HCS Clustering Algorithm A Clustering Algorithm Based on

HCS Clustering Algorithm A Clustering Algorithm Based on Graph Connectivity

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 2

The Problem • Clustering: – Group elements into subsets based on similarity between pairs of elements • Requirements: – Elements in the same cluster are highly similar to each other – Elements in different clusters have low similarity to each other • Challenges: – Large sets of data – Inaccurate and noisy measurements ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 3

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 4

HCS Algorithm Overview • Highly Connected Subgraphs Algorithm – Uses graph theoretic techniques • Basic Idea – Uses similarity information to construct a similarity graph – Groups elements that are highly connected with each other ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 5

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 6
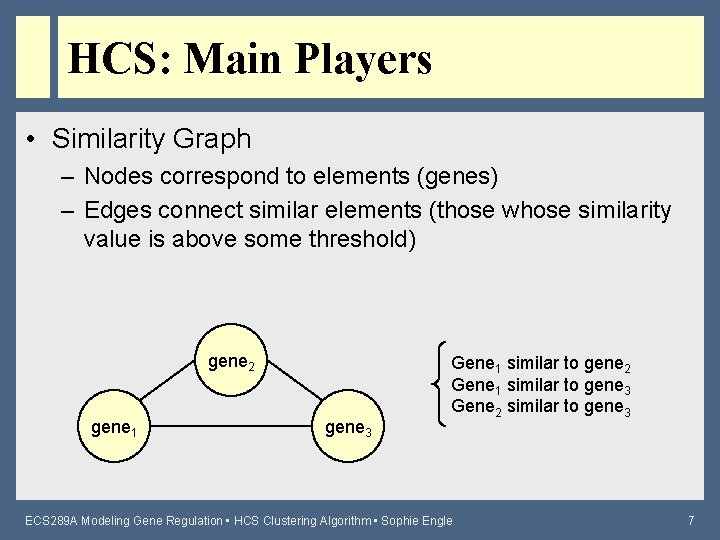
HCS: Main Players • Similarity Graph – Nodes correspond to elements (genes) – Edges connect similar elements (those whose similarity value is above some threshold) gene 2 gene 1 gene 3 Gene 1 similar to gene 2 Gene 1 similar to gene 3 Gene 2 similar to gene 3 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 7
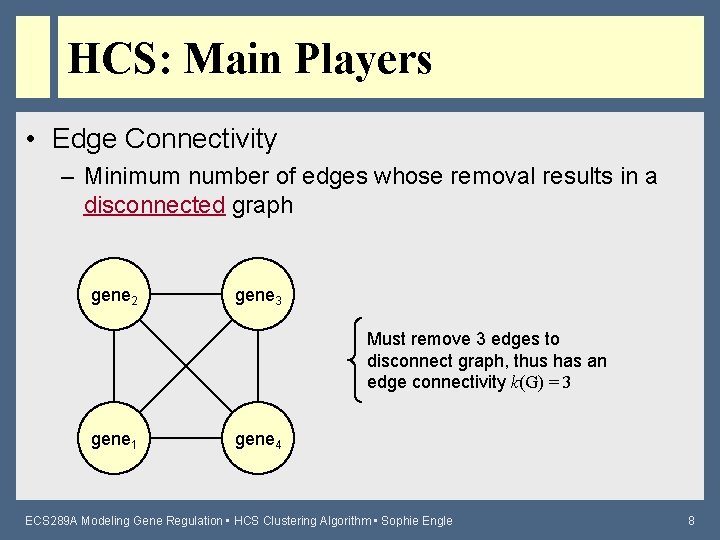
HCS: Main Players • Edge Connectivity – Minimum number of edges whose removal results in a disconnected graph gene 2 gene 3 Must remove 3 edges to disconnect graph, thus has an edge connectivity k(G) = 3 gene 1 gene 4 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 8

HCS: Main Players • Edge Connectivity – Minimum number of edges whose removal results in a disconnected graph gene 2 gene 3 Must remove 3 edges to disconnect graph, thus has an edge connectivity k(G) = 3 gene 1 gene 4 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 9
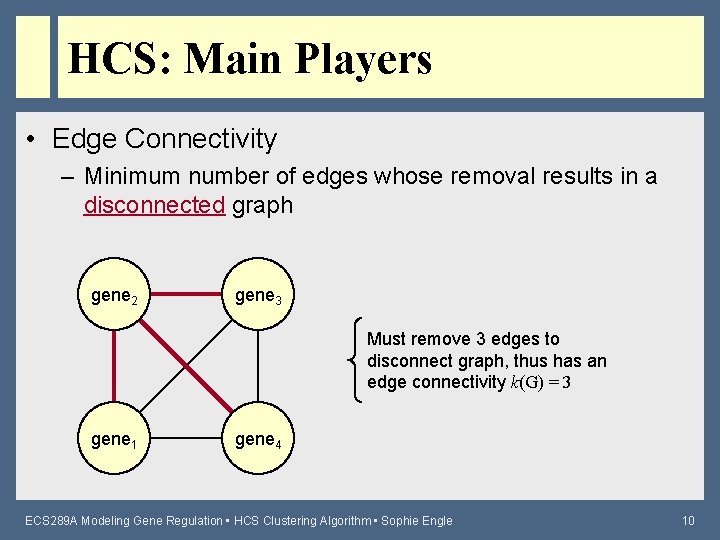
HCS: Main Players • Edge Connectivity – Minimum number of edges whose removal results in a disconnected graph gene 2 gene 3 Must remove 3 edges to disconnect graph, thus has an edge connectivity k(G) = 3 gene 1 gene 4 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 10
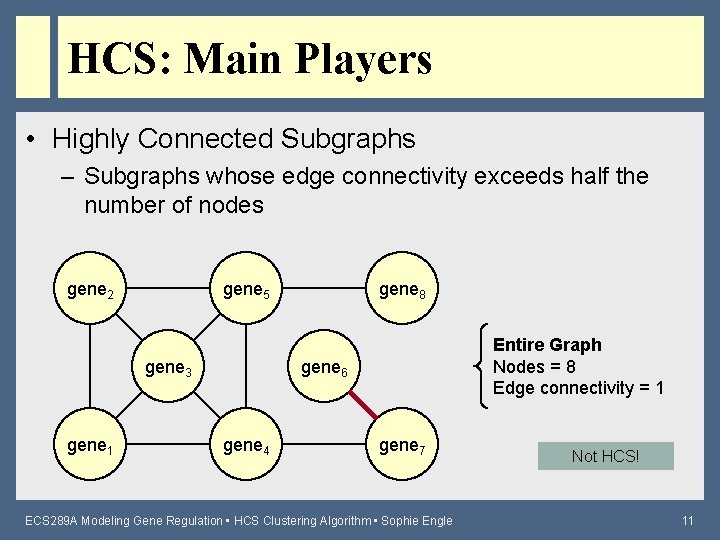
HCS: Main Players • Highly Connected Subgraphs – Subgraphs whose edge connectivity exceeds half the number of nodes gene 2 gene 5 gene 3 gene 1 gene 8 Entire Graph Nodes = 8 Edge connectivity = 1 gene 6 gene 4 gene 7 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle Not HCS! 11
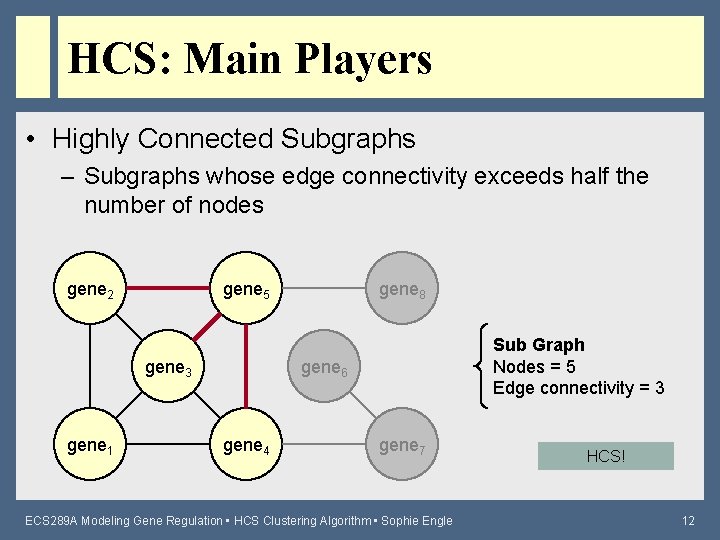
HCS: Main Players • Highly Connected Subgraphs – Subgraphs whose edge connectivity exceeds half the number of nodes gene 2 gene 5 gene 3 gene 1 gene 8 Sub Graph Nodes = 5 Edge connectivity = 3 gene 6 gene 4 gene 7 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle HCS! 12

HCS: Main Players • Cut – A set of edges whose removal disconnects the graph gene 2 gene 5 gene 3 gene 1 gene 8 gene 6 gene 4 gene 7 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 13

HCS: Main Players • Minimum Cut – A cut with a minimum number of edges gene 2 gene 5 gene 3 gene 1 gene 8 gene 6 gene 4 gene 7 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 14

HCS: Main Players • Minimum Cut – A cut with a minimum number of edges gene 2 gene 5 gene 3 gene 1 gene 8 gene 6 gene 4 gene 7 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 15
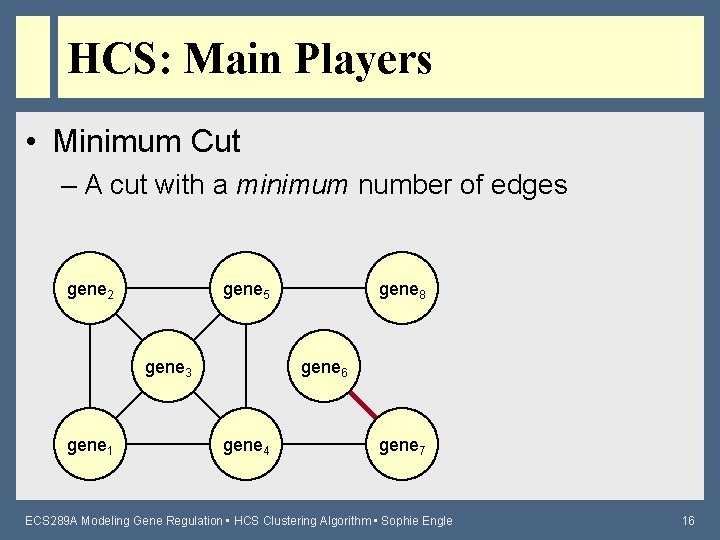
HCS: Main Players • Minimum Cut – A cut with a minimum number of edges gene 2 gene 5 gene 3 gene 1 gene 8 gene 6 gene 4 gene 7 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 16

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 17

HCS: Algorithm (by example) 5 2 3 4 6 12 11 10 7 9 8 1 find and remove a minimum cut ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 18
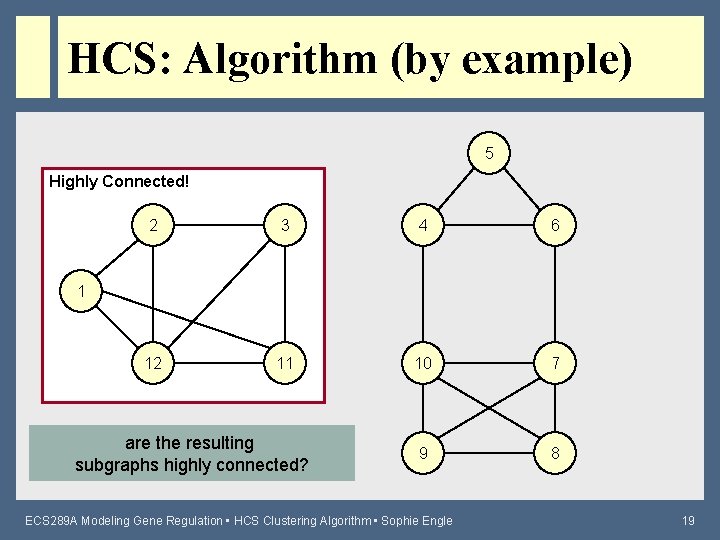
HCS: Algorithm (by example) 5 Highly Connected! 2 3 4 6 12 11 10 7 9 8 1 are the resulting subgraphs highly connected? ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 19
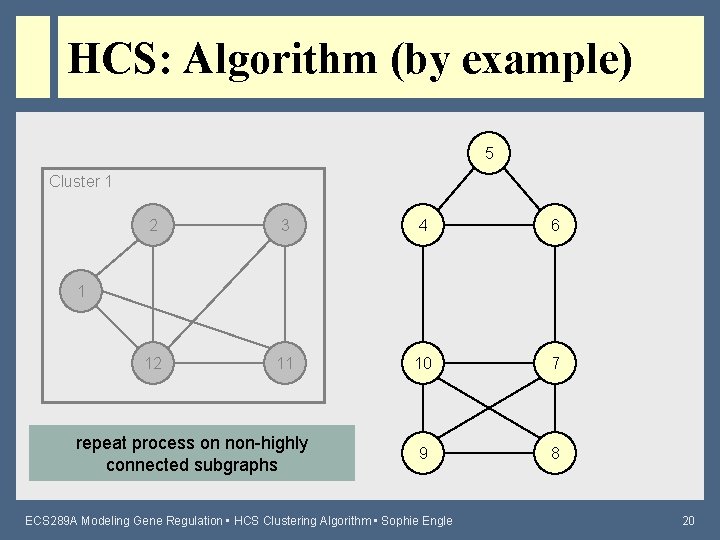
HCS: Algorithm (by example) 5 Cluster 1 2 3 4 6 12 11 10 7 9 8 1 repeat process on non-highly connected subgraphs ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 20
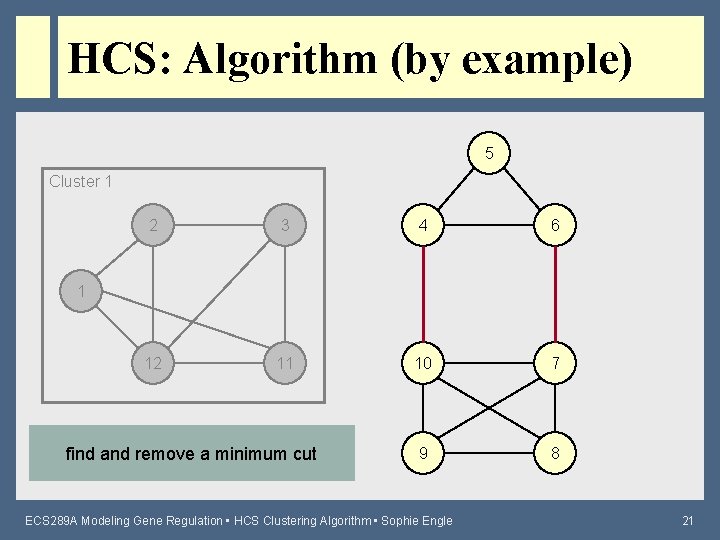
HCS: Algorithm (by example) 5 Cluster 1 2 3 4 6 12 11 10 7 9 8 1 find and remove a minimum cut ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 21

HCS: Algorithm (by example) 5 Cluster 1 2 3 4 6 Highly Connected! 12 11 are the resulting subgraphs highly connected? 10 7 9 8 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 22
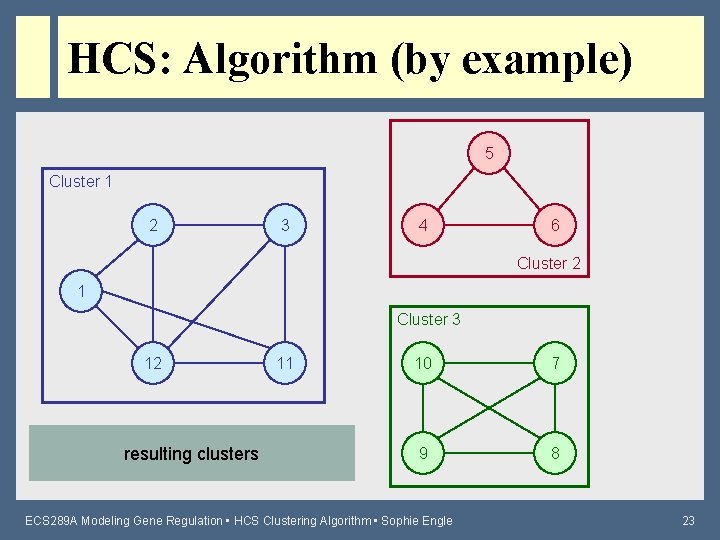
HCS: Algorithm (by example) 5 Cluster 1 2 3 4 6 Cluster 2 1 Cluster 3 12 resulting clusters 11 10 7 9 8 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 23

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > n ÷ 2 return Hi else HCS( Hi ) } } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 24

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > n ÷ 2 return Hi else Find a minimum cut in graph G. This returns a set of subgraphs HCS( Hi ) { H , … , H } resulting from the removal of the cut set. } 1 t } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 25

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > n ÷ 2 return Hi else HCS( Hi ) For each subgraph… } } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 26

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > n ÷ 2 return Hi If the subgraph is highly else connected, then return that subgraph as a cluster. HCS( Hi ) (Note: k( H ) denotes edge connectivity of graph H , n } i i } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle denotes number of nodes) 27

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > n ÷ 2 return Hi Otherwise, repeat the algorithm else on the subgraph. (recursive function) HCS( Hi ) This continues until there are } no more subgraphs, and all } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle clusters have been found. 28

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > n ÷ 2 return Hi Running time is bounded by else 2 N × f( n, m ) where N is the number of clusters found, HCS( Hi ) and f( n, m ) is the time complexity of computing a } minimum cut in a graph with n } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle nodes and m edges. 29

HCS: Algorithm HCS( G ) { MINCUT( G ) = { H 1, … , Ht } for each Hi, i = [ 1, t ] { if k( Hi ) > Deterministic n ÷ 2 for Graph: return Un-weighted Hi takes O(nm) steps else where n is the number nodes and m is the HCS( Hiof ) number of edges } } ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 30

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 31

HCS: Properties • Homogeneity – Each cluster has a diameter of at most 2 • Distance is the minimum length path between two nodes – Determined by number of EDGES traveled between nodes • Diameter is the longest distance in the graph – Each cluster is at least half as dense as a clique • Clique is a graph with maximum possible edge connectivity a b f c e d Dist( a, d ) = 2 Dist( a, e ) = 3 Diam( G ) = 4 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle clique 32

HCS: Properties • Separation – Any non-trivial split is unlikely to have diameter of two – Number of edges removed by each iteration is linear in the size of the underlying subgraph • Compared to quadratic number of edges within final clusters • Indicates separation unless sizes are small • Does not imply number of edges removed overall ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 33

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 34
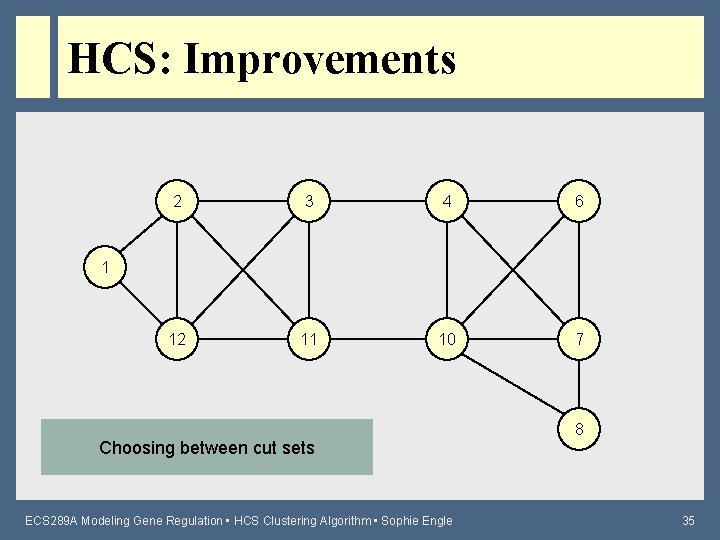
HCS: Improvements 2 3 4 6 12 11 10 7 1 Choosing between cut sets ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 8 35
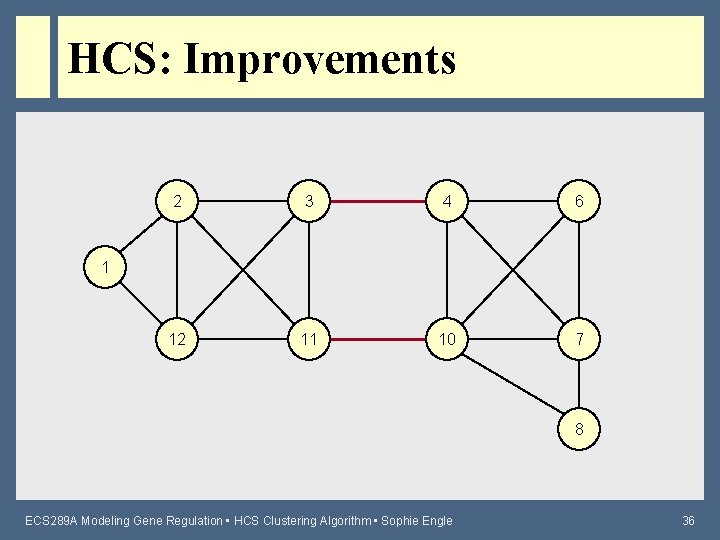
HCS: Improvements 2 3 4 6 12 11 10 7 1 8 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 36

HCS: Improvements 2 3 4 6 12 11 10 7 1 8 ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 37

HCS: Improvements • Iterated HCS – Sometimes there are multiple minimum cuts to choose from • Some cuts may create “singletons” or nodes that become disconnected from the rest of the graph – Performs several iterations of HCS until no new cluster is found (to find best final clusters) • Theoretically adds another O(n) factor to running time, but typically only needs 1 – 5 more iterations ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 38

HCS: Improvements • Remove low degree nodes first – If node has low degree, likely will just be separated from rest of graph – Calculating separation for those nodes is expensive – Removal helps eliminate unnecessary iterations and significantly reduces running time ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 39

Presentation Outline • The Problem • HCS Algorithm Overview – Main Players – General Algorithm – Properties – Improvements • Conclusion ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 40

Conclusion • Performance – With improvements, can handle problems with up to thousands of elements in reasonable computing time – Generates clusters with high homogeneity and separation – More robust (responds better when noise is introduced) than other approaches based on connectivity ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 41

References “A Clustering Algorithm based on Graph Connectivity” By Erez Hartuv and Ron Shamir March 1999 ( Revised December 1999) http: //www. math. tau. ac. il/~rshamir/papers. html ECS 289 A Modeling Gene Regulation • HCS Clustering Algorithm • Sophie Engle 42
- Slides: 42