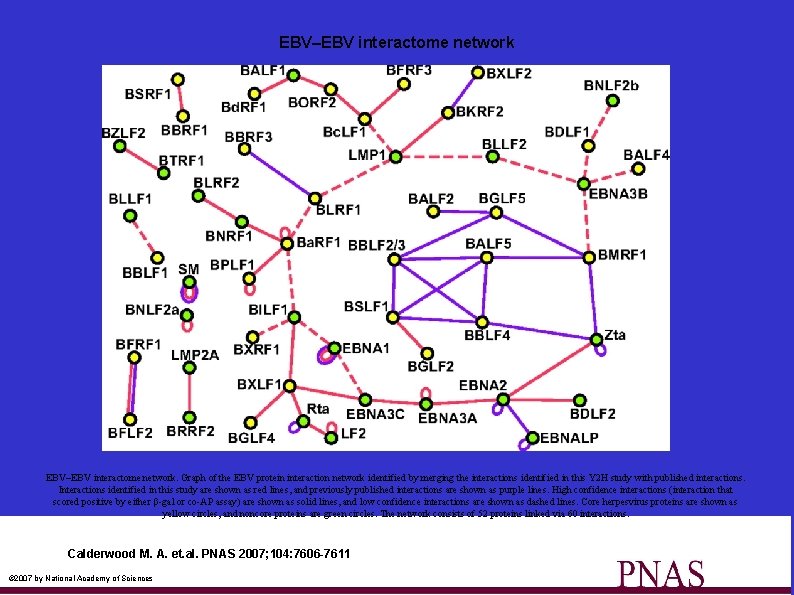
EBV–EBV interactome network. Graph of the EBV protein interaction network identified by merging the interactions identified in this Y 2 H study with published interactions. Interactions identified in this study are shown as red lines, and previously published interactions are shown as purple lines. High confidence interactions (interaction that scored positive by either β-gal or co-AP assay) are shown as solid lines, and low confidence interactions are shown as dashed lines. Core herpesvirus proteins are shown as yellow circles, and noncore proteins are green circles. The network consists of 52 proteins linked via 60 interactions. Calderwood M. A. et. al. PNAS 2007; 104: 7606 -7611 © 2007 by National Academy of Sciences

EBV–human interactome network graph of the EBV–human protein interaction network as determined by our Y 2 H screen. Core herpesvirus proteins are shown as yellow circles, and noncore herpesvirus proteins are green circles. Human proteins are shown as blue squares. Interactions identified in this screen are shown as red lines. This interactome represents 40 EBV proteins and 112 human proteins connected by 173 interactions. Calderwood M. A. et. al. PNAS 2007; 104: 7606 -7611 © 2007 by National Academy of Sciences
- Slides: 15